Report error Found 22 Enz. Inhib. hit(s) with all data for entry = 50020872
TargetHistone-lysine N-methyltransferase EHMT2(Human)
National Institute of Technology Durgapur
Curated by ChEMBL
National Institute of Technology Durgapur
Curated by ChEMBL
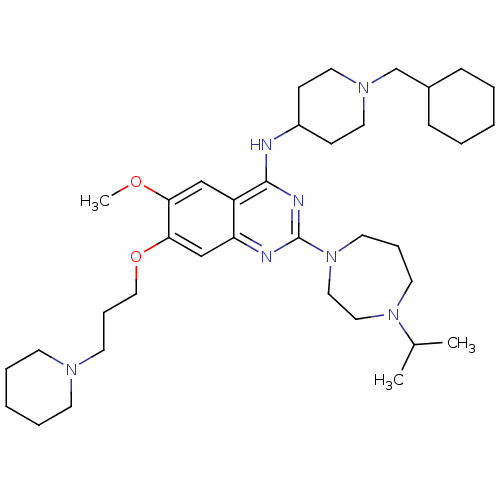
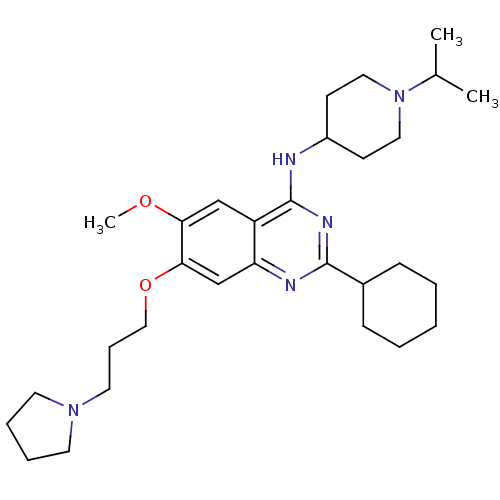
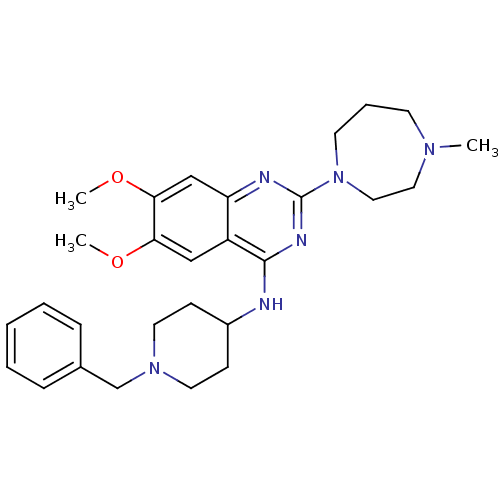
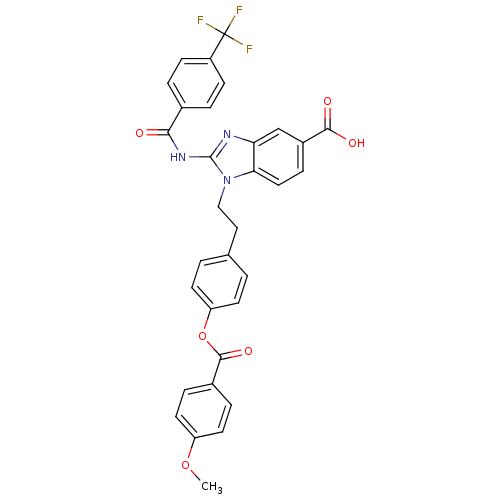
Affinity DataIC50: 1.80nMAssay Description:Inhibition of N-terminal GST-tagged human recombinant G9a (785 to 1210 residues) expressed in baculovirus-infected Sf9 cells using biotinylated histo...More data for this Ligand-Target Pair
TargetHistone-lysine N-methyltransferase EHMT2(Human)
National Institute of Technology Durgapur
Curated by ChEMBL
National Institute of Technology Durgapur
Curated by ChEMBL
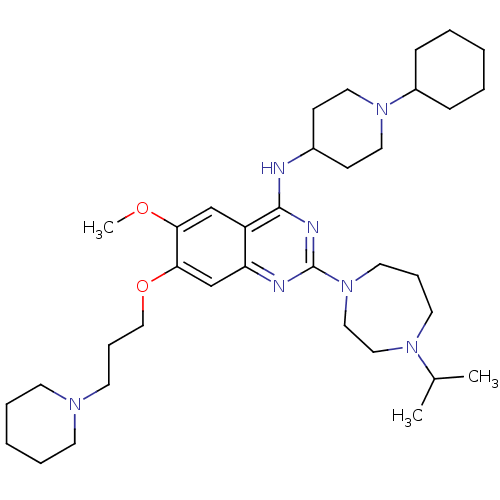
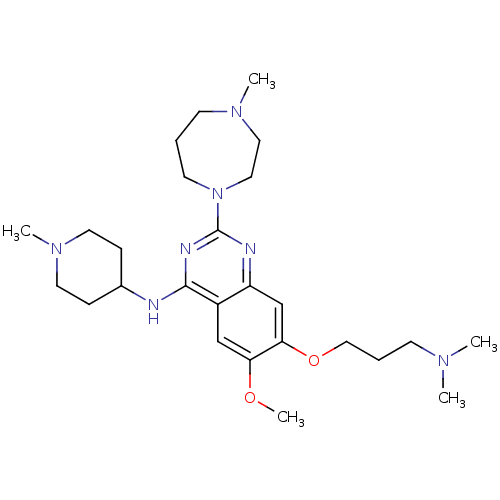
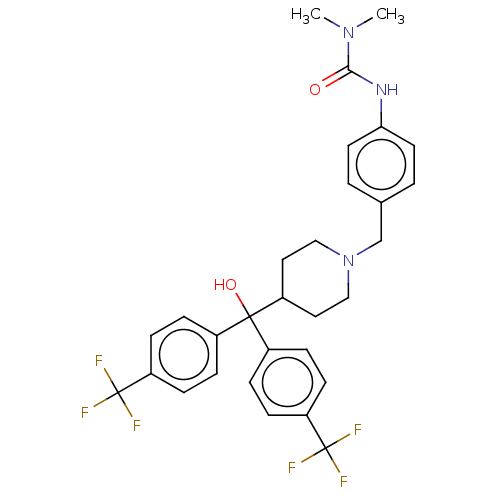
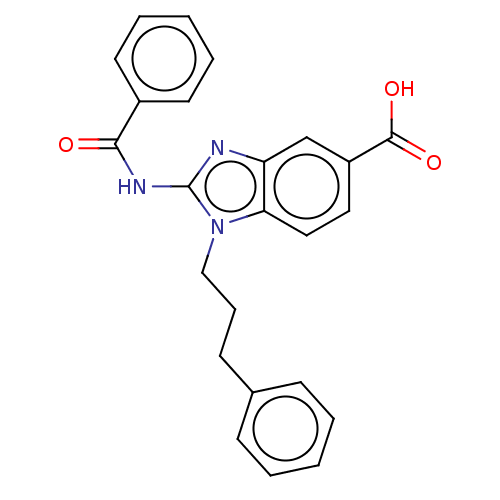
Affinity DataIC50: 2.10nMAssay Description:Inhibition of N-terminal GST-tagged human recombinant G9a (785 to 1210 residues) expressed in baculovirus-infected Sf9 cells using biotinylated histo...More data for this Ligand-Target Pair
TargetHistone-lysine N-methyltransferase EHMT2(Human)
National Institute of Technology Durgapur
Curated by ChEMBL
National Institute of Technology Durgapur
Curated by ChEMBL
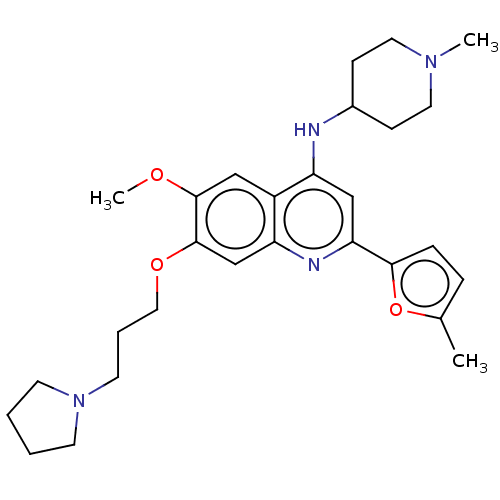
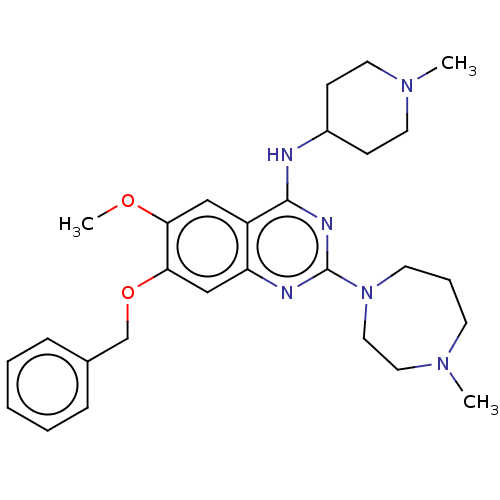
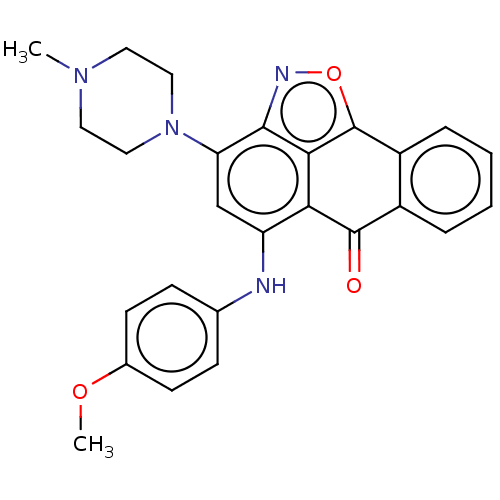
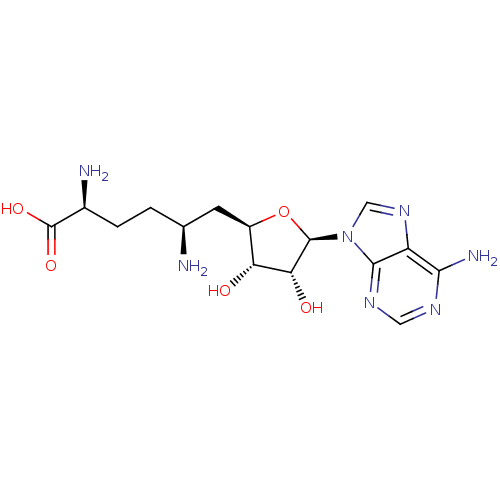
