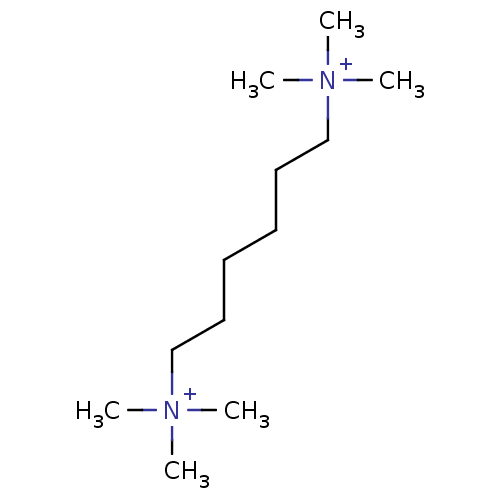
Report error Found 32 Enz. Inhib. hit(s) with all data for entry = 4856
TargetNicotinic acetylcholine receptor(Cape York rat)
Georgetown University
Curated by PDSP Ki Database
Georgetown University
Curated by PDSP Ki Database
TargetNicotinic acetylcholine receptor(Cape York rat)
Georgetown University
Curated by PDSP Ki Database
Georgetown University
Curated by PDSP Ki Database
TargetNicotinic acetylcholine receptor(Cape York rat)
Georgetown University
Curated by PDSP Ki Database
Georgetown University
Curated by PDSP Ki Database
TargetNicotinic acetylcholine receptor(Cape York rat)
Georgetown University
Curated by PDSP Ki Database
Georgetown University
Curated by PDSP Ki Database
TargetNicotinic acetylcholine receptor(Cape York rat)
Georgetown University
Curated by PDSP Ki Database
Georgetown University
Curated by PDSP Ki Database
TargetNicotinic acetylcholine receptor(Cape York rat)
Georgetown University
Curated by PDSP Ki Database
Georgetown University
Curated by PDSP Ki Database
TargetNicotinic acetylcholine receptor(Cape York rat)
Georgetown University
Curated by PDSP Ki Database
Georgetown University
Curated by PDSP Ki Database
TargetNicotinic acetylcholine receptor(Cape York rat)
Georgetown University
Curated by PDSP Ki Database
Georgetown University
Curated by PDSP Ki Database
TargetNicotinic acetylcholine receptor(Cape York rat)
Georgetown University
Curated by PDSP Ki Database
Georgetown University
Curated by PDSP Ki Database
TargetNicotinic acetylcholine receptor(Cape York rat)
Georgetown University
Curated by PDSP Ki Database
Georgetown University
Curated by PDSP Ki Database
TargetNicotinic acetylcholine receptor(Cape York rat)
Georgetown University
Curated by PDSP Ki Database
Georgetown University
Curated by PDSP Ki Database
TargetNicotinic acetylcholine receptor(Cape York rat)
Georgetown University
Curated by PDSP Ki Database
Georgetown University
Curated by PDSP Ki Database
TargetNicotinic acetylcholine receptor(Cape York rat)
Georgetown University
Curated by PDSP Ki Database
Georgetown University
Curated by PDSP Ki Database
TargetNicotinic acetylcholine receptor(Cape York rat)
Georgetown University
Curated by PDSP Ki Database
Georgetown University
Curated by PDSP Ki Database
TargetNicotinic acetylcholine receptor(Cape York rat)
Georgetown University
Curated by PDSP Ki Database
Georgetown University
Curated by PDSP Ki Database
TargetNicotinic acetylcholine receptor(Cape York rat)
Georgetown University
Curated by PDSP Ki Database
Georgetown University
Curated by PDSP Ki Database
TargetNicotinic acetylcholine receptor(Cape York rat)
Georgetown University
Curated by PDSP Ki Database
Georgetown University
Curated by PDSP Ki Database
TargetNicotinic acetylcholine receptor(Cape York rat)
Georgetown University
Curated by PDSP Ki Database
Georgetown University
Curated by PDSP Ki Database
TargetNicotinic acetylcholine receptor(Cape York rat)
Georgetown University
Curated by PDSP Ki Database
Georgetown University
Curated by PDSP Ki Database
TargetNicotinic acetylcholine receptor(Cape York rat)
Georgetown University
Curated by PDSP Ki Database
Georgetown University
Curated by PDSP Ki Database
TargetNicotinic acetylcholine receptor(Cape York rat)
Georgetown University
Curated by PDSP Ki Database
Georgetown University
Curated by PDSP Ki Database
TargetNicotinic acetylcholine receptor(Cape York rat)
Georgetown University
Curated by PDSP Ki Database
Georgetown University
Curated by PDSP Ki Database
TargetNicotinic acetylcholine receptor(Cape York rat)
Georgetown University
Curated by PDSP Ki Database
Georgetown University
Curated by PDSP Ki Database
TargetNicotinic acetylcholine receptor(Cape York rat)
Georgetown University
Curated by PDSP Ki Database
Georgetown University
Curated by PDSP Ki Database
TargetNicotinic acetylcholine receptor(Cape York rat)
Georgetown University
Curated by PDSP Ki Database
Georgetown University
Curated by PDSP Ki Database
TargetNicotinic acetylcholine receptor(Cape York rat)
Georgetown University
Curated by PDSP Ki Database
Georgetown University
Curated by PDSP Ki Database
TargetNicotinic acetylcholine receptor(Cape York rat)
Georgetown University
Curated by PDSP Ki Database
Georgetown University
Curated by PDSP Ki Database
TargetNicotinic acetylcholine receptor(Cape York rat)
Georgetown University
Curated by PDSP Ki Database
Georgetown University
Curated by PDSP Ki Database
TargetNicotinic acetylcholine receptor(Cape York rat)
Georgetown University
Curated by PDSP Ki Database
Georgetown University
Curated by PDSP Ki Database
TargetNicotinic acetylcholine receptor(Cape York rat)
Georgetown University
Curated by PDSP Ki Database
Georgetown University
Curated by PDSP Ki Database
TargetNicotinic acetylcholine receptor(Cape York rat)
Georgetown University
Curated by PDSP Ki Database
Georgetown University
Curated by PDSP Ki Database
TargetNicotinic acetylcholine receptor(Cape York rat)
Georgetown University
Curated by PDSP Ki Database
Georgetown University
Curated by PDSP Ki Database