Report error Found 15 Enz. Inhib. hit(s) with all data for entry = 50020049
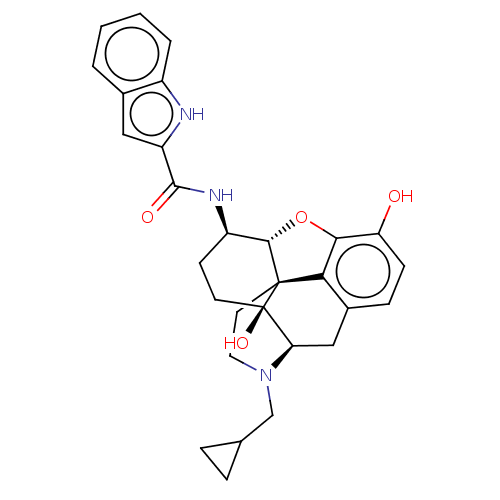
Affinity DataIC50: 0.210nMAssay Description:Inhibition of mu opioid receptor (unknown origin)More data for this Ligand-Target Pair
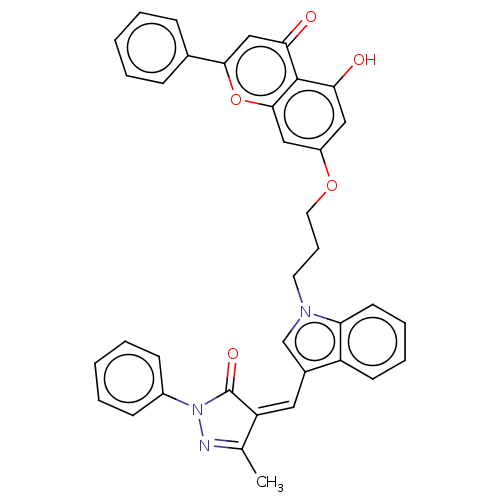
Ligand InfoSimilars
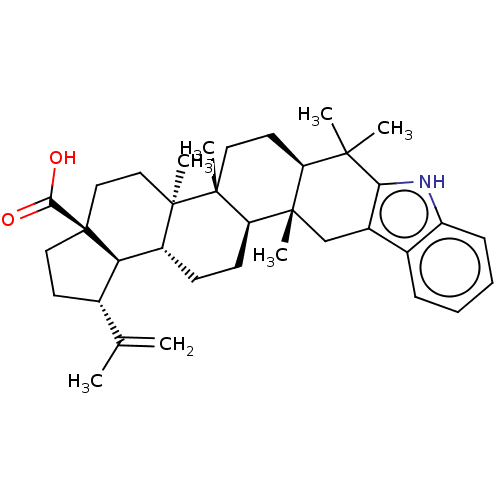
Affinity DataIC50: 40nMAssay Description:Inhibition of alpha glucosidase (unknown origin) measured after 60 minsMore data for this Ligand-Target Pair
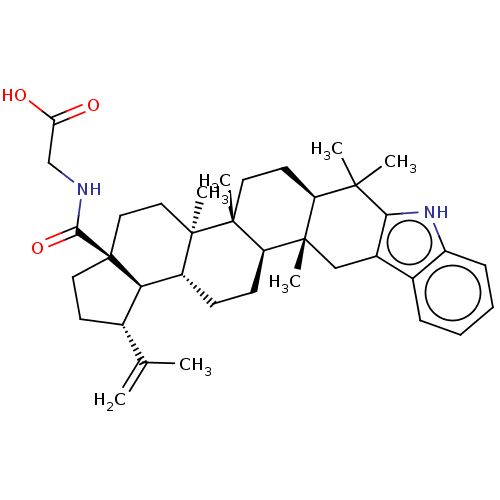
Ligand InfoSimilars
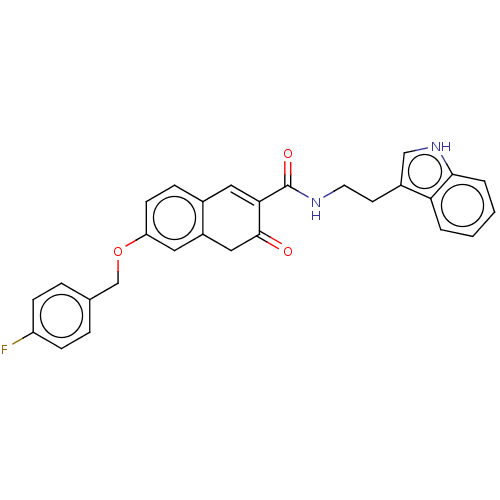
Affinity DataIC50: 160nMAssay Description:Inhibition of Acetylcholinesterase (unknown origin)More data for this Ligand-Target Pair
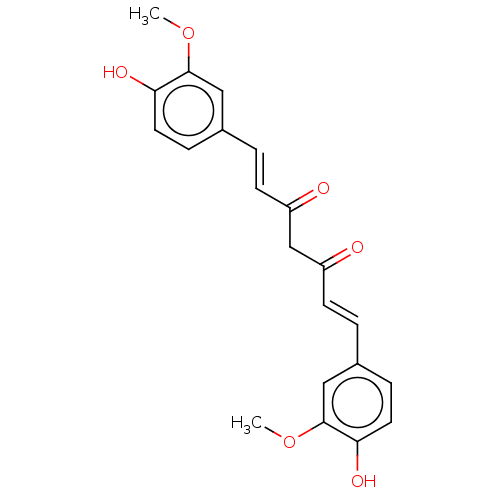
Ligand InfoSimilars
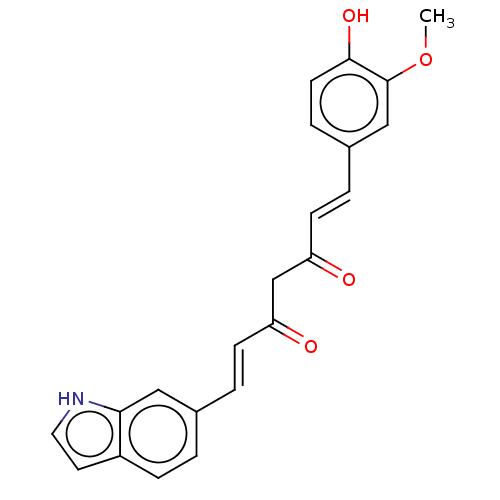
Affinity DataIC50: 420nMAssay Description:Inhibition of amyloid-beta (unknown origin)More data for this Ligand-Target Pair
Affinity DataIC50: 440nMAssay Description:Inhibition of tau phosphorylation (unknown origin)More data for this Ligand-Target Pair
Affinity DataIC50: 700nMAssay Description:Inhibition of COX-2 (unknown origin)More data for this Ligand-Target Pair
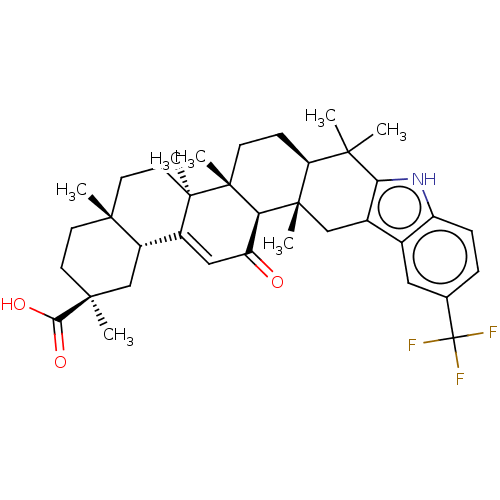
Ligand InfoSimilars
Affinity DataIC50: 1.80E+3nMAssay Description:Inhibition of alpha glucosidase (unknown origin) measured after 60 minsMore data for this Ligand-Target Pair
Ligand InfoSimilars