Affinity DataIC50: 2.5nMAssay Description:Inhibition of N-terminal GST-tagged human recombinant G9a (785 to 1210 residues) expressed in baculovirus-infected Sf9 cells using biotinylated histo...More data for this Ligand-Target Pair
TargetHistone-lysine N-methyltransferase EHMT2(Human)
National Institute of Technology Durgapur
Curated by ChEMBL
National Institute of Technology Durgapur
Curated by ChEMBL
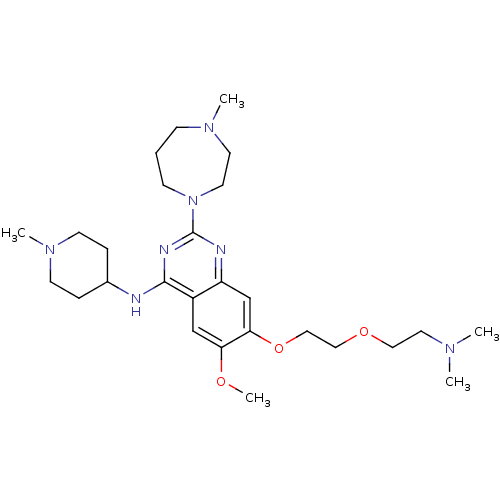
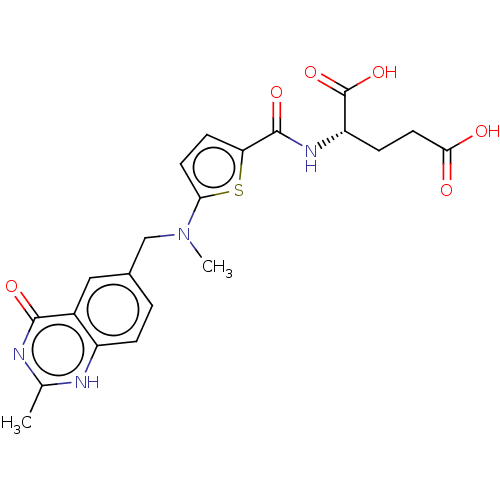
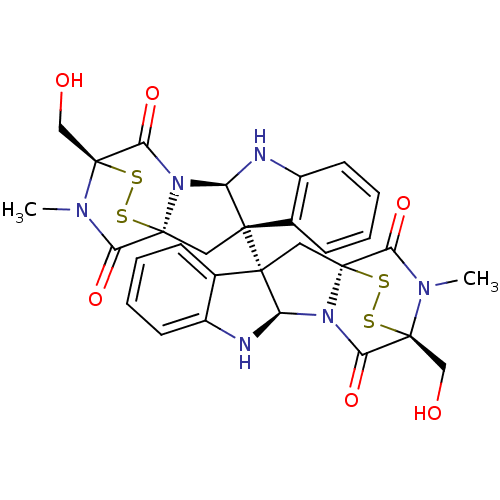
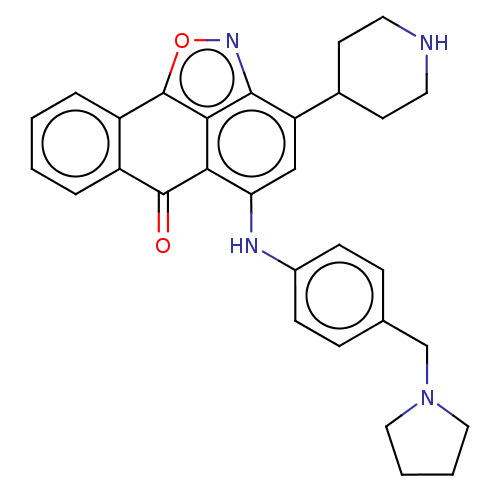
Affinity DataIC50: 2.5nMAssay Description:Inhibition of G9a (unknown origin)More data for this Ligand-Target Pair
TargetHistone-lysine N-methyltransferase EHMT2(Human)
National Institute of Technology Durgapur
Curated by ChEMBL
National Institute of Technology Durgapur
Curated by ChEMBL
Affinity DataIC50: 3.30nMAssay Description:Inhibition of N-terminal GST-tagged human recombinant G9a (785 to 1210 residues) expressed in baculovirus-infected Sf9 cells using biotinylated histo...More data for this Ligand-Target Pair
TargetHistone-lysine N-methyltransferase EHMT2(Human)
National Institute of Technology Durgapur
Curated by ChEMBL
National Institute of Technology Durgapur
Curated by ChEMBL
Affinity DataIC50: 4nMAssay Description:Inhibition of G9a (unknown origin)More data for this Ligand-Target Pair
TargetHistone-lysine N-methyltransferase EHMT2(Human)
National Institute of Technology Durgapur
Curated by ChEMBL
National Institute of Technology Durgapur
Curated by ChEMBL
Affinity DataIC50: 6nMAssay Description:Inhibition of G9a (unknown origin)More data for this Ligand-Target Pair
