Report error Found 46 Enz. Inhib. hit(s) with all data for entry = 50021193
TargetSignal peptidase I(Escherichia coli (strain K12))
Eberhard Karls University Tubingen
Curated by ChEMBL
Eberhard Karls University Tubingen
Curated by ChEMBL
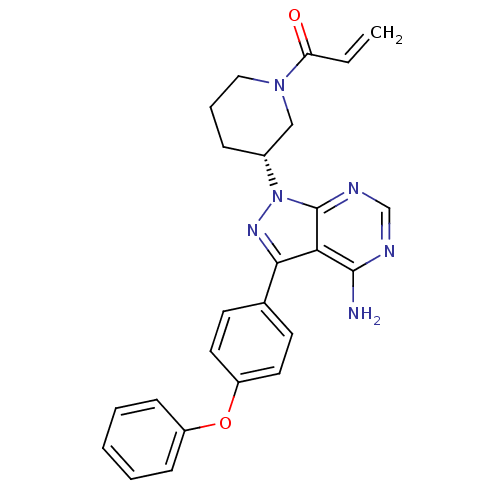
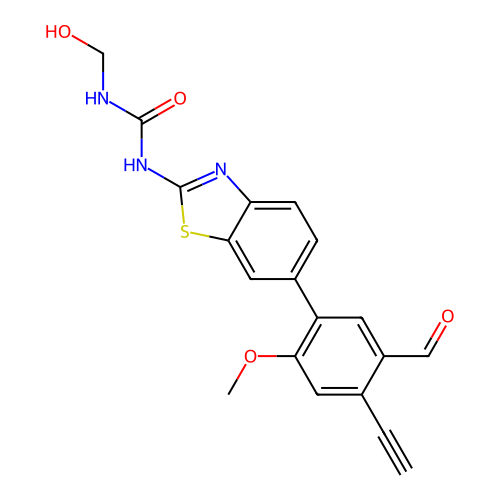
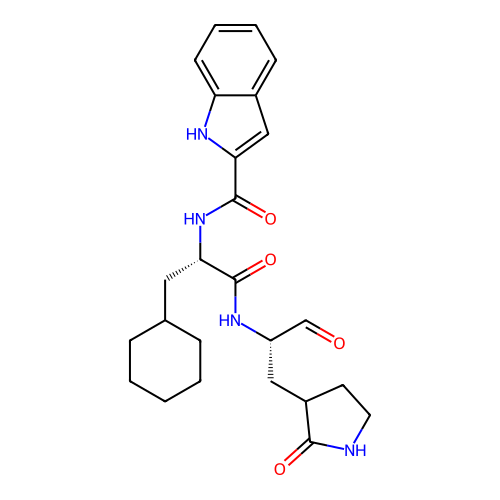
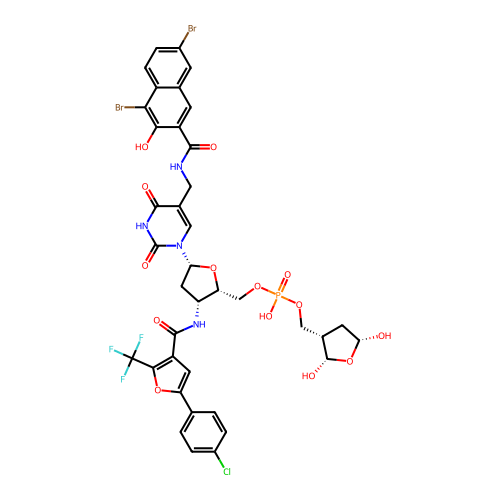
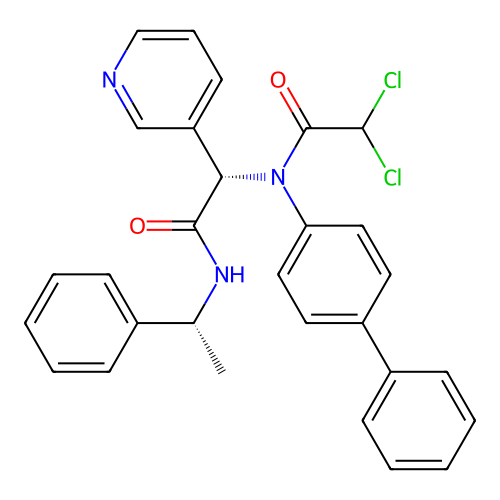
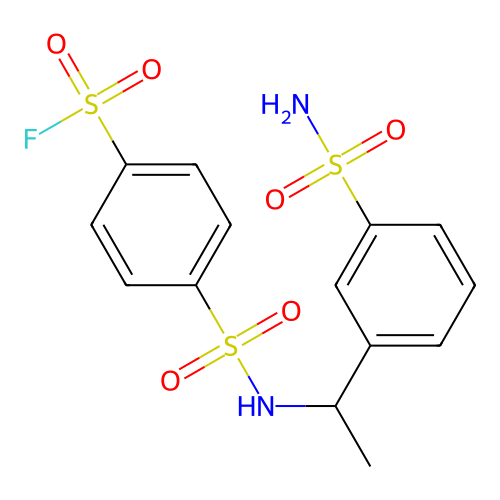
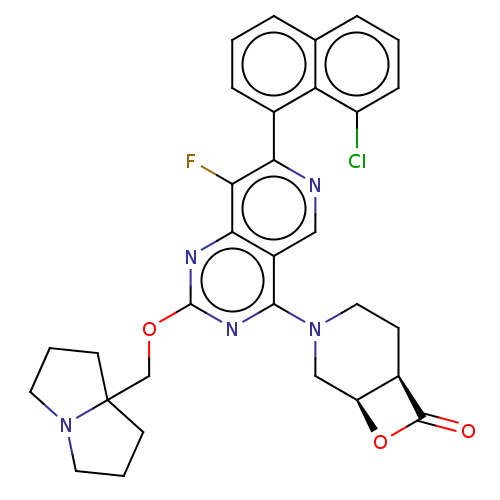
Affinity DataKi: 0.440nMAssay Description:Binding affinity to Escherichia coli type I signal peptidase LepB assessed as inhibition constantMore data for this Ligand-Target Pair
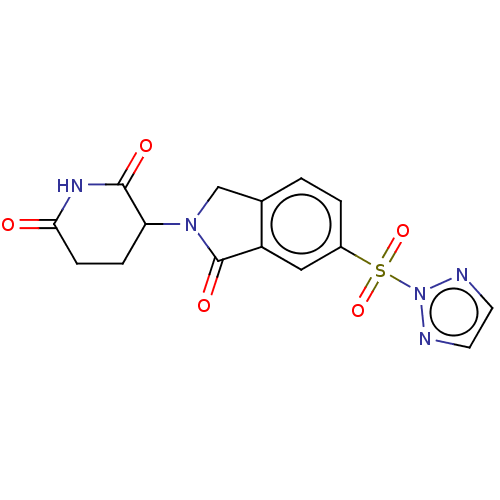
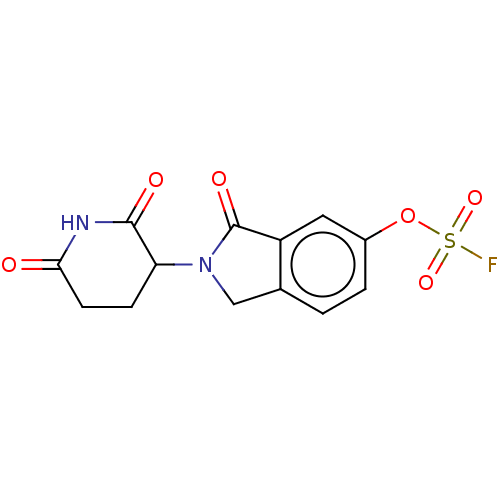
Affinity DataIC50: 0.880nMAssay Description:Inhibition of CRBN (unknown origin)More data for this Ligand-Target Pair
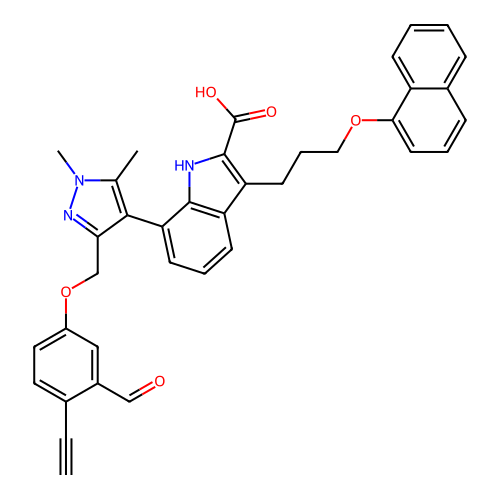
Affinity DataIC50: 2.80nMAssay Description:Inhibition of wild type N-terminal His tagged ABL kinase (unknown origin) using fluorescence-labelled substrate by off-chip mobility shift assayMore data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit delta isoform(Human)
Eberhard Karls University Tubingen
Curated by ChEMBL
Eberhard Karls University Tubingen
Curated by ChEMBL
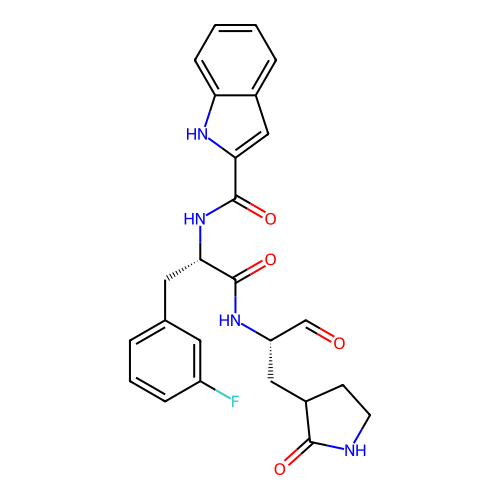
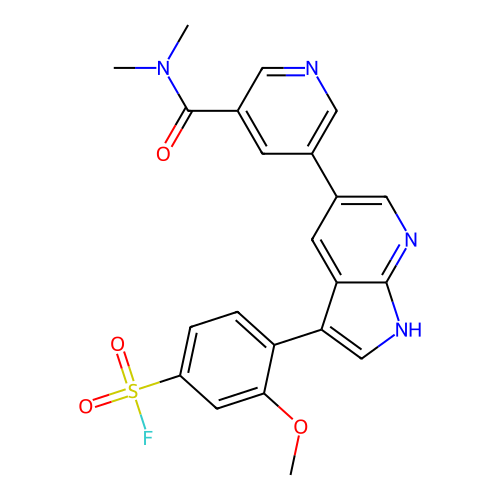
Affinity DataKi: 3.60nMAssay Description:Inhibition of PI3Kdelta (unknown origin) assessed as inhibition constantMore data for this Ligand-Target Pair
Affinity DataKi: 4.20nMAssay Description:Binding affinity to human ALDH1A2 assessed as inhibition constantMore data for this Ligand-Target Pair
