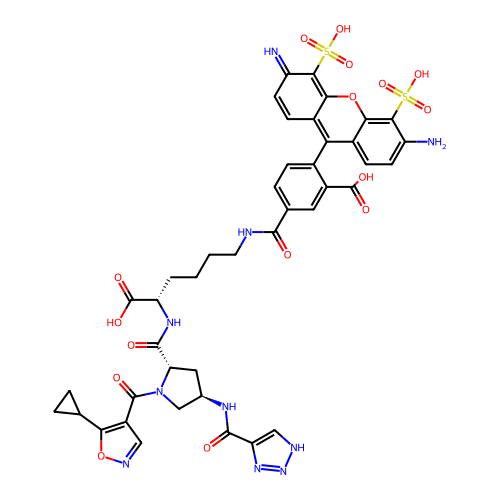
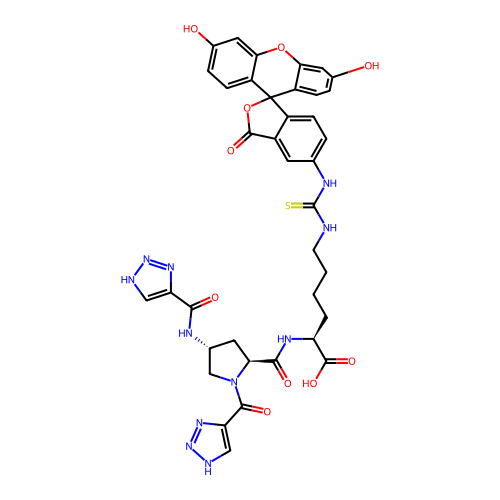
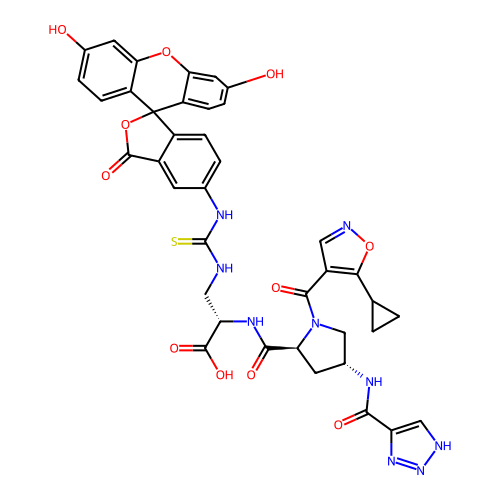
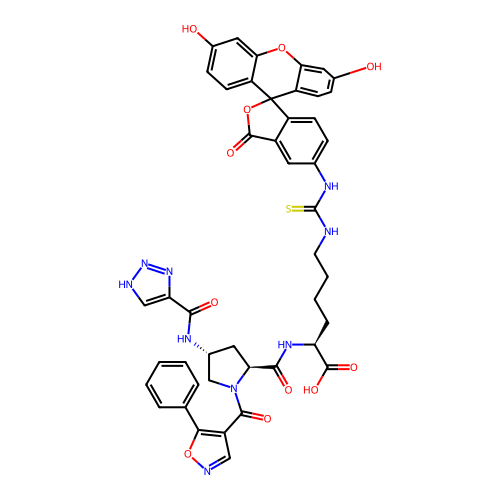
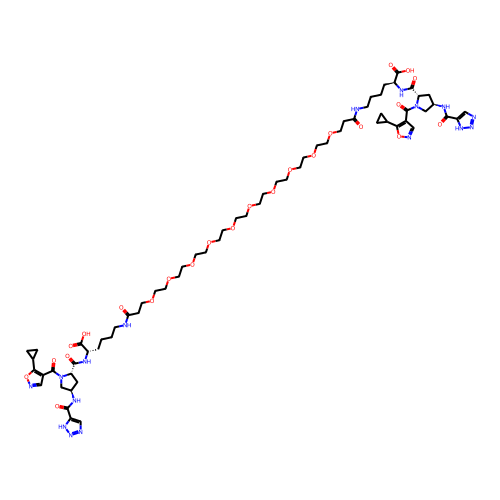
Report error Found 77 Enz. Inhib. hit(s) with all data for entry = 50021073
Affinity DataKd: 1.5nMAssay Description:Displacement of [177Lu]LU-PSMA-617 from human recombinant GCPIII assessed as dissociation constant by fluorescence polarization assayMore data for this Ligand-Target Pair
Affinity DataKd: 1.70nMAssay Description:Binding affinity to human recombinant GCPIII assessed as dissociation constant by fluorescence polarization assayMore data for this Ligand-Target Pair
Affinity DataKd: >2nMAssay Description:Binding affinity to human recombinant PSMA assessed as dissociation constant by fluorescence polarization assayMore data for this Ligand-Target Pair
Affinity DataKd: 4.30nMAssay Description:Binding affinity to human recombinant GCPIII assessed as dissociation constant by fluorescence polarization assayMore data for this Ligand-Target Pair
Affinity DataKd: 4.5nMAssay Description:Binding affinity to human recombinant GCPIII assessed as dissociation constant by fluorescence polarization assayMore data for this Ligand-Target Pair
Affinity DataKd: 4.80nMAssay Description:Binding affinity to human recombinant GCPIII assessed as dissociation constant by fluorescence polarization assayMore data for this Ligand-Target Pair
Affinity DataKd: 6.30nMAssay Description:Binding affinity to human recombinant GCPIII assessed as dissociation constant by fluorescence polarization assayMore data for this Ligand-Target Pair
Affinity DataKd: 6.40nMAssay Description:Binding affinity to human recombinant GCPIII assessed as dissociation constant by fluorescence polarization assayMore data for this Ligand-Target Pair
Affinity DataKd: 7nMAssay Description:Binding affinity to human recombinant GCPIII assessed as dissociation constant by fluorescence polarization assayMore data for this Ligand-Target Pair
Affinity DataKd: 8nMAssay Description:Displacement of [177Lu]LU-PSMA-617 from human recombinant GCPIII assessed as dissociation constant by fluorescence polarization assayMore data for this Ligand-Target Pair
Affinity DataKd: 12nMAssay Description:Displacement of [177Lu]LU-PSMA-617 from human recombinant GCPIII assessed as dissociation constant by fluorescence polarization assayMore data for this Ligand-Target Pair
Affinity DataKd: 12nMAssay Description:Binding affinity to human recombinant GCPIII assessed as dissociation constant by fluorescence polarization assayMore data for this Ligand-Target Pair
Affinity DataKd: 12nMAssay Description:Displacement of [177Lu]LU-PSMA-617 from human recombinant GCPIII assessed as dissociation constant by fluorescence polarization assayMore data for this Ligand-Target Pair
Affinity DataKd: 14nMAssay Description:Binding affinity to human recombinant GCPIII assessed as dissociation constant by fluorescence polarization assayMore data for this Ligand-Target Pair
Affinity DataKd: 15nMAssay Description:Binding affinity to human recombinant GCPIII assessed as dissociation constant by fluorescence polarization assayMore data for this Ligand-Target Pair
Affinity DataKd: 23nMAssay Description:Binding affinity to human recombinant PSMA assessed as dissociation constant by fluorescence polarization assayMore data for this Ligand-Target Pair
