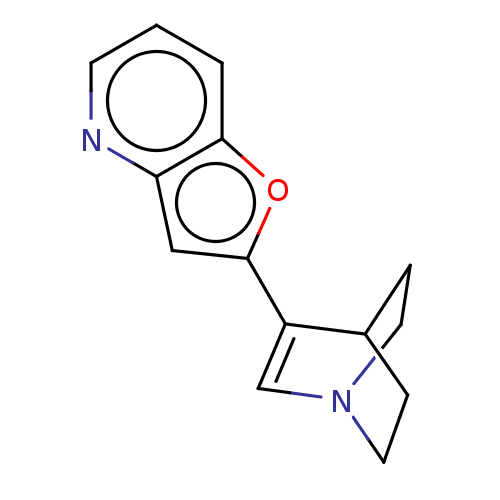
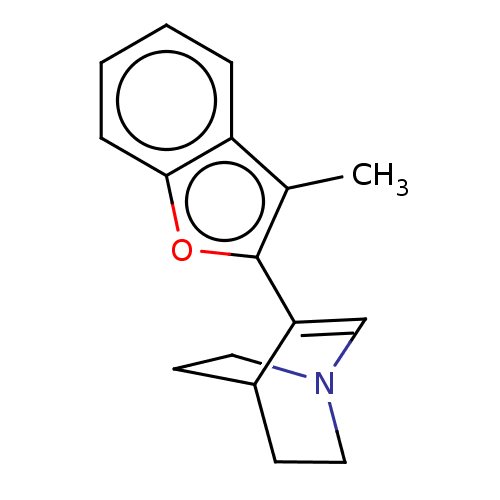
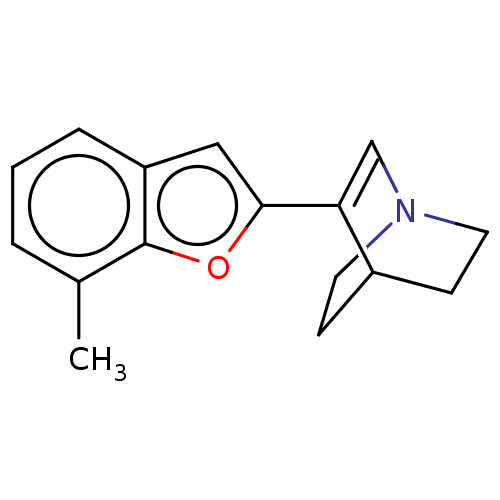
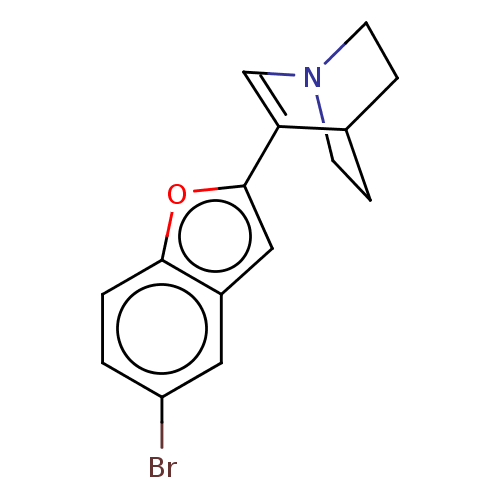
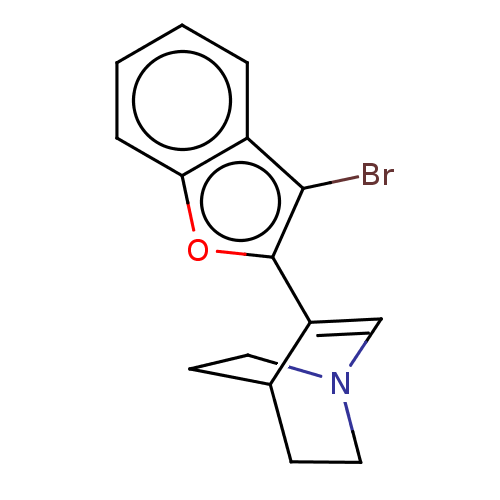
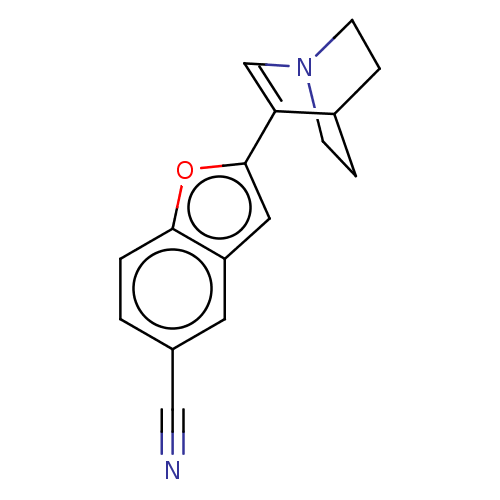
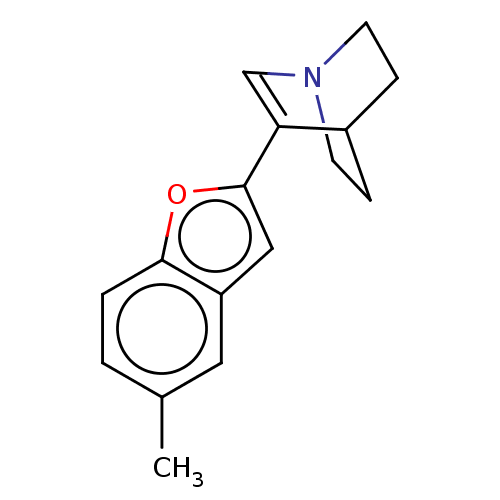
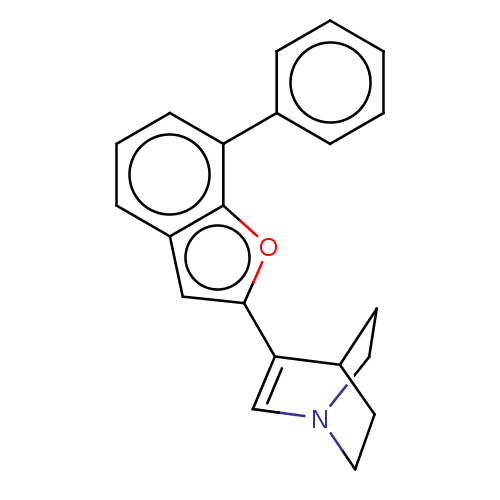
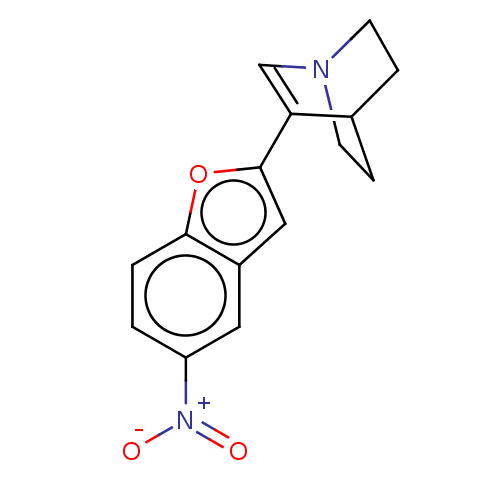
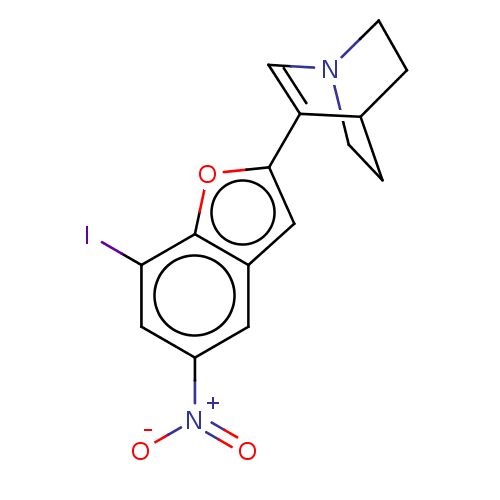
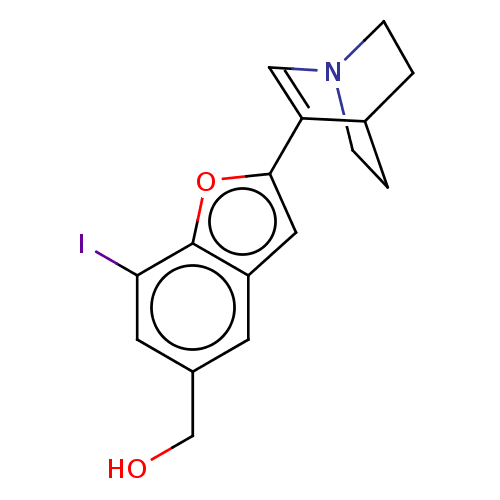
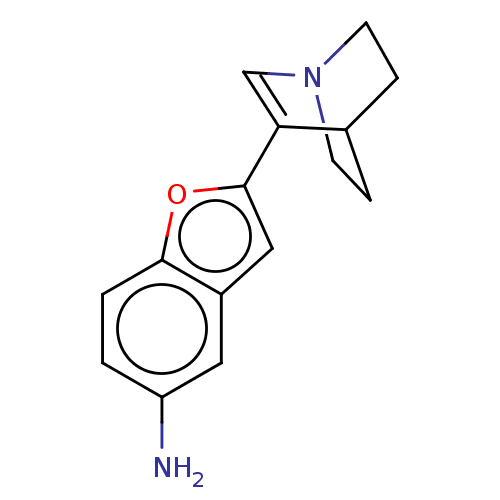
Report error Found 100 Enz. Inhib. hit(s) with all data for entry = 50003434
Affinity DataKi: 6.30nMAssay Description:Receptor binding affinity against Muscarinic acetylcholine receptor from guinea pig cerebral cortex was determined using [3H]- QNB as radioligandMore data for this Ligand-Target Pair
Affinity DataKi: 9.60nMAssay Description:Receptor binding affinity (Ki) against Muscarinic acetylcholine receptor (cerebral cortex) was determined by competition radioligand -[3H]- QNB bindi...More data for this Ligand-Target Pair
Affinity DataKi: 15nMAssay Description:Receptor binding affinity (Ki) against Muscarinic acetylcholine receptor (cerebral cortex) was determined by competition radioligand -[3H]- QNB bindi...More data for this Ligand-Target Pair
Affinity DataKi: 20nMAssay Description:Receptor binding affinity (Ki) against Muscarinic acetylcholine receptor (cerebral cortex) was determined by competition radioligand -[3H]- QNB bindi...More data for this Ligand-Target Pair
Affinity DataKi: 21nMAssay Description:Receptor binding affinity against Muscarinic acetylcholine receptor from guinea pig cerebral cortex was determined using [3H]- QNB as radioligandMore data for this Ligand-Target Pair
Affinity DataKi: 25nMAssay Description:Receptor binding affinity (Ki) against Muscarinic acetylcholine receptor (cerebral cortex) was determined by competition radioligand -[3H]- QNB bindi...More data for this Ligand-Target Pair
Affinity DataKi: 25nMAssay Description:Receptor binding affinity (Ki) against Muscarinic acetylcholine receptor (cerebral cortex) was determined by competition radioligand -[3H]- QNB bindi...More data for this Ligand-Target Pair
Affinity DataKi: 30nMAssay Description:Receptor binding affinity against Muscarinic acetylcholine receptor from guinea pig parotid gland was determined using [3H]- QNB as radioligandMore data for this Ligand-Target Pair