Affinity DataKi: 4.40nMAssay Description:Binding affinity to BTK (unknown origin) assessed as inhibition constantMore data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit delta isoform(Human)
Eberhard Karls University Tubingen
Curated by ChEMBL
Eberhard Karls University Tubingen
Curated by ChEMBL
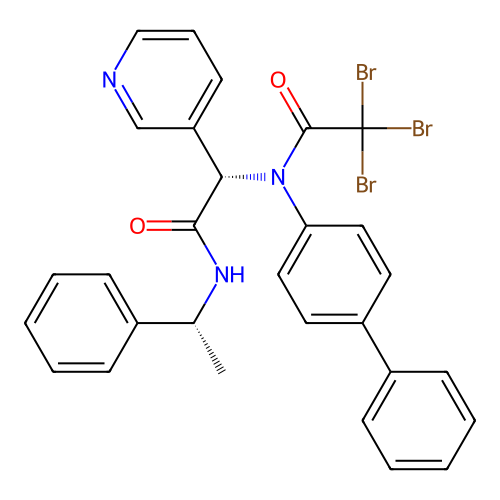
Affinity DataKi: 5.70nMAssay Description:Inhibition of PI3Kdelta (unknown origin) assessed as inhibition constantMore data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit delta isoform(Human)
Eberhard Karls University Tubingen
Curated by ChEMBL
Eberhard Karls University Tubingen
Curated by ChEMBL
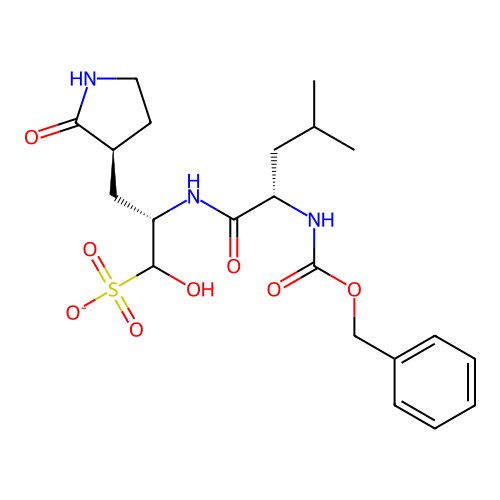
Affinity DataKi: 6nMAssay Description:Inhibition of PI3Kdelta (unknown origin) assessed as inhibition constantMore data for this Ligand-Target Pair
TargetPeptidyl-prolyl cis-trans isomerase E(Homo sapiens)
Eberhard Karls University Tubingen
Curated by ChEMBL
Eberhard Karls University Tubingen
Curated by ChEMBL
Affinity DataIC50: 13nMAssay Description:Inhibition of CypE (unknown origin) expressed in Escherichia coli BL21(DE3) by FLIPR based Chymotrypsin coupled prolyl-isomerase assayMore data for this Ligand-Target Pair
Affinity DataKi: 13nMAssay Description:Inhibition of recombinant ABL kinase (unknown origin) assessed as inhibition constantMore data for this Ligand-Target Pair
Affinity DataIC50: 21nMAssay Description:Inhibition of CRBN (unknown origin)More data for this Ligand-Target Pair
TargetInduced myeloid leukemia cell differentiation protein Mcl-1(Human)
Eberhard Karls University Tubingen
Curated by ChEMBL
Eberhard Karls University Tubingen
Curated by ChEMBL