Affinity DataKd: 23nMAssay Description:Binding affinity to human recombinant GCPIII assessed as dissociation constant by fluorescence polarization assayMore data for this Ligand-Target Pair
Affinity DataKd: 24nMAssay Description:Binding affinity to human recombinant GCPIII assessed as dissociation constant by fluorescence polarization assayMore data for this Ligand-Target Pair
Affinity DataKd: 30nMAssay Description:Binding affinity to human recombinant GCPIII assessed as dissociation constant by fluorescence polarization assayMore data for this Ligand-Target Pair
Affinity DataKd: 41nMAssay Description:Displacement of [177Lu]LU-PSMA-617 from human recombinant GCPIII assessed as dissociation constant by fluorescence polarization assayMore data for this Ligand-Target Pair
Affinity DataKd: 48nMAssay Description:Binding affinity to human recombinant PSMA assessed as dissociation constant by fluorescence polarization assayMore data for this Ligand-Target Pair
Affinity DataKd: 53nMAssay Description:Displacement of [177Lu]LU-PSMA-617 from human recombinant GCPIII assessed as dissociation constant by fluorescence polarization assayMore data for this Ligand-Target Pair
Affinity DataKd: 67nMAssay Description:Displacement of [177Lu]LU-PSMA-617 from human recombinant GCPIII assessed as dissociation constant by fluorescence polarization assayMore data for this Ligand-Target Pair
Affinity DataKd: 87nMAssay Description:Binding affinity to human recombinant GCPIII assessed as dissociation constant by fluorescence polarization assayMore data for this Ligand-Target Pair
Affinity DataKd: 106nMAssay Description:Binding affinity to human recombinant GCPIII assessed as dissociation constant by fluorescence polarization assayMore data for this Ligand-Target Pair
Affinity DataKd: 169nMAssay Description:Binding affinity to human recombinant GCPIII assessed as dissociation constant by fluorescence polarization assayMore data for this Ligand-Target Pair
Affinity DataKd: 178nMAssay Description:Binding affinity to human recombinant PSMA assessed as dissociation constant by fluorescence polarization assayMore data for this Ligand-Target Pair
Affinity DataKd: 231nMAssay Description:Displacement of [177Lu]LU-PSMA-617 from human recombinant PSMA assessed as dissociation constant by fluorescence polarization assayMore data for this Ligand-Target Pair
Affinity DataKd: >300nMAssay Description:Binding affinity to human recombinant GCPIII assessed as dissociation constant by fluorescence polarization assayMore data for this Ligand-Target Pair
Affinity DataKd: >300nMAssay Description:Binding affinity to human recombinant GCPIII assessed as dissociation constant by fluorescence polarization assayMore data for this Ligand-Target Pair
Affinity DataKd: 300nMAssay Description:Displacement of [177Lu]LU-PSMA-617 from human recombinant GCPIII assessed as dissociation constant by fluorescence polarization assayMore data for this Ligand-Target Pair
Affinity DataKd: >300nMAssay Description:Binding affinity to human recombinant GCPIII assessed as dissociation constant by fluorescence polarization assayMore data for this Ligand-Target Pair
Affinity DataKd: 310nMAssay Description:Binding affinity to human recombinant PSMA assessed as dissociation constant by fluorescence polarization assayMore data for this Ligand-Target Pair
