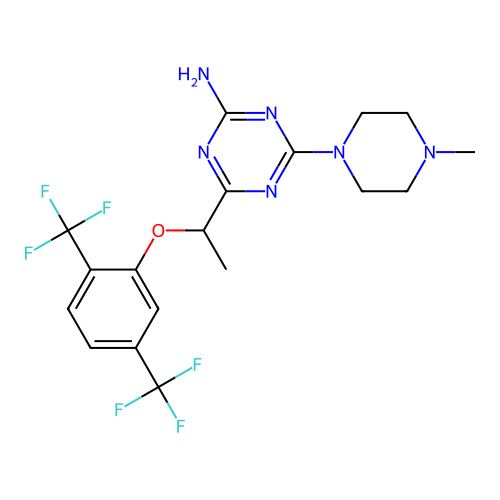
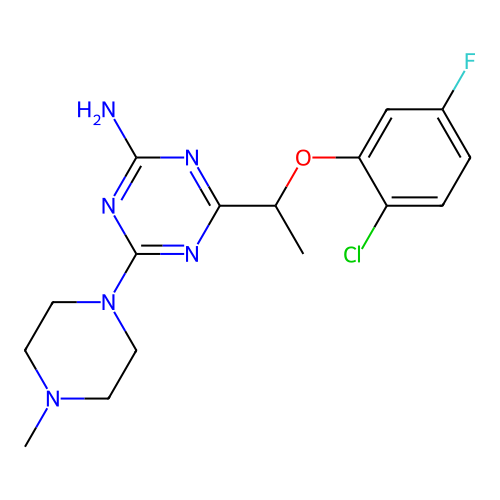
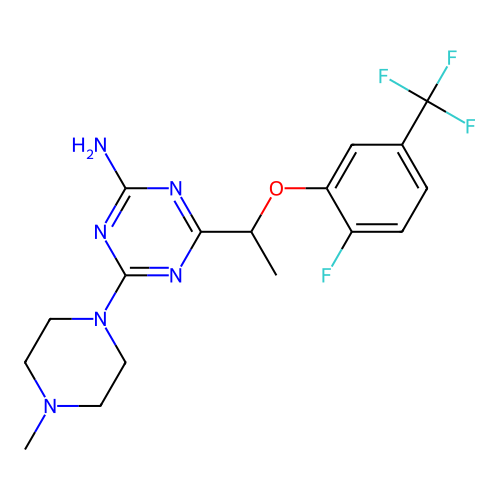
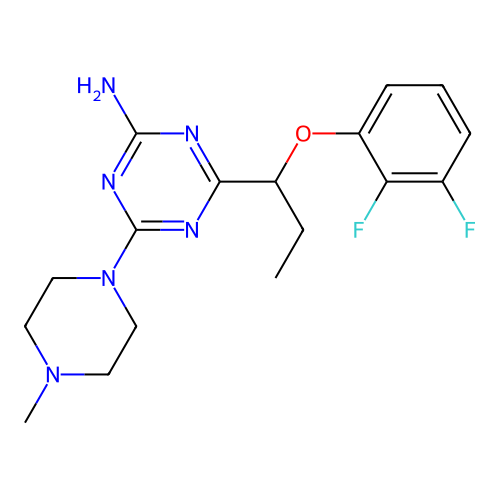
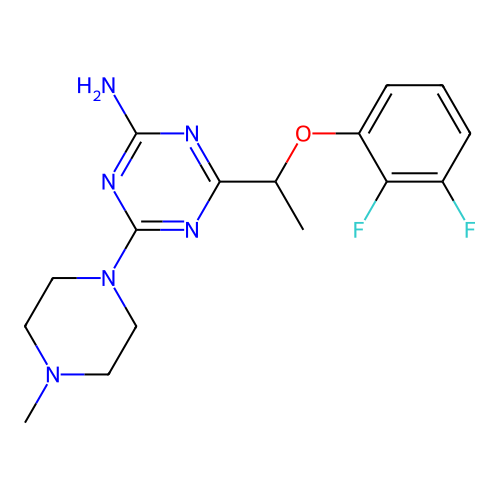
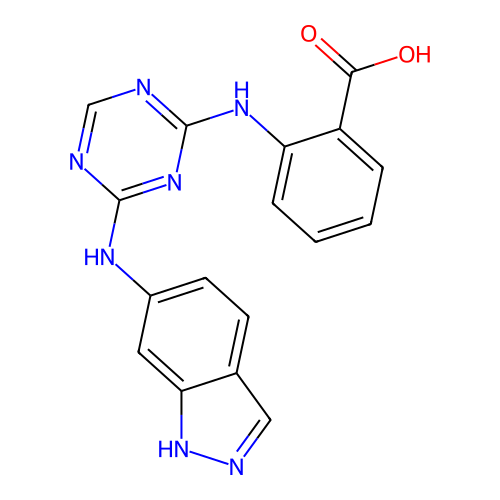
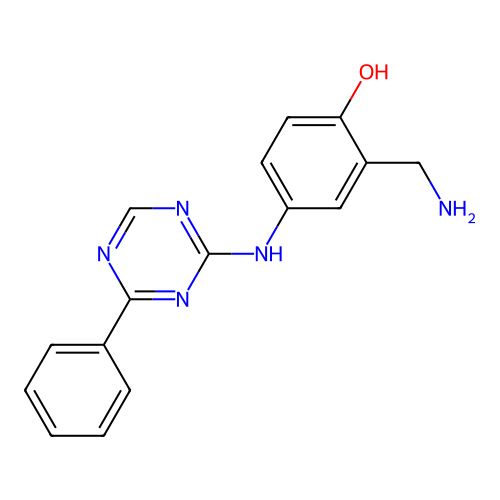
Report error Found 141 Enz. Inhib. hit(s) with all data for entry = 50022417
TargetPlatelet-derived growth factor receptor alpha/beta(Mouse)
Jagiellonian University
Curated by ChEMBL
Jagiellonian University
Curated by ChEMBL
Affinity DataIC50: 3nMAssay Description:Inhibition of CDK5/p25 (unknown origin)More data for this Ligand-Target Pair
Affinity DataKi: 5nMAssay Description:Displacement of [3H]-LSD from human 5-HT6 receptor expressed in HEK293 cell membranes assessed as inhibition constant incubated for 1 hr by Microbeta...More data for this Ligand-Target Pair
Affinity DataKi: 6nMAssay Description:Displacement of [3H]-LSD from human 5-HT6 receptor expressed in HEK293 cell membranes assessed as inhibition constant incubated for 1 hr by Microbeta...More data for this Ligand-Target Pair
Affinity DataKi: 6nMAssay Description:Displacement of [3H]-LSD from human 5-HT6 receptor expressed in HEK293 cell membranes assessed as inhibition constant incubated for 1 hr by Microbeta...More data for this Ligand-Target Pair
Affinity DataKi: 7nMAssay Description:Displacement of [3H]-LSD from human 5-HT6 receptor expressed in HEK293 cell membranes assessed as inhibition constant incubated for 1 hr by Microbeta...More data for this Ligand-Target Pair
Affinity DataKi: 7nMAssay Description:Displacement of [3H]-LSD from human 5-HT6 receptor expressed in HEK293 cell membranes assessed as inhibition constant incubated for 1 hr by Microbeta...More data for this Ligand-Target Pair
Affinity DataKi: 8nMAssay Description:Displacement of [3H]-LSD from human 5-HT6 receptor expressed in HEK293 cell membranes assessed as inhibition constant incubated for 1 hr by Microbeta...More data for this Ligand-Target Pair
Affinity DataKi: 9nMAssay Description:Displacement of [3H]-raclopride from human D2 receptor expressed in HEK293 cell membranes assessed as inhibition constant incubated for 1 hr by Micro...More data for this Ligand-Target Pair
Affinity DataKi: 11nMAssay Description:Displacement of [3H]-LSD from human 5-HT6 receptor expressed in HEK293 cell membranes assessed as inhibition constant incubated for 1 hr by Microbeta...More data for this Ligand-Target Pair