TargetHistone-lysine N-methyltransferase EHMT2(Human)
National Institute of Technology Durgapur
Curated by ChEMBL
National Institute of Technology Durgapur
Curated by ChEMBL
Affinity DataIC50: 8nMAssay Description:Inhibition of G9a (unknown origin)More data for this Ligand-Target Pair
TargetHistone-lysine N-methyltransferase EHMT2(Human)
National Institute of Technology Durgapur
Curated by ChEMBL
National Institute of Technology Durgapur
Curated by ChEMBL
Affinity DataIC50: 9nMAssay Description:Inhibition of G9a (unknown origin)More data for this Ligand-Target Pair
TargetHistone-lysine N-methyltransferase EHMT2(Human)
National Institute of Technology Durgapur
Curated by ChEMBL
National Institute of Technology Durgapur
Curated by ChEMBL
Affinity DataIC50: 9.40nMAssay Description:Inhibition of G9a (unknown origin)More data for this Ligand-Target Pair
TargetHistone-lysine N-methyltransferase EHMT2(Human)
National Institute of Technology Durgapur
Curated by ChEMBL
National Institute of Technology Durgapur
Curated by ChEMBL
Affinity DataIC50: 12nMAssay Description:Inhibition of human recombinant G9a (785 to 1210 residues) using Histone H3 and SAM as substrate incubated for 2 hrs by chemiluminescence analysisMore data for this Ligand-Target Pair
TargetHistone-lysine N-methyltransferase EHMT2(Human)
National Institute of Technology Durgapur
Curated by ChEMBL
National Institute of Technology Durgapur
Curated by ChEMBL
Affinity DataIC50: 15nMAssay Description:Inhibition of G9a (unknown origin)More data for this Ligand-Target Pair
TargetHistone-lysine N-methyltransferase EHMT2(Human)
National Institute of Technology Durgapur
