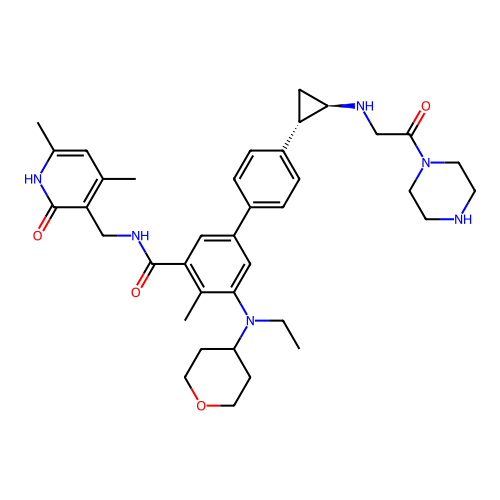
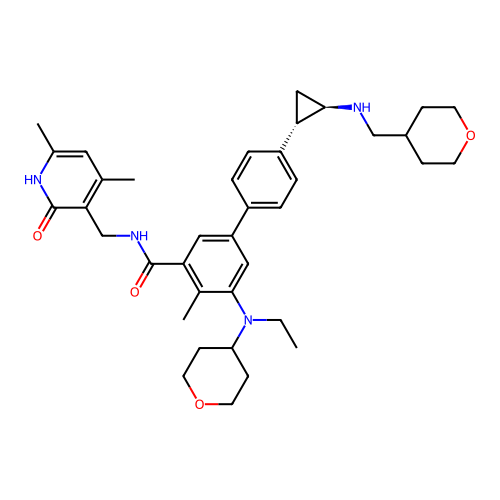
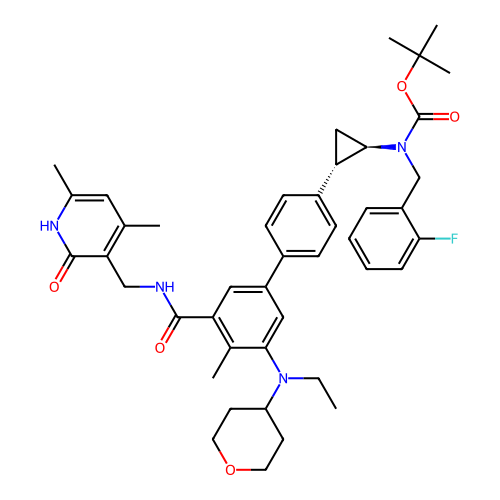
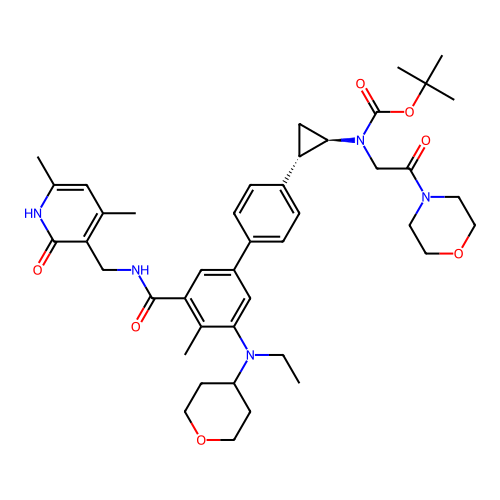
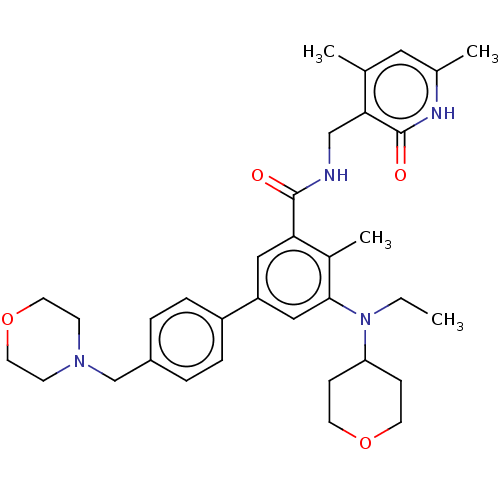
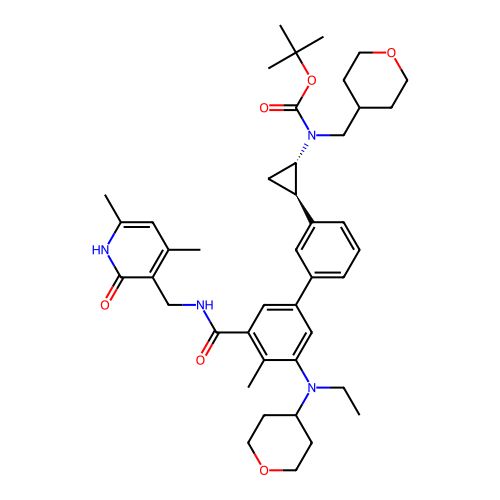
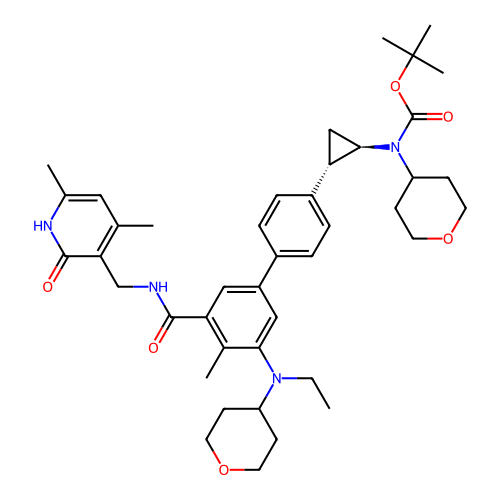
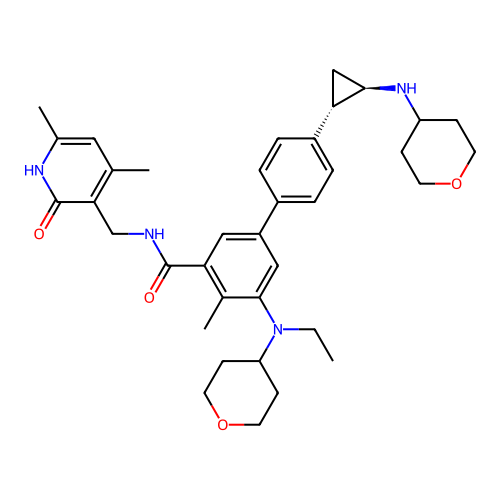
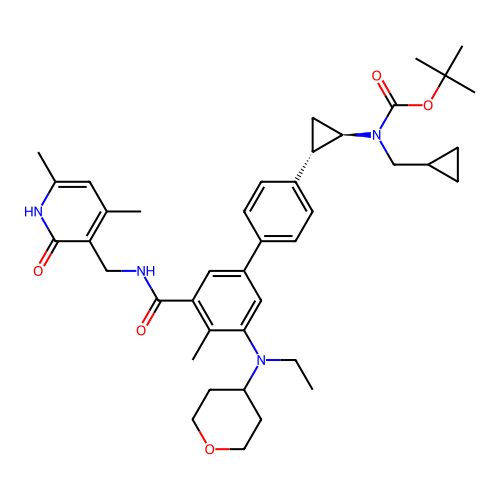
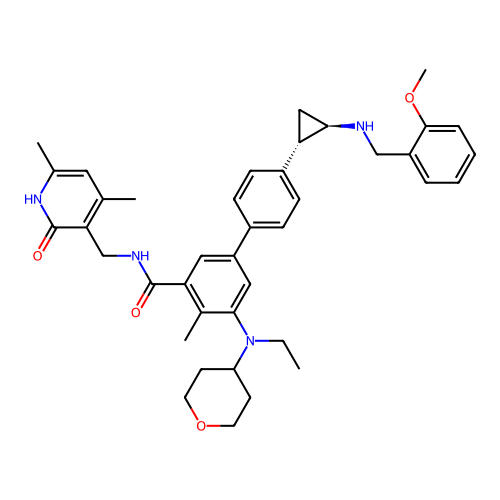
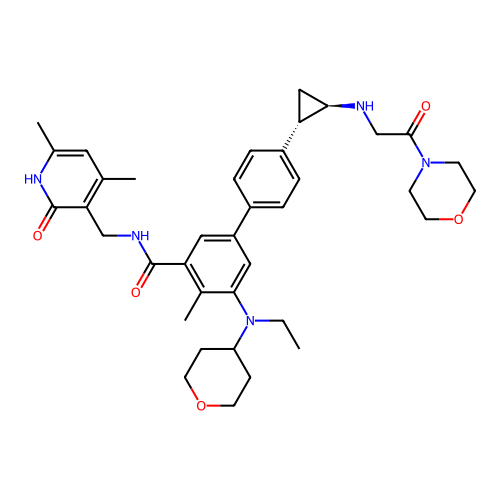
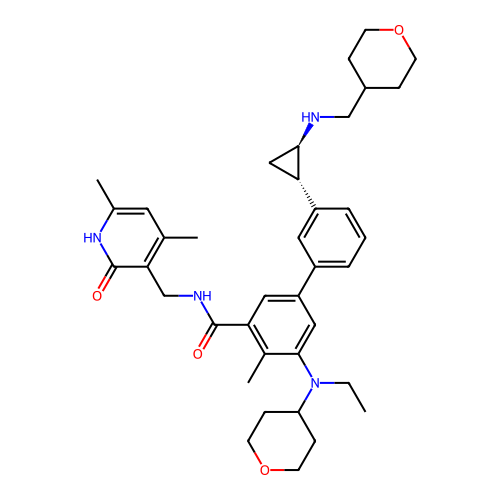
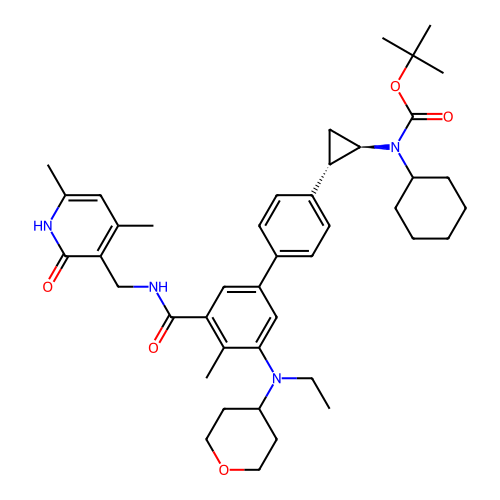
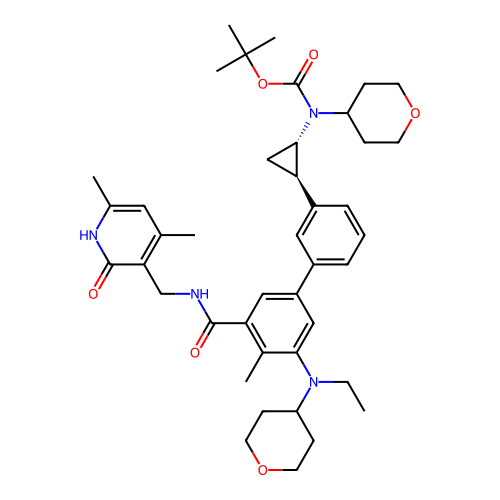
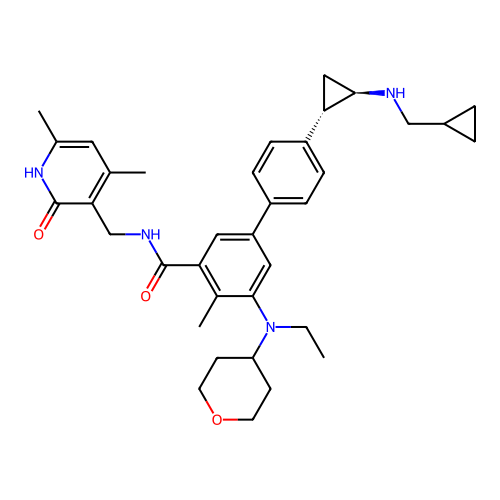
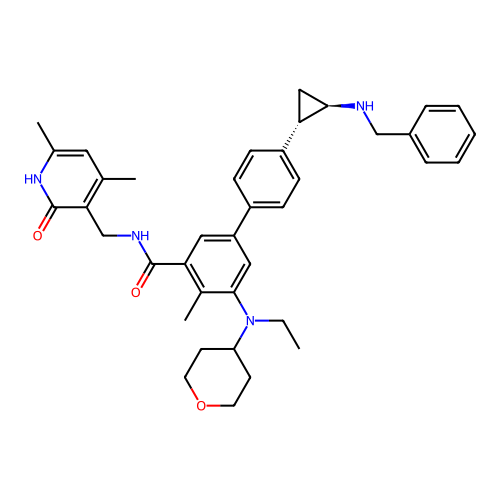
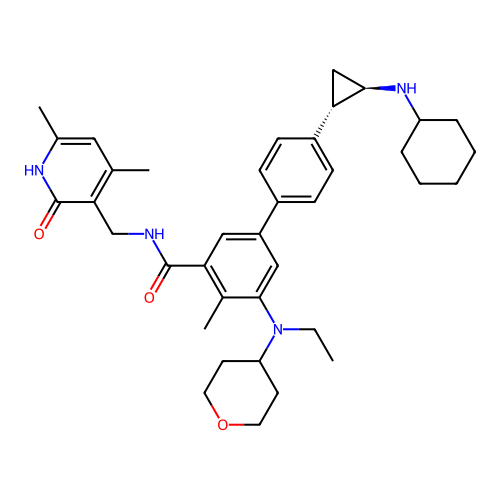
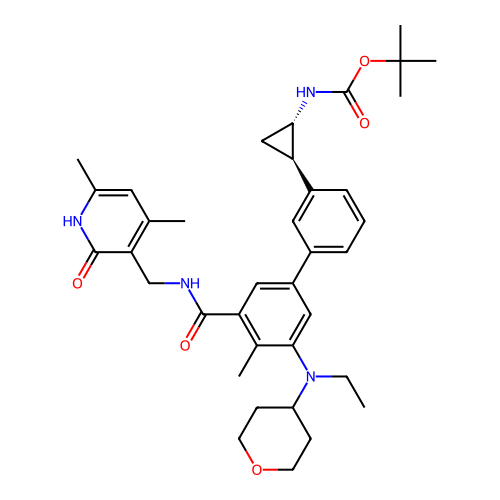
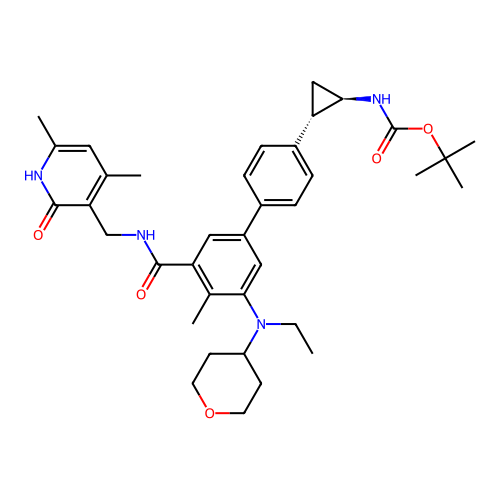
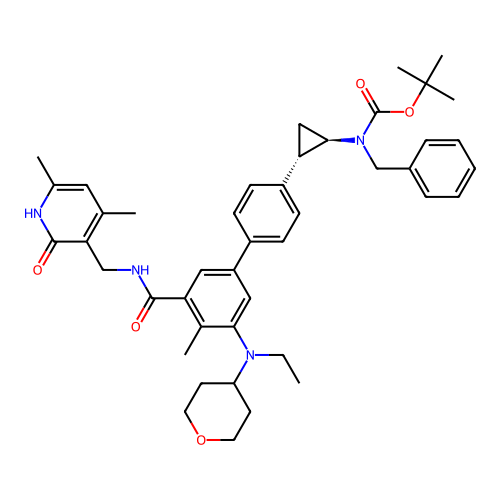
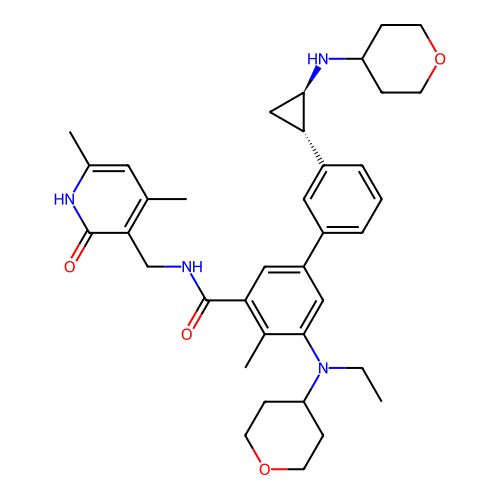
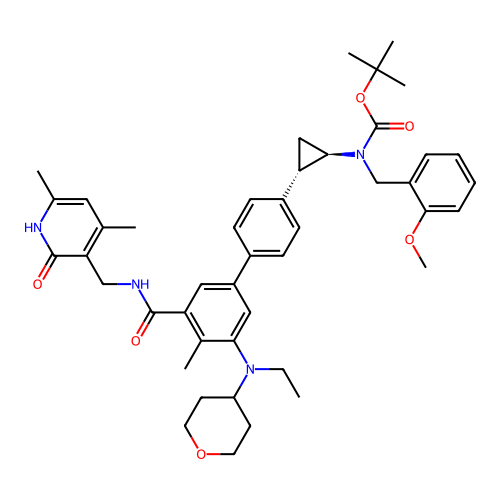
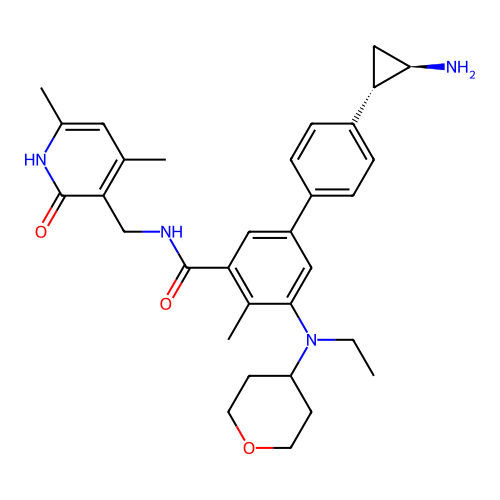
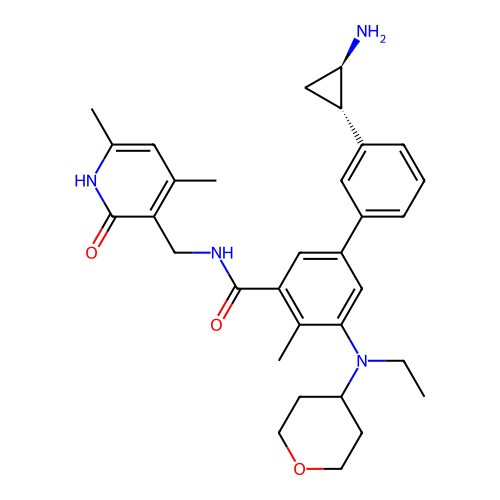
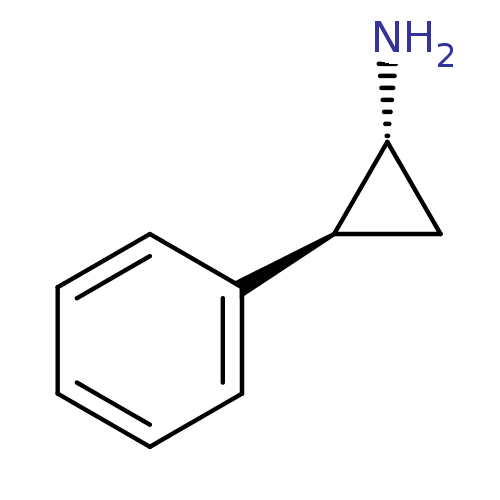
Report error Found 61 Enz. Inhib. hit(s) with all data for entry = 50022098
TargetLysine-specific histone demethylase 1A(Human)
Sun Yat-sen University Cancer Center
Curated by ChEMBL
Sun Yat-sen University Cancer Center
Curated by ChEMBL
Affinity DataIC50: 20nMAssay Description:Inhibition of recombinant human LSD1 using Histone H3 as fluorometric substrate incubated for 30 mins by HRP based plate reader assayMore data for this Ligand-Target Pair
TargetHistone-lysine N-methyltransferase EZH2(Human)
Sun Yat-sen University Cancer Center
Curated by ChEMBL
Sun Yat-sen University Cancer Center
Curated by ChEMBL
Affinity DataIC50: 50nMAssay Description:Inhibition of recombinant EZH2 (unknown origin) using SAM and H3 peptide as substrate incubated for 60 mins by MTase-Glo reagent analysisMore data for this Ligand-Target Pair
TargetLysine-specific histone demethylase 1A(Human)
Sun Yat-sen University Cancer Center
Curated by ChEMBL
Sun Yat-sen University Cancer Center
Curated by ChEMBL
Affinity DataIC50: 90nMAssay Description:Inhibition of recombinant human LSD1 using Histone H3 as fluorometric substrate incubated for 30 mins by HRP based plate reader assayMore data for this Ligand-Target Pair
TargetHistone-lysine N-methyltransferase EZH2(Human)
Sun Yat-sen University Cancer Center
Curated by ChEMBL
Sun Yat-sen University Cancer Center
Curated by ChEMBL