Affinity DataIC50: 3.00E+3nMAssay Description:Inhibition of tau phosphorylation (unknown origin)More data for this Ligand-Target Pair
Affinity DataIC50: 4.30E+3nMAssay Description:Inhibition of Protein tyrosine phosphatase 1B (unknown origin)More data for this Ligand-Target Pair
Ligand InfoSimilars
Affinity DataIC50: 5.40E+3nMAssay Description:Inhibition of amyloid-beta (unknown origin)More data for this Ligand-Target Pair
Affinity DataIC50: 8.50E+4nMAssay Description:Inhibition of Alpha-glucosidase (unknown origin) spectrophotometric analysisMore data for this Ligand-Target Pair
Ligand InfoSimilars
Affinity DataIC50: 1.15E+5nMAssay Description:Inhibition of Alpha-glucosidase (unknown origin) using p-nitrophenyl glucopyranoside as substrate by UV based microplate reader analysisMore data for this Ligand-Target Pair
Ligand InfoSimilars
Affinity DataIC50: 1.89E+5nMAssay Description:Inhibition of alpha glucosidase (unknown origin) measured after 60 minsMore data for this Ligand-Target Pair
Affinity DataIC50: 4.20E+5nMAssay Description:Inhibition of Alpha-glucosidase (unknown origin) using p-nitrophenyl glucopyranoside as substrate by UV based microplate reader analysisMore data for this Ligand-Target Pair
Affinity DataIC50: 6.04E+5nMAssay Description:Inhibition of Alpha-glucosidase (unknown origin) using p-nitrophenyl glucopyranoside as substrate by UV based microplate reader analysisMore data for this Ligand-Target Pair
Ligand InfoSimilars