Affinity DataKi: 31nMAssay Description:Receptor binding affinity (Ki) for Muscarinic acetylcholine receptor (heart)was determined by competition radioligand -[3H]- QNB binding assayMore data for this Ligand-Target Pair
Affinity DataKi: 34nMAssay Description:Receptor binding affinity against Muscarinic acetylcholine receptor from guinea pig heart was determined using [3H]- QNB as radioligandMore data for this Ligand-Target Pair
Affinity DataKi: 37nMAssay Description:Receptor binding affinity against Muscarinic acetylcholine receptor from guinea pig urinary bladder was determined using [3H]- QNB as radioligandMore data for this Ligand-Target Pair
Affinity DataKi: 39nMAssay Description:Receptor binding affinity (Ki) against Muscarinic acetylcholine receptor (cerebral cortex) was determined by competition radioligand -[3H]- QNB bindi...More data for this Ligand-Target Pair
Affinity DataKi: 40nMAssay Description:Receptor binding affinity (Ki) against Muscarinic acetylcholine receptor (cerebral cortex) was determined by competition radioligand -[3H]- QNB bindi...More data for this Ligand-Target Pair
Affinity DataKi: 44nMAssay Description:Receptor binding affinity (Ki) for Muscarinic acetylcholine receptor (heart)was determined by competition radioligand -[3H]- QNB binding assayMore data for this Ligand-Target Pair
Affinity DataKi: 56nMAssay Description:Receptor binding affinity (Ki) for Muscarinic acetylcholine receptor (parotid glands) was determined by competition radioligand -[3H]- QNB binding as...More data for this Ligand-Target Pair
Affinity DataKi: 59nMAssay Description:Receptor binding affinity (Ki) for Muscarinic acetylcholine receptor (parotid glands) was determined by competition radioligand -[3H]- QNB binding as...More data for this Ligand-Target Pair
Affinity DataKi: 60nMAssay Description:Receptor binding affinity (Ki) against Muscarinic acetylcholine receptor (cerebral cortex) was determined by competition radioligand -[3H]- QNB bindi...More data for this Ligand-Target Pair
Affinity DataKi: 61nMAssay Description:Receptor binding affinity against Muscarinic acetylcholine receptor from guinea pig heart was determined using [3H]- QNB as radioligandMore data for this Ligand-Target Pair
Affinity DataKi: 67nMAssay Description:Receptor binding affinity (Ki) for Muscarinic acetylcholine receptor (urinary bladder) was determined by competition radioligand -[3H]- QNB binding a...More data for this Ligand-Target Pair
