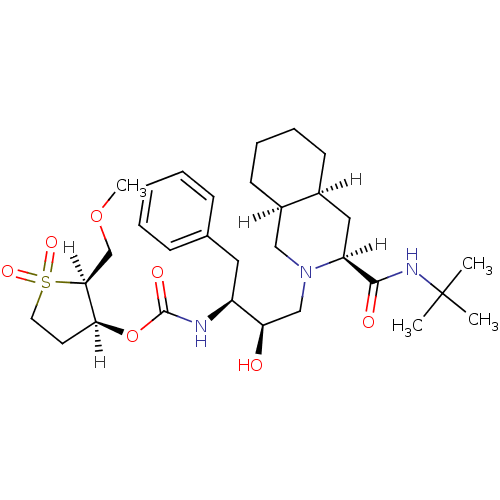
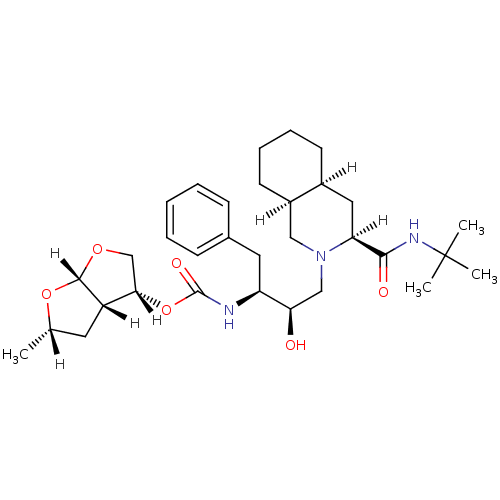
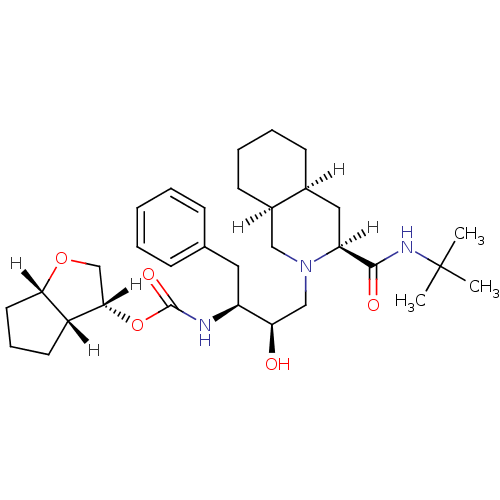
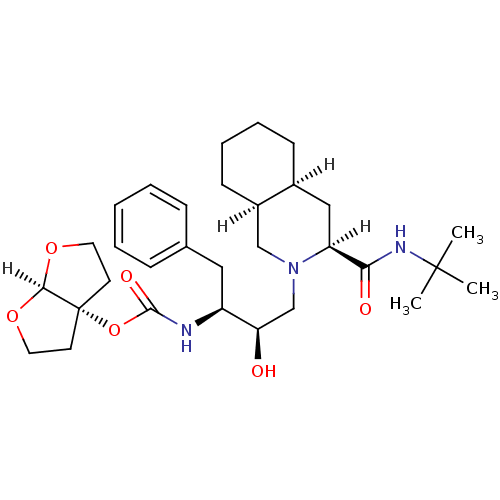
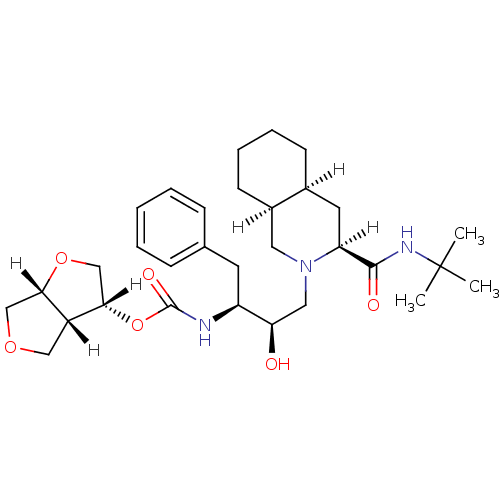
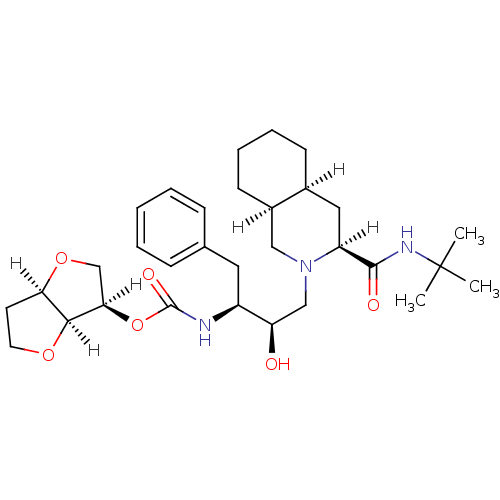
Report error Found 14 Enz. Inhib. hit(s) with all data for entry = 324
TargetDimer of Gag-Pol polyprotein [489-587](Human immunodeficiency virus type 1)
University of Illinois At Chicago
University of Illinois At Chicago
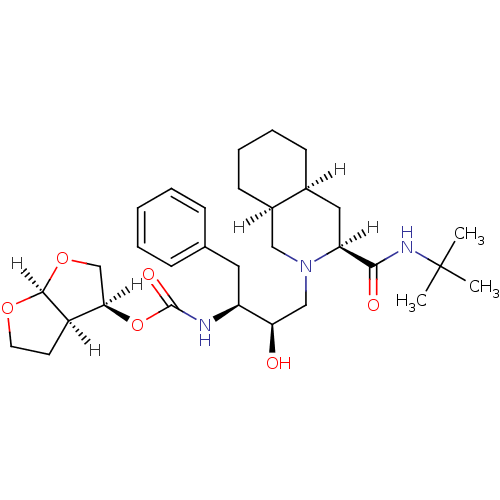
Affinity DataIC50: 1.20nMpH: 5.5 T: 2°CAssay Description:The IC50 values for the compounds were determined using purified HIV-1 Protease. Inhibition of the cleavage of the peptide H-Val-Ser-Gln-Asn-(L-beta-...More data for this Ligand-Target Pair
TargetDimer of Gag-Pol polyprotein [489-587](Human immunodeficiency virus type 1)
University of Illinois At Chicago
University of Illinois At Chicago
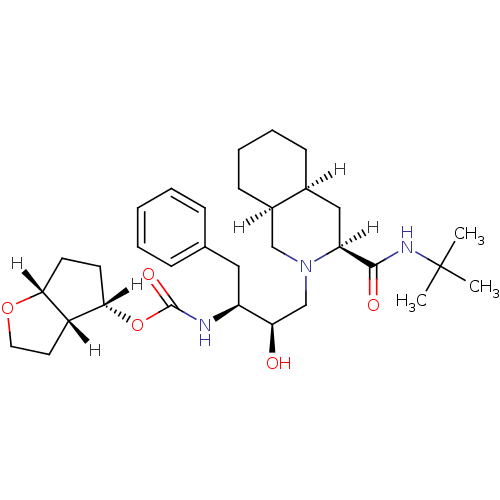
Affinity DataIC50: 1.80nMpH: 5.5 T: 2°CAssay Description:The IC50 values for the compounds were determined using purified HIV-1 Protease. Inhibition of the cleavage of the peptide H-Val-Ser-Gln-Asn-(L-beta-...More data for this Ligand-Target Pair
TargetDimer of Gag-Pol polyprotein [489-587](Human immunodeficiency virus type 1)
University of Illinois At Chicago
University of Illinois At Chicago
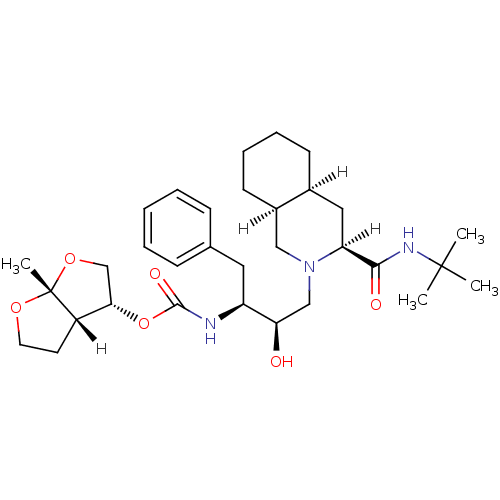
Affinity DataIC50: 4.10nMpH: 5.5 T: 2°CAssay Description:The IC50 values for the compounds were determined using purified HIV-1 Protease. Inhibition of the cleavage of the peptide H-Val-Ser-Gln-Asn-(L-beta-...More data for this Ligand-Target Pair
TargetDimer of Gag-Pol polyprotein [489-587](Human immunodeficiency virus type 1)
University of Illinois At Chicago
University of Illinois At Chicago
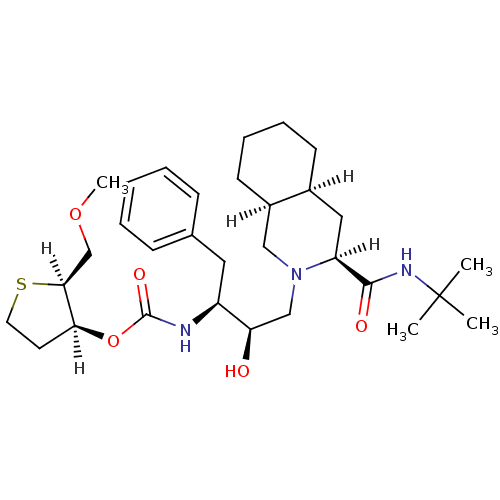
Affinity DataIC50: 4.20nMpH: 5.5 T: 2°CAssay Description:The IC50 values for the compounds were determined using purified HIV-1 Protease. Inhibition of the cleavage of the peptide H-Val-Ser-Gln-Asn-(L-beta-...More data for this Ligand-Target Pair