Affinity DataKi: 12nMAssay Description:Displacement of [3H]-LSD from human 5-HT6 receptor expressed in HEK293 cell membranes assessed as inhibition constant incubated for 1 hr by Microbeta...More data for this Ligand-Target Pair
Affinity DataKi: 15nMAssay Description:Displacement of [3H]-LSD from human 5-HT6 receptor expressed in HEK293 cell membranes assessed as inhibition constant incubated for 1 hr by Microbeta...More data for this Ligand-Target Pair
Affinity DataKi: 15nMAssay Description:Displacement of [3H]-LSD from human 5-HT6 receptor expressed in HEK293 cell membranes assessed as inhibition constant incubated for 1 hr by Microbeta...More data for this Ligand-Target Pair
Affinity DataKi: 18nMAssay Description:Displacement of [3H]-LSD from human 5-HT6 receptor expressed in HEK293 cell membranes assessed as inhibition constant incubated for 1 hr by Microbeta...More data for this Ligand-Target Pair
Affinity DataKi: 21nMAssay Description:Displacement of [3H]-ketanserin from human 5-HT2A receptor expressed in HEK293 cell membranes assessed as inhibition constant incubated for 1 hr by M...More data for this Ligand-Target Pair
Affinity DataKi: 21nMAssay Description:Displacement of [3H]-LSD from human 5-HT6 receptor expressed in HEK293 cell membranes assessed as inhibition constant incubated for 1 hr by Microbeta...More data for this Ligand-Target Pair
Affinity DataKi: 23nMAssay Description:Displacement of [3H]-LSD from human 5-HT6 receptor expressed in HEK293 cell membranes assessed as inhibition constant incubated for 1 hr by Microbeta...More data for this Ligand-Target Pair
Affinity DataKi: 30nMAssay Description:Displacement of [3H]-LSD from human 5-HT6 receptor expressed in HEK293 cell membranes assessed as inhibition constant incubated for 1 hr by Microbeta...More data for this Ligand-Target Pair
Affinity DataKi: 31nMAssay Description:Displacement of [3H]-LSD from human 5-HT6 receptor expressed in HEK293 cell membranes assessed as inhibition constant incubated for 1 hr by Microbeta...More data for this Ligand-Target Pair
Affinity DataKi: 31nMAssay Description:Displacement of [3H]-LSD from human 5-HT6 receptor expressed in HEK293 cell membranes assessed as inhibition constant incubated for 1 hr by Microbeta...More data for this Ligand-Target Pair
Affinity DataKi: 32nMAssay Description:Displacement of [3H]-8-OH-DPAT from human 5-HT1A receptor expressed in HEK293 cell membranes assessed as inhibition constant incubated for 1 hr by Mi...More data for this Ligand-Target Pair