Affinity DataKi: 72nMAssay Description:Receptor binding affinity (Ki) for Muscarinic acetylcholine receptor (heart)was determined by competition radioligand -[3H]- QNB binding assayMore data for this Ligand-Target Pair
Affinity DataKi: 76nMAssay Description:Receptor binding affinity (Ki) for Muscarinic acetylcholine receptor (parotid glands) was determined by competition radioligand -[3H]- QNB binding as...More data for this Ligand-Target Pair
Affinity DataKi: 76nMAssay Description:Receptor binding affinity (Ki) for Muscarinic acetylcholine receptor (heart)was determined by competition radioligand -[3H]- QNB binding assayMore data for this Ligand-Target Pair
Affinity DataKi: 78nMAssay Description:Receptor binding affinity (Ki) for Muscarinic acetylcholine receptor (heart)was determined by competition radioligand -[3H]- QNB binding assayMore data for this Ligand-Target Pair
Affinity DataKi: 83nMAssay Description:Receptor binding affinity against Muscarinic acetylcholine receptor from guinea pig urinary bladder was determined using [3H]- QNB as radioligandMore data for this Ligand-Target Pair
Affinity DataKi: 86nMAssay Description:Receptor binding affinity (Ki) for Muscarinic acetylcholine receptor (heart)was determined by competition radioligand -[3H]- QNB binding assayMore data for this Ligand-Target Pair
Affinity DataKi: 97nMAssay Description:Receptor binding affinity (Ki) for Muscarinic acetylcholine receptor (urinary bladder) was determined by competition radioligand -[3H]- QNB binding a...More data for this Ligand-Target Pair
Affinity DataKi: 106nMAssay Description:Receptor binding affinity (Ki) for Muscarinic acetylcholine receptor (heart)was determined by competition radioligand -[3H]- QNB binding assayMore data for this Ligand-Target Pair
Affinity DataKi: 107nMAssay Description:Receptor binding affinity (Ki) for Muscarinic acetylcholine receptor (urinary bladder) was determined by competition radioligand -[3H]- QNB binding a...More data for this Ligand-Target Pair
Affinity DataKi: 109nMAssay Description:Receptor binding affinity (Ki) for Muscarinic acetylcholine receptor (parotid glands) was determined by competition radioligand -[3H]- QNB binding as...More data for this Ligand-Target Pair
