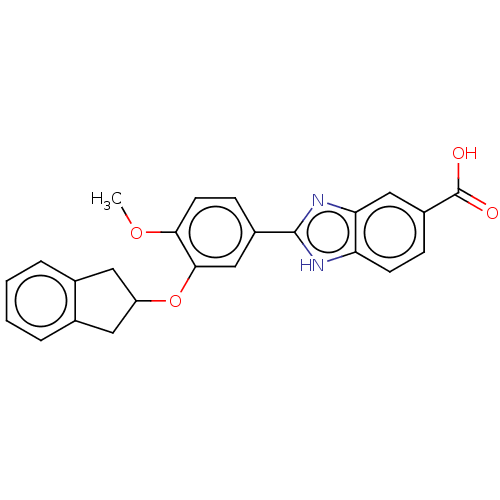
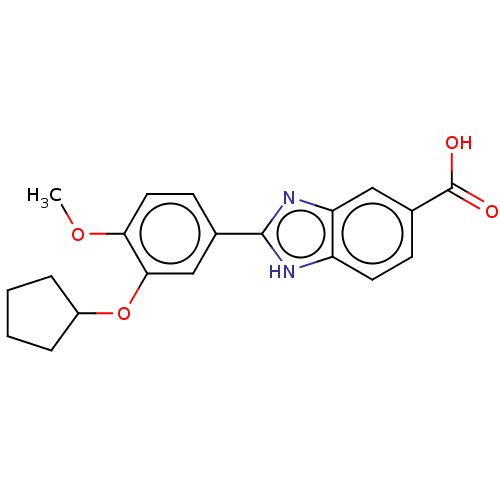
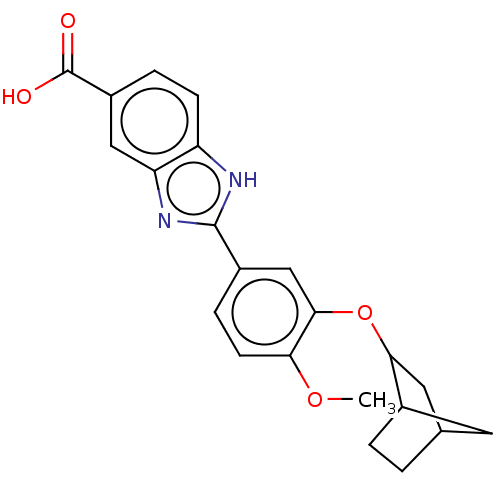
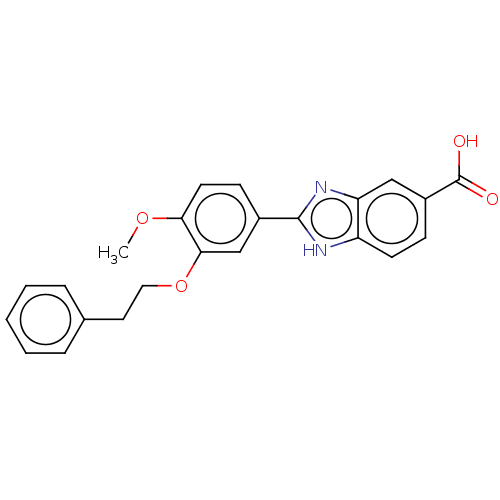
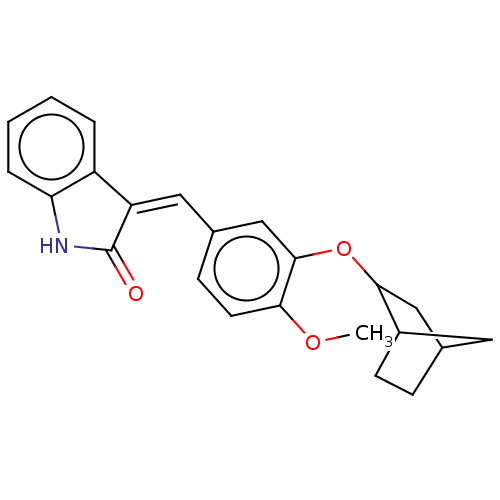
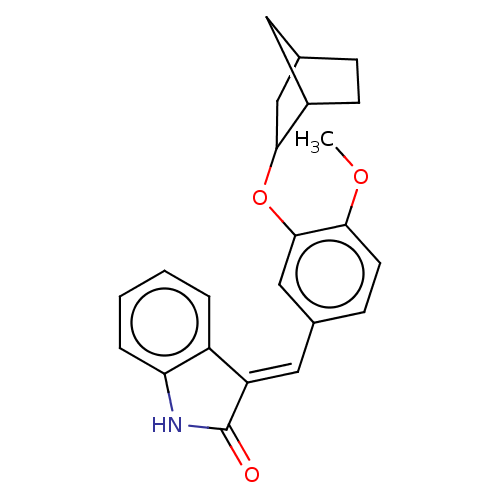
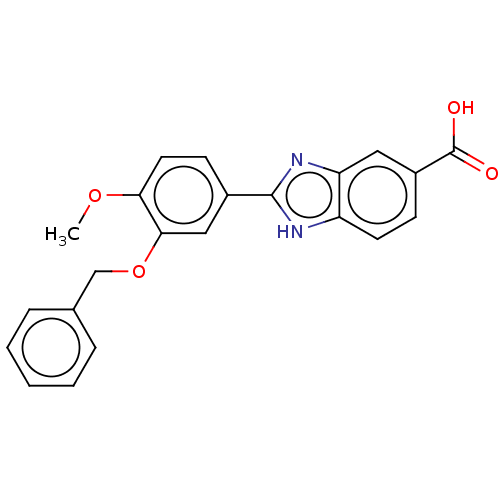
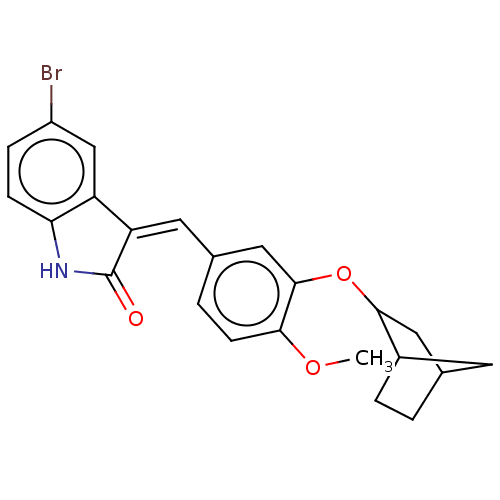
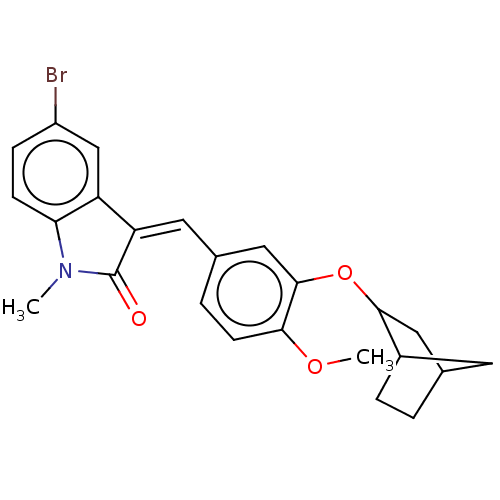
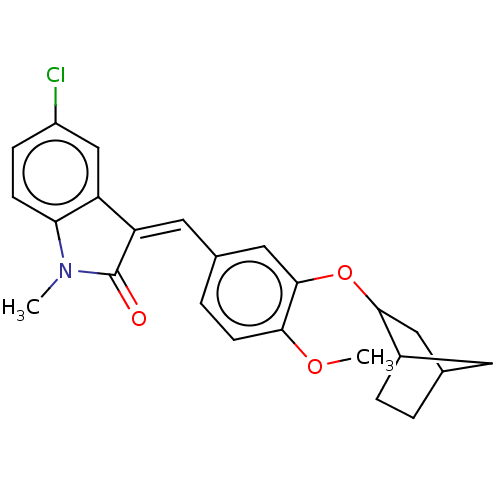
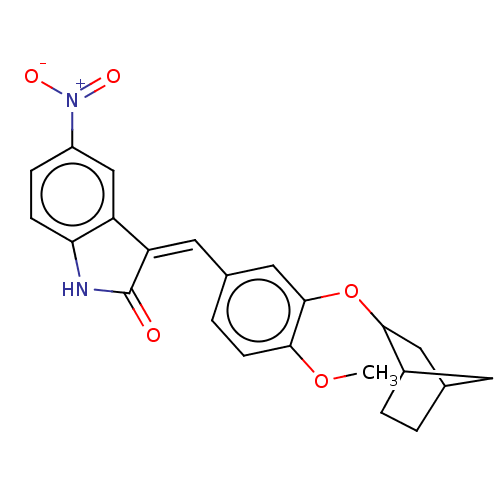
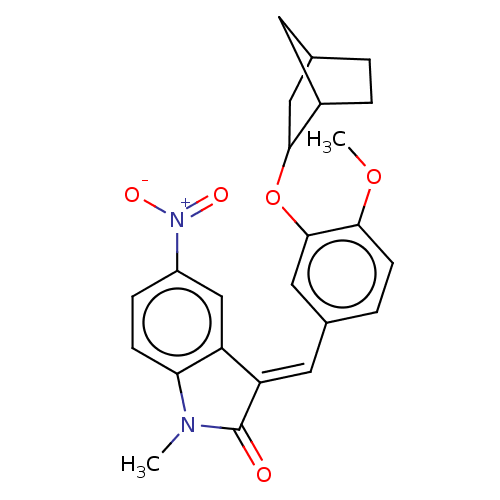
Affinity DataIC50: 4nMAssay Description:Inhibition of [3H]rolipram binding to Phosphodiesterase 4 (PDE IV)More data for this Ligand-Target Pair
Affinity DataIC50: 4nMAssay Description:Displacement of [3H]-Rolipram from mouse brain homogenatesMore data for this Ligand-Target Pair
Affinity DataIC50: 32nMAssay Description:Displacement of [3H]-Rolipram from mouse brain homogenatesMore data for this Ligand-Target Pair