Affinity DataKi: 35nMAssay Description:Displacement of [3H]-LSD from human 5-HT6 receptor expressed in HEK293 cell membranes assessed as inhibition constant incubated for 1 hr by Microbeta...More data for this Ligand-Target Pair
Affinity DataKi: 39nMAssay Description:Displacement of [3H]-LSD from human 5-HT6 receptor expressed in HEK293 cell membranes assessed as inhibition constant incubated for 1 hr by Microbeta...More data for this Ligand-Target Pair
Affinity DataKi: 52nMAssay Description:Displacement of [3H]-LSD from human 5-HT6 receptor expressed in HEK293 cell membranes assessed as inhibition constant incubated for 1 hr by Microbeta...More data for this Ligand-Target Pair
Affinity DataKi: 55nMAssay Description:Displacement of [3H]-LSD from human 5-HT6 receptor expressed in HEK293 cell membranes assessed as inhibition constant incubated for 1 hr by Microbeta...More data for this Ligand-Target Pair
TargetPlatelet-derived growth factor receptor alpha/beta(Mouse)
Jagiellonian University
Curated by ChEMBL
Jagiellonian University
Curated by ChEMBL
Affinity DataIC50: 55nMAssay Description:Inhibition of CDK5/p25 (unknown origin)More data for this Ligand-Target Pair
Affinity DataKi: 56nMAssay Description:Displacement of [3H]-LSD from human 5-HT6 receptor expressed in HEK293 cell membranes assessed as inhibition constant incubated for 1 hr by Microbeta...More data for this Ligand-Target Pair
Affinity DataKi: 58nMAssay Description:Displacement of [3H]-LSD from human 5-HT6 receptor expressed in HEK293 cell membranes assessed as inhibition constant incubated for 1 hr by Microbeta...More data for this Ligand-Target Pair
Affinity DataKi: 60nMAssay Description:Displacement of [3H]-LSD from human 5-HT6 receptor expressed in HEK293 cell membranes assessed as inhibition constant incubated for 1 hr by Microbeta...More data for this Ligand-Target Pair
Affinity DataKi: 62nMAssay Description:Displacement of [3H]-5-CT from human 5-HT7 receptor expressed in HEK293 cell membranes assessed as inhibition constant incubated for 1 hr by Microbet...More data for this Ligand-Target Pair
Affinity DataKi: 66nMAssay Description:Displacement of [3H]-LSD from human 5-HT6 receptor expressed in HEK293 cell membranes assessed as inhibition constant incubated for 1 hr by Microbeta...More data for this Ligand-Target Pair