TargetDimer of Gag-Pol polyprotein [489-587](Human immunodeficiency virus type 1)
University of Illinois At Chicago
University of Illinois At Chicago
Affinity DataIC50: 6.40nMpH: 5.5 T: 2°CAssay Description:The IC50 values for the compounds were determined using purified HIV-1 Protease. Inhibition of the cleavage of the peptide H-Val-Ser-Gln-Asn-(L-beta-...More data for this Ligand-Target Pair
TargetDimer of Gag-Pol polyprotein [489-587](Human immunodeficiency virus type 1)
University of Illinois At Chicago
University of Illinois At Chicago
Affinity DataIC50: 17nMpH: 5.5 T: 2°CAssay Description:The IC50 values for the compounds were determined using purified HIV-1 Protease. Inhibition of the cleavage of the peptide H-Val-Ser-Gln-Asn-(L-beta-...More data for this Ligand-Target Pair
TargetDimer of Gag-Pol polyprotein [489-587](Human immunodeficiency virus type 1)
University of Illinois At Chicago
University of Illinois At Chicago
Affinity DataIC50: 18.1nMpH: 5.5 T: 2°CAssay Description:The IC50 values for the compounds were determined using purified HIV-1 Protease. Inhibition of the cleavage of the peptide H-Val-Ser-Gln-Asn-(L-beta-...More data for this Ligand-Target Pair
TargetDimer of Gag-Pol polyprotein [489-587](Human immunodeficiency virus type 1)
University of Illinois At Chicago
University of Illinois At Chicago
Affinity DataIC50: 19.7nMpH: 5.5 T: 2°CAssay Description:The IC50 values for the compounds were determined using purified HIV-1 Protease. Inhibition of the cleavage of the peptide H-Val-Ser-Gln-Asn-(L-beta-...More data for this Ligand-Target Pair
TargetDimer of Gag-Pol polyprotein [489-587](Human immunodeficiency virus type 1)
University of Illinois At Chicago
University of Illinois At Chicago
Affinity DataIC50: 23.8nMpH: 5.5 T: 2°CAssay Description:The IC50 values for the compounds were determined using purified HIV-1 Protease. Inhibition of the cleavage of the peptide H-Val-Ser-Gln-Asn-(L-beta-...More data for this Ligand-Target Pair