Affinity DataKi: 110nMAssay Description:Receptor binding affinity against Muscarinic acetylcholine receptor from guinea pig parotid gland was determined using [3H]- QNB as radioligandMore data for this Ligand-Target Pair
Affinity DataKi: 116nMAssay Description:Receptor binding affinity (Ki) against Muscarinic acetylcholine receptor (cerebral cortex) was determined by competition radioligand -[3H]- QNB bindi...More data for this Ligand-Target Pair
Affinity DataKi: 116nMAssay Description:Receptor binding affinity (Ki) for Muscarinic acetylcholine receptor (urinary bladder) was determined by competition radioligand -[3H]- QNB binding a...More data for this Ligand-Target Pair
Affinity DataKi: 135nMAssay Description:Receptor binding affinity (Ki) against Muscarinic acetylcholine receptor (cerebral cortex) was determined by competition radioligand -[3H]- QNB bindi...More data for this Ligand-Target Pair
Affinity DataKi: 139nMAssay Description:Receptor binding affinity against Muscarinic acetylcholine receptor from guinea pig cerebral cortex was determined using [3H]- QNB as radioligandMore data for this Ligand-Target Pair
Affinity DataKi: >150nMAssay Description:Receptor binding affinity against Muscarinic acetylcholine receptor from guinea pig cerebral cortex was determined using [3H]- QNB as radioligandMore data for this Ligand-Target Pair
Affinity DataKi: 156nMAssay Description:Receptor binding affinity (Ki) for Muscarinic acetylcholine receptor (urinary bladder) was determined by competition radioligand -[3H]- QNB binding a...More data for this Ligand-Target Pair
Affinity DataKi: 160nMAssay Description:Receptor binding affinity (Ki) for Muscarinic acetylcholine receptor (heart)was determined by competition radioligand -[3H]- QNB binding assayMore data for this Ligand-Target Pair
Affinity DataKi: 161nMAssay Description:Receptor binding affinity (Ki) for Muscarinic acetylcholine receptor (parotid glands) was determined by competition radioligand -[3H]- QNB binding as...More data for this Ligand-Target Pair
Affinity DataKi: 197nMAssay Description:Receptor binding affinity against Muscarinic acetylcholine receptor from guinea pig cerebral cortex was determined using [3H]- QNB as radioligandMore data for this Ligand-Target Pair
Affinity DataKi: 204nMAssay Description:Receptor binding affinity (Ki) for Muscarinic acetylcholine receptor (parotid glands) was determined by competition radioligand -[3H]- QNB binding as...More data for this Ligand-Target Pair