Affinity DataKi: 72nMAssay Description:Displacement of [3H]-LSD from human 5-HT6 receptor expressed in HEK293 cell membranes assessed as inhibition constant incubated for 1 hr by Microbeta...More data for this Ligand-Target Pair
Affinity DataKi: 94nMAssay Description:Displacement of [3H]-LSD from human 5-HT6 receptor expressed in HEK293 cell membranes assessed as inhibition constant incubated for 1 hr by Microbeta...More data for this Ligand-Target Pair
Affinity DataKi: 94nMAssay Description:Displacement of [3H]-LSD from human 5-HT6 receptor expressed in HEK293 cell membranes assessed as inhibition constant incubated for 1 hr by Microbeta...More data for this Ligand-Target Pair
Affinity DataKi: 113nMAssay Description:Displacement of [3H]-ketanserin from human 5-HT2A receptor expressed in HEK293 cell membranes assessed as inhibition constant incubated for 1 hr by M...More data for this Ligand-Target Pair
Affinity DataKi: 117nMAssay Description:Displacement of [3H]-ketanserin from human 5-HT2A receptor expressed in HEK293 cell membranes assessed as inhibition constant incubated for 1 hr by M...More data for this Ligand-Target Pair
TargetPlatelet-derived growth factor receptor alpha/beta(Mouse)
Jagiellonian University
Curated by ChEMBL
Jagiellonian University
Curated by ChEMBL
Affinity DataIC50: 143nMAssay Description:Inhibition of full length human recombinant CDK5/p25 using histone H1 as substrate assessed as residual activity incubated for 15 mins by luciferase ...More data for this Ligand-Target Pair
TargetPlatelet-derived growth factor receptor alpha/beta(Mouse)
Jagiellonian University
Curated by ChEMBL
Jagiellonian University
Curated by ChEMBL
Affinity DataIC50: 160nMAssay Description:Inhibition of CDK5/p25 (unknown origin)More data for this Ligand-Target Pair
Affinity DataKi: 209nMAssay Description:Displacement of [3H]-ketanserin from human 5-HT2A receptor expressed in HEK293 cell membranes assessed as inhibition constant incubated for 1 hr by M...More data for this Ligand-Target Pair
TargetPlatelet-derived growth factor receptor alpha/beta(Mouse)
Jagiellonian University
Curated by ChEMBL
Jagiellonian University
Curated by ChEMBL
Affinity DataKi: 251nMAssay Description:Binding affinity to CDK5/p25 (unknown origin)More data for this Ligand-Target Pair
Affinity DataKi: 260nMAssay Description:Displacement of [3H]-raclopride from human D2 receptor expressed in HEK293 cell membranes assessed as inhibition constant incubated for 1 hr by Micro...More data for this Ligand-Target Pair