TargetDimer of Gag-Pol polyprotein [489-587](Human immunodeficiency virus type 1)
University of Illinois At Chicago
University of Illinois At Chicago
Affinity DataIC50: 27.1nMpH: 5.5 T: 2°CAssay Description:The IC50 values for the compounds were determined using purified HIV-1 Protease. Inhibition of the cleavage of the peptide H-Val-Ser-Gln-Asn-(L-beta-...More data for this Ligand-Target Pair
TargetDimer of Gag-Pol polyprotein [489-587](Human immunodeficiency virus type 1)
University of Illinois At Chicago
University of Illinois At Chicago
Affinity DataIC50: 190nMpH: 5.5 T: 2°CAssay Description:The IC50 values for the compounds were determined using purified HIV-1 Protease. Inhibition of the cleavage of the peptide H-Val-Ser-Gln-Asn-(L-beta-...More data for this Ligand-Target Pair
TargetDimer of Gag-Pol polyprotein [489-587](Human immunodeficiency virus type 1)
University of Illinois At Chicago
University of Illinois At Chicago
Affinity DataIC50: 410nMpH: 5.5 T: 2°CAssay Description:The IC50 values for the compounds were determined using purified HIV-1 Protease. Inhibition of the cleavage of the peptide H-Val-Ser-Gln-Asn-(L-beta-...More data for this Ligand-Target Pair
TargetDimer of Gag-Pol polyprotein [489-587](Human immunodeficiency virus type 1)
University of Illinois At Chicago
University of Illinois At Chicago
Affinity DataIC50: 1.70E+3nMpH: 5.5 T: 2°CAssay Description:The IC50 values for the compounds were determined using purified HIV-1 Protease. Inhibition of the cleavage of the peptide H-Val-Ser-Gln-Asn-(L-beta-...More data for this Ligand-Target Pair
TargetDimer of Gag-Pol polyprotein [489-587](Human immunodeficiency virus type 1)
University of Illinois At Chicago
University of Illinois At Chicago
Affinity DataIC50: 3.00E+3nMpH: 5.5 T: 2°CAssay Description:The IC50 values for the compounds were determined using purified HIV-1 Protease. Inhibition of the cleavage of the peptide H-Val-Ser-Gln-Asn-(L-beta-...More data for this Ligand-Target Pair