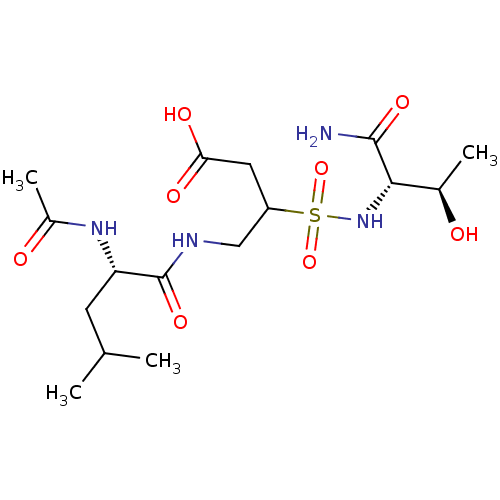
Report error Found 5 Enz. Inhib. hit(s) with all data for entry = 50030118
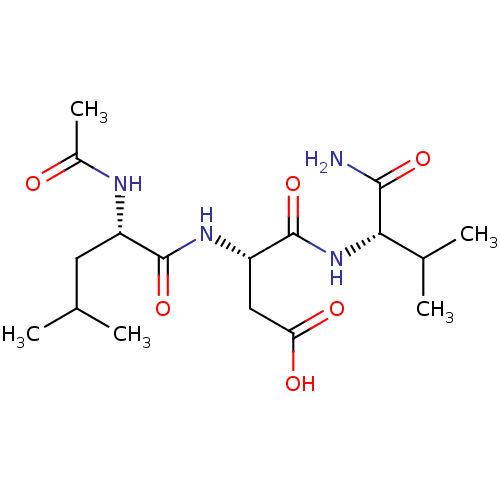
Affinity DataIC50: 1.26E+5nMAssay Description:The concentration of compound required to prevent 50 percent of cells from adhering to MAdCAM-1More data for this Ligand-Target Pair
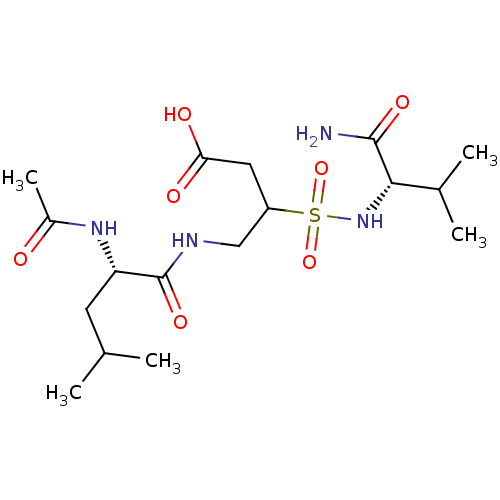
Affinity DataIC50: 2.19E+5nMAssay Description:The concentration of compound required to prevent 50 percent of cells from adhering to MAdCAM-1More data for this Ligand-Target Pair
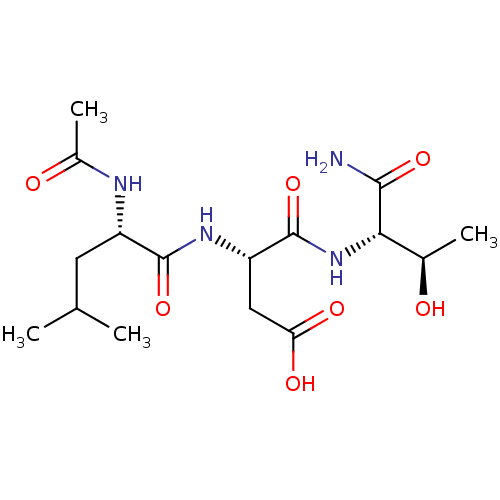
Affinity DataIC50: 2.76E+5nMAssay Description:The concentration of compound required to prevent 50 percent of cells from adhering to MAdCAM-1More data for this Ligand-Target Pair
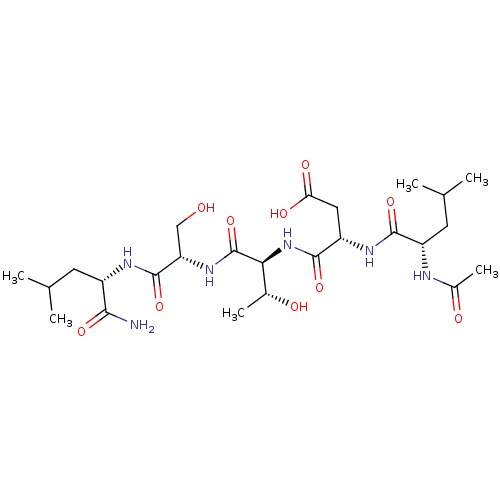
Affinity DataIC50: 2.98E+5nMAssay Description:The concentration of compound required to prevent 50 percent of cells from adhering to MAdCAM-1More data for this Ligand-Target Pair
Affinity DataIC50: 5.00E+5nMAssay Description:The concentration of compound required to prevent 50 percent of cells from adhering to MAdCAM-1More data for this Ligand-Target Pair