Affinity DataIC50: 90nMAssay Description:Inhibition of recombinant EZH2 (unknown origin) using SAM and H3 peptide as substrate incubated for 60 mins by MTase-Glo reagent analysisMore data for this Ligand-Target Pair
TargetLysine-specific histone demethylase 1A(Human)
Sun Yat-sen University Cancer Center
Curated by ChEMBL
Sun Yat-sen University Cancer Center
Curated by ChEMBL
Affinity DataIC50: 100nMAssay Description:Inhibition of recombinant human LSD1 using Histone H3 as fluorometric substrate incubated for 30 mins by HRP based plate reader assayMore data for this Ligand-Target Pair
TargetLysine-specific histone demethylase 1A(Human)
Sun Yat-sen University Cancer Center
Curated by ChEMBL
Sun Yat-sen University Cancer Center
Curated by ChEMBL
Affinity DataIC50: 120nMAssay Description:Inhibition of recombinant human LSD1 using Histone H3 as fluorometric substrate incubated for 30 mins by HRP based plate reader assayMore data for this Ligand-Target Pair
TargetHistone-lysine N-methyltransferase EZH2(Human)
Sun Yat-sen University Cancer Center
Curated by ChEMBL
Sun Yat-sen University Cancer Center
Curated by ChEMBL
Affinity DataIC50: 140nMAssay Description:Inhibition of recombinant EZH2 (unknown origin) using SAM and H3 peptide as substrate incubated for 60 mins by MTase-Glo reagent analysisMore data for this Ligand-Target Pair
TargetHistone-lysine N-methyltransferase EZH2(Human)
Sun Yat-sen University Cancer Center
Curated by ChEMBL
Sun Yat-sen University Cancer Center
Curated by ChEMBL
Affinity DataIC50: 140nMAssay Description:Inhibition of recombinant EZH2 (unknown origin) using SAM and H3 peptide as substrate incubated for 60 mins by MTase-Glo reagent analysisMore data for this Ligand-Target Pair
TargetHistone-lysine N-methyltransferase EZH2(Human)
Sun Yat-sen University Cancer Center
Curated by ChEMBL
Sun Yat-sen University Cancer Center
Curated by ChEMBL
Affinity DataIC50: 150nMAssay Description:Inhibition of recombinant EZH2 (unknown origin) using SAM and H3 peptide as substrate incubated for 60 mins by MTase-Glo reagent analysisMore data for this Ligand-Target Pair
TargetHistone-lysine N-methyltransferase EZH2(Human)
Sun Yat-sen University Cancer Center
Curated by ChEMBL
Sun Yat-sen University Cancer Center
Curated by ChEMBL
Affinity DataIC50: 170nMAssay Description:Inhibition of recombinant EZH2 (unknown origin) using SAM and H3 peptide as substrate incubated for 60 mins by MTase-Glo reagent analysisMore data for this Ligand-Target Pair