Curated by ChEMBL
National Institute of Technology Durgapur
Curated by ChEMBL
Affinity DataIC50: 43nMAssay Description:Inhibition of G9a (unknown origin)More data for this Ligand-Target Pair
TargetHistone-lysine N-methyltransferase EHMT2(Human)
National Institute of Technology Durgapur
Curated by ChEMBL
National Institute of Technology Durgapur
Curated by ChEMBL
Affinity DataIC50: 120nMAssay Description:Inhibition of human recombinant G9a (785 to 1210 residues) using Histone H3 and SAM as substrate incubated for 2 hrs by chemiluminescence analysisMore data for this Ligand-Target Pair
TargetHistone-lysine N-methyltransferase EHMT2(Human)
National Institute of Technology Durgapur
Curated by ChEMBL
National Institute of Technology Durgapur
Curated by ChEMBL
Affinity DataIC50: 1.70E+3nMAssay Description:Inhibition of N-terminal GST-tagged human recombinant G9a (785 to 1210 residues) expressed in baculovirus-infected Sf9 cells using biotinylated histo...More data for this Ligand-Target Pair
TargetHistone-lysine N-methyltransferase EHMT2(Human)
National Institute of Technology Durgapur
Curated by ChEMBL
National Institute of Technology Durgapur
Curated by ChEMBL
Affinity DataIC50: 1.70E+3nMAssay Description:Inhibition of G9a (unknown origin)More data for this Ligand-Target Pair
TargetHistone-lysine N-methyltransferase EHMT2(Human)
National Institute of Technology Durgapur
Curated by ChEMBL
National Institute of Technology Durgapur
Curated by ChEMBL
Affinity DataIC50: 2.18E+3nMAssay Description:Inhibition of G9a (unknown origin)More data for this Ligand-Target Pair
TargetHistone-lysine N-methyltransferase EHMT2(Human)