Affinity DataIC50: 27nMAssay Description:Irreversible inhibition of MCL-1 (unknown origin) measured after 8 hrsMore data for this Ligand-Target Pair
Affinity DataIC50: 40nMAssay Description:Inhibition of SARS-CoV-2 Main protease (Mpro) by FRET assayMore data for this Ligand-Target Pair
Affinity DataIC50: 53nMAssay Description:Inhibition of SARS-CoV-2 Main protease (Mpro) by FRET assayMore data for this Ligand-Target Pair
TargetPeptidyl-prolyl cis-trans isomerase E(Homo sapiens)
Eberhard Karls University Tubingen
Curated by ChEMBL
Eberhard Karls University Tubingen
Curated by ChEMBL
Affinity DataKi: 72nMAssay Description:Inhibition of CypE (unknown origin) assessed as apparent inhibition constant incubated for 24 hrs by Competition anisotropy binding assayMore data for this Ligand-Target Pair
Affinity DataIC50: 73nMAssay Description:Inhibition of wild type N-terminal His tagged ABL kinase (unknown origin) using fluorescence-labelled substrate by off-chip mobility shift assayMore data for this Ligand-Target Pair
Affinity DataIC50: 80nMAssay Description:Inhibition of SARS-CoV-2 cysteine protease main proteaseMore data for this Ligand-Target Pair
TargetMAP kinase-activated protein kinase 2(Human)
Eberhard Karls University Tubingen
Curated by ChEMBL
Eberhard Karls University Tubingen
Curated by ChEMBL
Affinity DataIC50: 156nMAssay Description:Inhibition of human MK2 recombinant assessed as phosphorylation of Sox peptide by continuous fluorescence read assayMore data for this Ligand-Target Pair
Ligand InfoSimilars
Affinity DataIC50: 190nMAssay Description:Inhibition of SARS-CoV-2 cysteine protease main proteaseMore data for this Ligand-Target Pair