TargetHistone-lysine N-methyltransferase EZH2(Human)
Sun Yat-sen University Cancer Center
Curated by ChEMBL
Sun Yat-sen University Cancer Center
Curated by ChEMBL
Affinity DataIC50: 180nMAssay Description:Inhibition of recombinant EZH2 (unknown origin) using SAM and H3 peptide as substrate incubated for 60 mins by MTase-Glo reagent analysisMore data for this Ligand-Target Pair
TargetHistone-lysine N-methyltransferase EZH2(Human)
Sun Yat-sen University Cancer Center
Curated by ChEMBL
Sun Yat-sen University Cancer Center
Curated by ChEMBL
Affinity DataIC50: 180nMAssay Description:Inhibition of recombinant EZH2 (unknown origin) using SAM and H3 peptide as substrate incubated for 60 mins by MTase-Glo reagent analysisMore data for this Ligand-Target Pair
TargetHistone-lysine N-methyltransferase EZH2(Human)
Sun Yat-sen University Cancer Center
Curated by ChEMBL
Sun Yat-sen University Cancer Center
Curated by ChEMBL
Affinity DataIC50: 190nMAssay Description:Inhibition of recombinant EZH2 (unknown origin) using SAM and H3 peptide as substrate incubated for 60 mins by MTase-Glo reagent analysisMore data for this Ligand-Target Pair
TargetHistone-lysine N-methyltransferase EZH2(Human)
Sun Yat-sen University Cancer Center
Curated by ChEMBL
Sun Yat-sen University Cancer Center
Curated by ChEMBL
Affinity DataIC50: 200nMAssay Description:Inhibition of recombinant EZH2 (unknown origin) using SAM and H3 peptide as substrate incubated for 60 mins by MTase-Glo reagent analysisMore data for this Ligand-Target Pair
TargetHistone-lysine N-methyltransferase EZH2(Human)
Sun Yat-sen University Cancer Center
Curated by ChEMBL
Sun Yat-sen University Cancer Center
Curated by ChEMBL
Affinity DataIC50: 220nMAssay Description:Inhibition of recombinant EZH2 (unknown origin) using SAM and H3 peptide as substrate incubated for 60 mins by MTase-Glo reagent analysisMore data for this Ligand-Target Pair
TargetHistone-lysine N-methyltransferase EZH2(Human)
Sun Yat-sen University Cancer Center
Curated by ChEMBL
Sun Yat-sen University Cancer Center
Curated by ChEMBL
Affinity DataIC50: 220nMAssay Description:Inhibition of recombinant EZH2 (unknown origin) using SAM and H3 peptide as substrate incubated for 60 mins by MTase-Glo reagent analysisMore data for this Ligand-Target Pair
TargetLysine-specific histone demethylase 1A(Human)
Sun Yat-sen University Cancer Center
Curated by ChEMBL
Sun Yat-sen University Cancer Center
Curated by ChEMBL