National Institute of Technology Durgapur
Curated by ChEMBL
National Institute of Technology Durgapur
Curated by ChEMBL
Affinity DataIC50: 2.50E+3nMAssay Description:Inhibition of N-terminal GST-tagged human recombinant G9a (785 to 1210 residues) expressed in baculovirus-infected Sf9 cells using biotinylated histo...More data for this Ligand-Target Pair
TargetHistone-lysine N-methyltransferase EHMT2(Human)
National Institute of Technology Durgapur
Curated by ChEMBL
National Institute of Technology Durgapur
Curated by ChEMBL
Affinity DataIC50: 4.70E+3nMAssay Description:Inhibition of G9a (unknown origin)More data for this Ligand-Target Pair
TargetHistone-lysine N-methyltransferase EHMT2(Human)
National Institute of Technology Durgapur
Curated by ChEMBL
National Institute of Technology Durgapur
Curated by ChEMBL
Affinity DataIC50: 6.30E+3nMAssay Description:Inhibition of G9a (unknown origin)More data for this Ligand-Target Pair
TargetHistone-lysine N-methyltransferase EHMT2(Human)
National Institute of Technology Durgapur
Curated by ChEMBL
National Institute of Technology Durgapur
Curated by ChEMBL
Affinity DataIC50: 3.01E+4nMAssay Description:Inhibition of G9a (unknown origin)More data for this Ligand-Target Pair
TargetHistone-lysine N-methyltransferase EHMT2(Human)
National Institute of Technology Durgapur
Curated by ChEMBL
National Institute of Technology Durgapur
Curated by ChEMBL
Affinity DataIC50: 3.92E+4nMAssay Description:Inhibition of G9a (unknown origin)More data for this Ligand-Target Pair
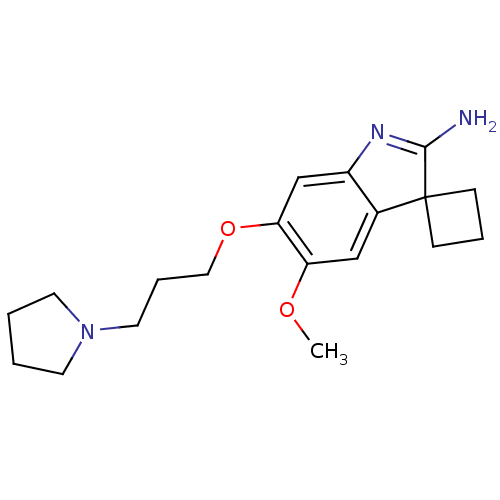
 3D Structure (crystal)
3D Structure (crystal)