Affinity DataIC50: 204nMAssay Description:Inhibition of DNA polymerase beta (unknown origin) measured after 30 minsMore data for this Ligand-Target Pair
Affinity DataKi: 221nMAssay Description:Binding affinity to BTK (unknown origin) assessed as inhibition constantMore data for this Ligand-Target Pair
Affinity DataIC50: 256nMAssay Description:Inhibition of CRBN (unknown origin)More data for this Ligand-Target Pair
Affinity DataEC50: 290nMAssay Description:Allosteric activation at human liver pyruvate kinaseMore data for this Ligand-Target Pair
Affinity DataIC50: 430nMAssay Description:Inhibition of SARS-CoV-2 cysteine protease main proteaseMore data for this Ligand-Target Pair
Affinity DataIC50: 800nMAssay Description:Inhibition of SARS-CoV-2 papain-like protease (PLpro) using fluorogenic peptide substrate by fluorescent assayMore data for this Ligand-Target Pair
Affinity DataKi: 1.34E+3nMAssay Description:Binding affinity to SARS-CoV-2 cysteine protease-Mpro assessed as inhibition constantMore data for this Ligand-Target Pair
Affinity DataKi: 1.80E+3nMAssay Description:Irreversible inhibition of DNA polymerase beta (unknown origin) assessed as inhibition constant by Kitz-Wilson plot analysisMore data for this Ligand-Target Pair
Affinity DataKi: 3.20E+3nMAssay Description:Inhibition of recombinant ABL kinase (unknown origin) assessed as inhibition constantMore data for this Ligand-Target Pair
Affinity DataIC50: 4.90E+3nMAssay Description:Inhibition of SARS-CoV-2 papain-like protease (PLpro) using fluorogenic peptide substrate by fluorescent assayMore data for this Ligand-Target Pair
Affinity DataKi: <5.00E+3nMAssay Description:Rate of protein modification at carbonic anhydrase II (unknown origin) fragment assessed as inhibition constantMore data for this Ligand-Target Pair