Affinity DataIC50: 250nMAssay Description:Inhibition of recombinant human LSD1 using Histone H3 as fluorometric substrate incubated for 30 mins by HRP based plate reader assayMore data for this Ligand-Target Pair
TargetLysine-specific histone demethylase 1A(Human)
Sun Yat-sen University Cancer Center
Curated by ChEMBL
Sun Yat-sen University Cancer Center
Curated by ChEMBL
Affinity DataIC50: 250nMAssay Description:Inhibition of recombinant human LSD1 using Histone H3 as fluorometric substrate incubated for 30 mins by HRP based plate reader assayMore data for this Ligand-Target Pair
TargetHistone-lysine N-methyltransferase EZH2(Human)
Sun Yat-sen University Cancer Center
Curated by ChEMBL
Sun Yat-sen University Cancer Center
Curated by ChEMBL
Affinity DataIC50: 260nMAssay Description:Inhibition of recombinant EZH2 (unknown origin) using SAM and H3 peptide as substrate incubated for 60 mins by MTase-Glo reagent analysisMore data for this Ligand-Target Pair
TargetHistone-lysine N-methyltransferase EZH2(Human)
Sun Yat-sen University Cancer Center
Curated by ChEMBL
Sun Yat-sen University Cancer Center
Curated by ChEMBL
Affinity DataIC50: 260nMAssay Description:Inhibition of recombinant EZH2 (unknown origin) using SAM and H3 peptide as substrate incubated for 60 mins by MTase-Glo reagent analysisMore data for this Ligand-Target Pair
TargetHistone-lysine N-methyltransferase EZH2(Human)
Sun Yat-sen University Cancer Center
Curated by ChEMBL
Sun Yat-sen University Cancer Center
Curated by ChEMBL
Affinity DataIC50: 270nMAssay Description:Inhibition of recombinant EZH2 (unknown origin) using SAM and H3 peptide as substrate incubated for 60 mins by MTase-Glo reagent analysisMore data for this Ligand-Target Pair
TargetHistone-lysine N-methyltransferase EZH2(Human)
Sun Yat-sen University Cancer Center
Curated by ChEMBL
Sun Yat-sen University Cancer Center
Curated by ChEMBL
Affinity DataIC50: 280nMAssay Description:Inhibition of recombinant EZH2 (unknown origin) using SAM and H3 peptide as substrate incubated for 60 mins by MTase-Glo reagent analysisMore data for this Ligand-Target Pair
TargetLysine-specific histone demethylase 1A(Human)
Sun Yat-sen University Cancer Center
Curated by ChEMBL
Sun Yat-sen University Cancer Center
Curated by ChEMBL
Affinity DataIC50: 280nMAssay Description:Inhibition of recombinant human LSD1 using Histone H3 as fluorometric substrate incubated for 30 mins by HRP based plate reader assayMore data for this Ligand-Target Pair