Affinity DataKd: 332nMAssay Description:Binding affinity to human recombinant PSMA assessed as dissociation constant by fluorescence polarization assayMore data for this Ligand-Target Pair
Affinity DataKd: 338nMAssay Description:Displacement of [177Lu]LU-PSMA-617 from human recombinant GCPIII assessed as dissociation constant by fluorescence polarization assayMore data for this Ligand-Target Pair
Affinity DataKd: >500nMAssay Description:Binding affinity to human recombinant GCPIII assessed as dissociation constant by fluorescence polarization assayMore data for this Ligand-Target Pair
Affinity DataKd: >500nMAssay Description:Binding affinity to human recombinant GCPIII assessed as dissociation constant by fluorescence polarization assayMore data for this Ligand-Target Pair
Affinity DataKd: >500nMAssay Description:Binding affinity to human recombinant GCPIII assessed as dissociation constant by fluorescence polarization assayMore data for this Ligand-Target Pair
Affinity DataKd: >500nMAssay Description:Binding affinity to human recombinant GCPIII assessed as dissociation constant by fluorescence polarization assayMore data for this Ligand-Target Pair
Affinity DataKd: 505nMAssay Description:Displacement of [177Lu]LU-PSMA-617 from human recombinant GCPIII assessed as dissociation constant by fluorescence polarization assayMore data for this Ligand-Target Pair
Affinity DataKd: >700nMAssay Description:Binding affinity to mouse FAP assessed as dissociation constant by fluorescence polarization assayMore data for this Ligand-Target Pair
Affinity DataKd: >1.00E+3nMAssay Description:Binding affinity to human recombinant PSMA assessed as dissociation constant by fluorescence polarization assayMore data for this Ligand-Target Pair
Affinity DataKd: >1.00E+3nMAssay Description:Binding affinity to human recombinant PSMA assessed as dissociation constant by fluorescence polarization assayMore data for this Ligand-Target Pair
Affinity DataKd: >1.00E+3nMAssay Description:Binding affinity to human recombinant PSMA assessed as dissociation constant by fluorescence polarization assayMore data for this Ligand-Target Pair
Affinity DataKd: >1.00E+3nMAssay Description:Binding affinity to human recombinant PSMA assessed as dissociation constant by fluorescence polarization assayMore data for this Ligand-Target Pair
Affinity DataKd: >1.00E+3nMAssay Description:Binding affinity to human recombinant PSMA assessed as dissociation constant by fluorescence polarization assayMore data for this Ligand-Target Pair
Affinity DataKd: >1.00E+3nMAssay Description:Binding affinity to human recombinant PSMA assessed as dissociation constant by fluorescence polarization assayMore data for this Ligand-Target Pair
Affinity DataKd: >1.00E+3nMAssay Description:Binding affinity to human recombinant PSMA assessed as dissociation constant by fluorescence polarization assayMore data for this Ligand-Target Pair
Affinity DataKd: >1.00E+3nMAssay Description:Binding affinity to human recombinant PSMA assessed as dissociation constant by fluorescence polarization assayMore data for this Ligand-Target Pair
Affinity DataKd: >1.00E+3nMAssay Description:Binding affinity to human recombinant PSMA assessed as dissociation constant by fluorescence polarization assayMore data for this Ligand-Target Pair