Affinity DataKi: 212nMAssay Description:Receptor binding affinity against Muscarinic acetylcholine receptor from guinea pig cerebral cortex was determined using [3H]- QNB as radioligandMore data for this Ligand-Target Pair
Affinity DataKi: 216nMAssay Description:Receptor binding affinity (Ki) for Muscarinic acetylcholine receptor (parotid glands) was determined by competition radioligand -[3H]- QNB binding as...More data for this Ligand-Target Pair
Affinity DataKi: 221nMAssay Description:Receptor binding affinity (Ki) against Muscarinic acetylcholine receptor (cerebral cortex) was determined by competition radioligand -[3H]- QNB bindi...More data for this Ligand-Target Pair
Affinity DataKi: 236nMAssay Description:Receptor binding affinity (Ki) for Muscarinic acetylcholine receptor (parotid glands) was determined by competition radioligand -[3H]- QNB binding as...More data for this Ligand-Target Pair
Affinity DataKi: 238nMAssay Description:Receptor binding affinity (Ki) for Muscarinic acetylcholine receptor (heart)was determined by competition radioligand -[3H]- QNB binding assayMore data for this Ligand-Target Pair
Affinity DataKi: 260nMAssay Description:Receptor binding affinity (Ki) for Muscarinic acetylcholine receptor (heart)was determined by competition radioligand -[3H]- QNB binding assayMore data for this Ligand-Target Pair
Affinity DataKi: 289nMAssay Description:Receptor binding affinity (Ki) against Muscarinic acetylcholine receptor (cerebral cortex) was determined by competition radioligand -[3H]- QNB bindi...More data for this Ligand-Target Pair
Affinity DataKi: 295nMAssay Description:Receptor binding affinity (Ki) for Muscarinic acetylcholine receptor (parotid glands) was determined by competition radioligand -[3H]- QNB binding as...More data for this Ligand-Target Pair
Affinity DataKi: 314nMAssay Description:Receptor binding affinity (Ki) for Muscarinic acetylcholine receptor (heart)was determined by competition radioligand -[3H]- QNB binding assayMore data for this Ligand-Target Pair
Affinity DataKi: 330nMAssay Description:Receptor binding affinity (Ki) for Muscarinic acetylcholine receptor (urinary bladder) was determined by competition radioligand -[3H]- QNB binding a...More data for this Ligand-Target Pair