TargetHistone-lysine N-methyltransferase EZH2(Human)
Sun Yat-sen University Cancer Center
Curated by ChEMBL
Sun Yat-sen University Cancer Center
Curated by ChEMBL
Affinity DataIC50: 290nMAssay Description:Inhibition of recombinant EZH2 (unknown origin) using SAM and H3 peptide as substrate incubated for 60 mins by MTase-Glo reagent analysisMore data for this Ligand-Target Pair
TargetHistone-lysine N-methyltransferase EZH2(Human)
Sun Yat-sen University Cancer Center
Curated by ChEMBL
Sun Yat-sen University Cancer Center
Curated by ChEMBL
Affinity DataIC50: 330nMAssay Description:Inhibition of recombinant EZH2 (unknown origin) using SAM and H3 peptide as substrate incubated for 60 mins by MTase-Glo reagent analysisMore data for this Ligand-Target Pair
TargetHistone-lysine N-methyltransferase EZH2(Human)
Sun Yat-sen University Cancer Center
Curated by ChEMBL
Sun Yat-sen University Cancer Center
Curated by ChEMBL
Affinity DataIC50: 350nMAssay Description:Inhibition of recombinant EZH2 (unknown origin) using SAM and H3 peptide as substrate incubated for 60 mins by MTase-Glo reagent analysisMore data for this Ligand-Target Pair
TargetLysine-specific histone demethylase 1A(Human)
Sun Yat-sen University Cancer Center
Curated by ChEMBL
Sun Yat-sen University Cancer Center
Curated by ChEMBL
Affinity DataIC50: 400nMAssay Description:Inhibition of recombinant human LSD1 using Histone H3 as fluorometric substrate incubated for 30 mins by HRP based plate reader assayMore data for this Ligand-Target Pair
TargetHistone-lysine N-methyltransferase EZH2(Human)
Sun Yat-sen University Cancer Center
Curated by ChEMBL
Sun Yat-sen University Cancer Center
Curated by ChEMBL
Affinity DataIC50: 410nMAssay Description:Inhibition of recombinant EZH2 (unknown origin) using SAM and H3 peptide as substrate incubated for 60 mins by MTase-Glo reagent analysisMore data for this Ligand-Target Pair
TargetHistone-lysine N-methyltransferase EZH2(Human)
Sun Yat-sen University Cancer Center
Curated by ChEMBL
Sun Yat-sen University Cancer Center
Curated by ChEMBL
Affinity DataIC50: 460nMAssay Description:Inhibition of recombinant EZH2 (unknown origin) using SAM and H3 peptide as substrate incubated for 60 mins by MTase-Glo reagent analysisMore data for this Ligand-Target Pair
TargetHistone-lysine N-methyltransferase EZH2(Human)
Sun Yat-sen University Cancer Center
Curated by ChEMBL
Sun Yat-sen University Cancer Center