Affinity DataKi: <5.00E+3nMAssay Description:Rate of protein modification at carbonic anhydrase II (unknown origin) fragment assessed as inhibition constantMore data for this Ligand-Target Pair
Affinity DataKi: <5.00E+3nMAssay Description:Rate of protein modification at carbonic anhydrase II (unknown origin) fragment assessed as inhibition constantMore data for this Ligand-Target Pair
Affinity DataKi: <5.00E+3nMAssay Description:Rate of protein modification at carbonic anhydrase II (unknown origin) fragment assessed as inhibition constantMore data for this Ligand-Target Pair
Affinity DataKi: <5.00E+3nMAssay Description:Rate of protein modification at carbonic anhydrase II (unknown origin) fragment assessed as inhibition constantMore data for this Ligand-Target Pair
Affinity DataKi: <5.00E+3nMAssay Description:Rate of protein modification at carbonic anhydrase II (unknown origin) fragment assessed as inhibition constantMore data for this Ligand-Target Pair
Affinity DataKi: <5.00E+3nMAssay Description:Rate of protein modification at carbonic anhydrase II (unknown origin) fragment assessed as inhibition constantMore data for this Ligand-Target Pair
Affinity DataKi: <5.00E+3nMAssay Description:Rate of protein modification at carbonic anhydrase II (unknown origin) fragment assessed as inhibition constantMore data for this Ligand-Target Pair
Affinity DataKi: <5.00E+3nMAssay Description:Rate of protein modification at carbonic anhydrase II (unknown origin) fragment assessed as inhibition constantMore data for this Ligand-Target Pair