Affinity DataIC50: 78nMAssay Description:Displacement of [3H]-Rolipram from mouse brain homogenatesMore data for this Ligand-Target Pair
Affinity DataIC50: 130nMAssay Description:Displacement of [3H]-Rolipram from mouse brain homogenatesMore data for this Ligand-Target Pair
Affinity DataIC50: 220nMAssay Description:Displacement of [3H]-Rolipram from mouse brain homogenatesMore data for this Ligand-Target Pair
Affinity DataIC50: 3.20E+3nMAssay Description:Inhibition of [3H]rolipram binding to Phosphodiesterase 4 (PDE IV)More data for this Ligand-Target Pair
Affinity DataIC50: 4.60E+3nMAssay Description:Inhibition of [3H]rolipram binding to Phosphodiesterase 4 (PDE IV)More data for this Ligand-Target Pair
Affinity DataIC50: 5.91E+3nMAssay Description:Displacement of [3H]-Rolipram from mouse brain homogenatesMore data for this Ligand-Target Pair
Affinity DataIC50: 5.00E+4nMAssay Description:Inhibition of [3H]rolipram binding to Phosphodiesterase 4 (PDE IV)More data for this Ligand-Target Pair
Affinity DataIC50: 5.70E+4nMAssay Description:Inhibition of [3H]rolipram binding to Phosphodiesterase 4 (PDE IV)More data for this Ligand-Target Pair
Affinity DataIC50: 5.80E+4nMAssay Description:Inhibition of [3H]rolipram binding to Phosphodiesterase 4 (PDE IV)More data for this Ligand-Target Pair
Affinity DataIC50: 6.20E+4nMAssay Description:Inhibition of [3H]rolipram binding to Phosphodiesterase 4 (PDE IV)More data for this Ligand-Target Pair
Affinity DataIC50: 6.60E+4nMAssay Description:Inhibition of [3H]rolipram binding to Phosphodiesterase 4 (PDE IV)More data for this Ligand-Target Pair
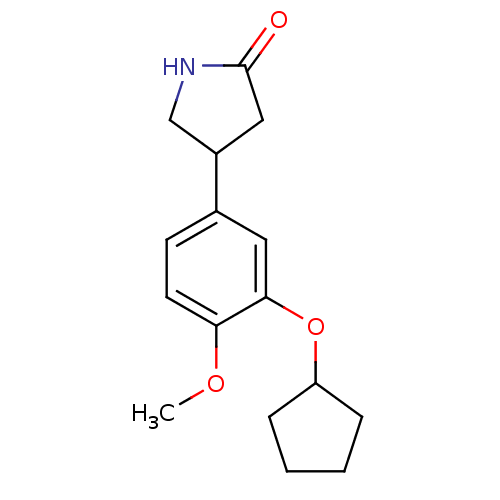
 3D Structure (crystal)
3D Structure (crystal)