Curated by ChEMBL
Affinity DataIC50: 480nMAssay Description:Inhibition of recombinant EZH2 (unknown origin) using SAM and H3 peptide as substrate incubated for 60 mins by MTase-Glo reagent analysisMore data for this Ligand-Target Pair
TargetAmine oxidase [flavin-containing] A(Human)
Sun Yat-sen University Cancer Center
Curated by ChEMBL
Sun Yat-sen University Cancer Center
Curated by ChEMBL
Affinity DataIC50: 510nMAssay Description:Inhibition of MAO-A (unknown origin)More data for this Ligand-Target Pair
TargetHistone-lysine N-methyltransferase EZH2(Human)
Sun Yat-sen University Cancer Center
Curated by ChEMBL
Sun Yat-sen University Cancer Center
Curated by ChEMBL
Affinity DataIC50: 520nMAssay Description:Inhibition of recombinant EZH2 (unknown origin) using SAM and H3 peptide as substrate incubated for 60 mins by MTase-Glo reagent analysisMore data for this Ligand-Target Pair
TargetHistone-lysine N-methyltransferase EZH2(Human)
Sun Yat-sen University Cancer Center
Curated by ChEMBL
Sun Yat-sen University Cancer Center
Curated by ChEMBL
Affinity DataIC50: 540nMAssay Description:Inhibition of recombinant EZH2 (unknown origin) using SAM and H3 peptide as substrate incubated for 60 mins by MTase-Glo reagent analysisMore data for this Ligand-Target Pair
TargetHistone-lysine N-methyltransferase EZH2(Human)
Sun Yat-sen University Cancer Center
Curated by ChEMBL
Sun Yat-sen University Cancer Center
Curated by ChEMBL
Affinity DataIC50: 580nMAssay Description:Inhibition of recombinant EZH2 (unknown origin) using SAM and H3 peptide as substrate incubated for 60 mins by MTase-Glo reagent analysisMore data for this Ligand-Target Pair
TargetHistone-lysine N-methyltransferase EZH2(Human)
Sun Yat-sen University Cancer Center
Curated by ChEMBL
Sun Yat-sen University Cancer Center
Curated by ChEMBL
Affinity DataIC50: 610nMAssay Description:Inhibition of recombinant EZH2 (unknown origin) using SAM and H3 peptide as substrate incubated for 60 mins by MTase-Glo reagent analysisMore data for this Ligand-Target Pair
TargetHistone-lysine N-methyltransferase EZH2(Human)
Sun Yat-sen University Cancer Center
Curated by ChEMBL
Sun Yat-sen University Cancer Center
Curated by ChEMBL
Affinity DataIC50: 620nMAssay Description:Inhibition of recombinant EZH2 (unknown origin) using SAM and H3 peptide as substrate incubated for 60 mins by MTase-Glo reagent analysisMore data for this Ligand-Target Pair