Affinity DataIC50: 5.20E+3nMAssay Description:Inhibition of SARS-CoV-2 papain-like protease (PLpro) using fluorogenic peptide substrate by fluorescent assayMore data for this Ligand-Target Pair
Affinity DataKi: 1.20E+4nMAssay Description:Rate of protein modification at carbonic anhydrase II (unknown origin) fragment assessed as inhibition constantMore data for this Ligand-Target Pair
Affinity DataKi: 2.20E+4nMAssay Description:Rate of protein modification at carbonic anhydrase II (unknown origin) fragment assessed as inhibition constantMore data for this Ligand-Target Pair
Affinity DataKi: 2.60E+4nMAssay Description:Binding affinity to human KRAS G12S mutant assessed as inhibition constantMore data for this Ligand-Target Pair
Ligand InfoSimilars
Affinity DataKi: 3.30E+4nMAssay Description:Rate of protein modification at carbonic anhydrase II (unknown origin) fragment assessed as inhibition constantMore data for this Ligand-Target Pair
Affinity DataKi: 5.30E+4nMAssay Description:Rate of protein modification at carbonic anhydrase II (unknown origin) fragment assessed as inhibition constantMore data for this Ligand-Target Pair
Affinity DataKi: 5.36E+4nMAssay Description:Binding affinity to SARS-CoV-2 cysteine protease-Mpro assessed as inhibition constantMore data for this Ligand-Target Pair
Affinity DataKi: 6.47E+4nMAssay Description:Binding affinity to SARS-CoV-2 cysteine protease-Mpro assessed as inhibition constantMore data for this Ligand-Target Pair
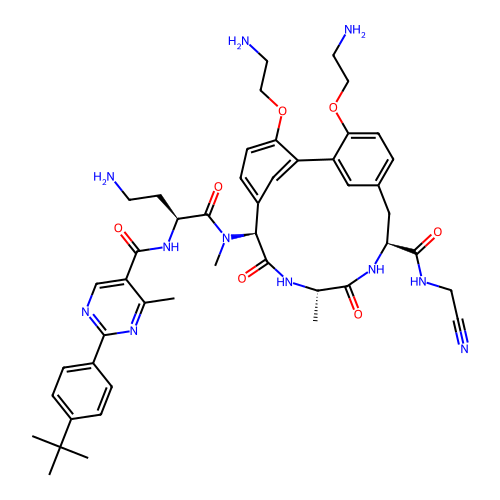
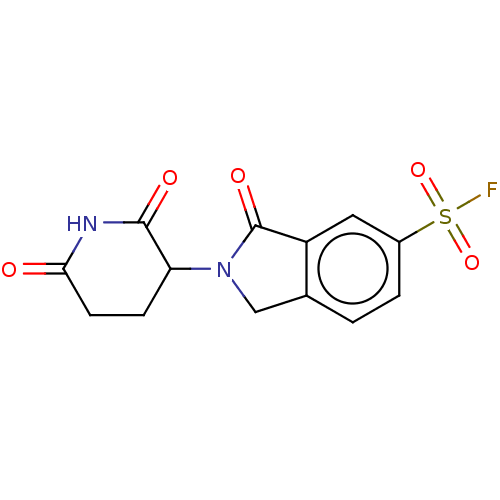
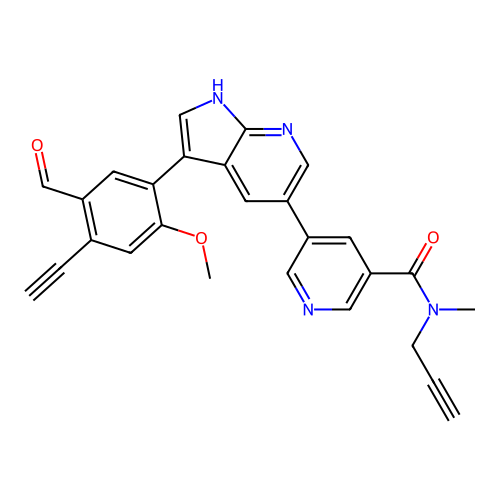
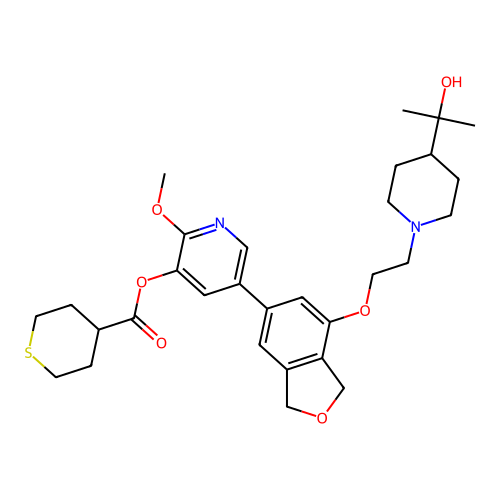
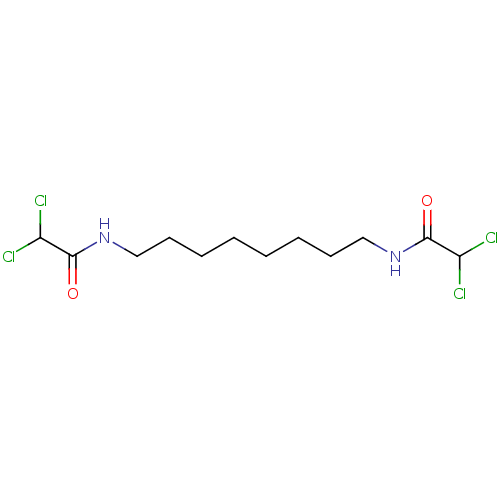
 3D Structure (crystal)
3D Structure (crystal)