TargetLysine-specific histone demethylase 1A(Human)
Sun Yat-sen University Cancer Center
Curated by ChEMBL
Sun Yat-sen University Cancer Center
Curated by ChEMBL
Affinity DataIC50: 700nMAssay Description:Inhibition of recombinant human LSD1 using Histone H3 as fluorometric substrate incubated for 30 mins by HRP based plate reader assayMore data for this Ligand-Target Pair
TargetLysine-specific histone demethylase 1A(Human)
Sun Yat-sen University Cancer Center
Curated by ChEMBL
Sun Yat-sen University Cancer Center
Curated by ChEMBL
Affinity DataIC50: 870nMAssay Description:Inhibition of recombinant human LSD1 using Histone H3 as fluorometric substrate incubated for 30 mins by HRP based plate reader assayMore data for this Ligand-Target Pair
TargetLysine-specific histone demethylase 1A(Human)
Sun Yat-sen University Cancer Center
Curated by ChEMBL
Sun Yat-sen University Cancer Center
Curated by ChEMBL
Affinity DataIC50: 920nMAssay Description:Inhibition of recombinant human LSD1 using Histone H3 as fluorometric substrate incubated for 30 mins by HRP based plate reader assayMore data for this Ligand-Target Pair
TargetLysine-specific histone demethylase 1A(Human)
Sun Yat-sen University Cancer Center
Curated by ChEMBL
Sun Yat-sen University Cancer Center
Curated by ChEMBL
Affinity DataIC50: 1.04E+3nMAssay Description:Inhibition of recombinant human LSD1 using Histone H3 as fluorometric substrate incubated for 30 mins by HRP based plate reader assayMore data for this Ligand-Target Pair
TargetLysine-specific histone demethylase 1A(Human)
Sun Yat-sen University Cancer Center
Curated by ChEMBL
Sun Yat-sen University Cancer Center
Curated by ChEMBL
Affinity DataIC50: 1.16E+3nMAssay Description:Inhibition of recombinant human LSD1 using Histone H3 as fluorometric substrate incubated for 30 mins by HRP based plate reader assayMore data for this Ligand-Target Pair
TargetHistone-lysine N-methyltransferase EZH1(Human)
Sun Yat-sen University Cancer Center
Curated by ChEMBL
Sun Yat-sen University Cancer Center
Curated by ChEMBL
Affinity DataIC50: 1.17E+3nMAssay Description:Inhibition of EZH1 (unknown origin)More data for this Ligand-Target Pair
TargetAmine oxidase [flavin-containing] A(Human)
Sun Yat-sen University Cancer Center
Curated by ChEMBL
Sun Yat-sen University Cancer Center
Curated by ChEMBL
Affinity DataIC50: 1.54E+3nMAssay Description:Inhibition of MAO-A (unknown origin)More data for this Ligand-Target Pair
TargetAmine oxidase [flavin-containing] A(Human)
Sun Yat-sen University Cancer Center
Curated by ChEMBL
Sun Yat-sen University Cancer Center
Curated by ChEMBL
Affinity DataIC50: 2.05E+3nMAssay Description:Inhibition of MAO-A (unknown origin)More data for this Ligand-Target Pair
TargetHistone-lysine N-methyltransferase EZH1(Human)
Sun Yat-sen University Cancer Center
Curated by ChEMBL
Sun Yat-sen University Cancer Center
Curated by ChEMBL
Affinity DataIC50: 2.16E+3nMAssay Description:Inhibition of EZH1 (unknown origin)More data for this Ligand-Target Pair
TargetLysine-specific histone demethylase 1A(Human)
Sun Yat-sen University Cancer Center
Curated by ChEMBL
Sun Yat-sen University Cancer Center
Curated by ChEMBL
Affinity DataIC50: 3.00E+3nMAssay Description:Inhibition of recombinant human LSD1 using Histone H3 as fluorometric substrate incubated for 30 mins by HRP based plate reader assayMore data for this Ligand-Target Pair
TargetLysine-specific histone demethylase 1A(Human)
Sun Yat-sen University Cancer Center
Curated by ChEMBL
Sun Yat-sen University Cancer Center
Curated by ChEMBL
Affinity DataIC50: 3.00E+3nMAssay Description:Inhibition of recombinant human LSD1 using Histone H3 as fluorometric substrate incubated for 30 mins by HRP based plate reader assayMore data for this Ligand-Target Pair
TargetLysine-specific histone demethylase 1A(Human)
Sun Yat-sen University Cancer Center
Curated by ChEMBL
Sun Yat-sen University Cancer Center
Curated by ChEMBL
Affinity DataIC50: 3.00E+3nMAssay Description:Inhibition of recombinant human LSD1 using Histone H3 as fluorometric substrate incubated for 30 mins by HRP based plate reader assayMore data for this Ligand-Target Pair
TargetLysine-specific histone demethylase 1A(Human)
Sun Yat-sen University Cancer Center
Curated by ChEMBL
Sun Yat-sen University Cancer Center
Curated by ChEMBL
Affinity DataIC50: 3.00E+3nMAssay Description:Inhibition of recombinant human LSD1 using Histone H3 as fluorometric substrate incubated for 30 mins by HRP based plate reader assayMore data for this Ligand-Target Pair
TargetLysine-specific histone demethylase 1A(Human)
Sun Yat-sen University Cancer Center
Curated by ChEMBL
Sun Yat-sen University Cancer Center
Curated by ChEMBL
Affinity DataIC50: 3.00E+3nMAssay Description:Inhibition of recombinant human LSD1 using Histone H3 as fluorometric substrate incubated for 30 mins by HRP based plate reader assayMore data for this Ligand-Target Pair