Affinity DataKi: 281nMAssay Description:Displacement of [3H]-ketanserin from human 5-HT2A receptor expressed in HEK293 cell membranes assessed as inhibition constant incubated for 1 hr by M...More data for this Ligand-Target Pair
Affinity DataKi: 287nMAssay Description:Displacement of [3H]-ketanserin from human 5-HT2A receptor expressed in HEK293 cell membranes assessed as inhibition constant incubated for 1 hr by M...More data for this Ligand-Target Pair
Affinity DataKi: 311nMAssay Description:Displacement of [3H]-ketanserin from human 5-HT2A receptor expressed in HEK293 cell membranes assessed as inhibition constant incubated for 1 hr by M...More data for this Ligand-Target Pair
Affinity DataKi: 320nMAssay Description:Displacement of [3H]-raclopride from human D2 receptor expressed in HEK293 cell membranes assessed as inhibition constant incubated for 1 hr by Micro...More data for this Ligand-Target Pair
Affinity DataKi: 338nMAssay Description:Displacement of [3H]-ketanserin from human 5-HT2A receptor expressed in HEK293 cell membranes assessed as inhibition constant incubated for 1 hr by M...More data for this Ligand-Target Pair
TargetPlatelet-derived growth factor receptor alpha/beta(Mouse)
Jagiellonian University
Curated by ChEMBL
Jagiellonian University
Curated by ChEMBL
Affinity DataKi: 398nMAssay Description:Binding affinity to CDK5/p25 (unknown origin)More data for this Ligand-Target Pair
Affinity DataKi: 408nMAssay Description:Displacement of [3H]-raclopride from human D2 receptor expressed in HEK293 cell membranes assessed as inhibition constant incubated for 1 hr by Micro...More data for this Ligand-Target Pair
Affinity DataKi: 411nMAssay Description:Displacement of [3H]-raclopride from human D2 receptor expressed in HEK293 cell membranes assessed as inhibition constant incubated for 1 hr by Micro...More data for this Ligand-Target Pair
Affinity DataKi: 416nMAssay Description:Displacement of [3H]-raclopride from human D2 receptor expressed in HEK293 cell membranes assessed as inhibition constant incubated for 1 hr by Micro...More data for this Ligand-Target Pair
Affinity DataKi: 421nMAssay Description:Displacement of [3H]-raclopride from human D2 receptor expressed in HEK293 cell membranes assessed as inhibition constant incubated for 1 hr by Micro...More data for this Ligand-Target Pair
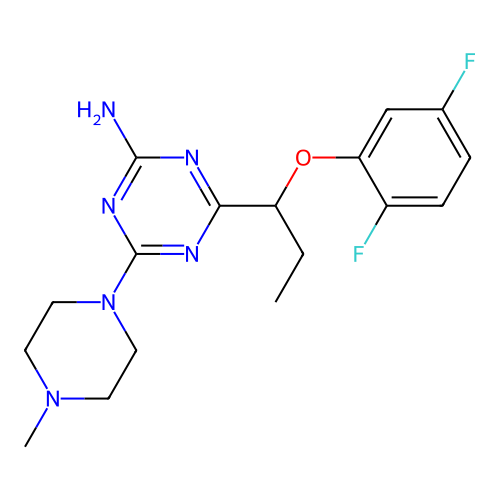
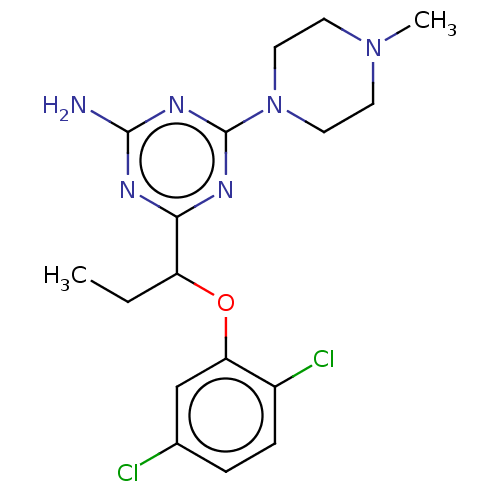
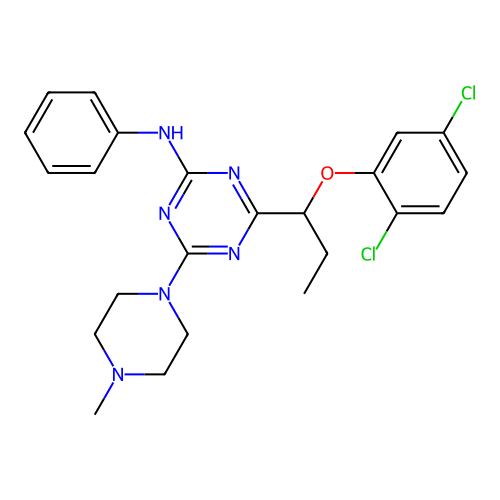
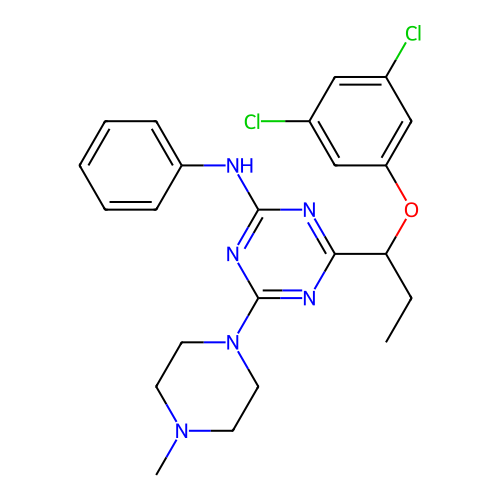
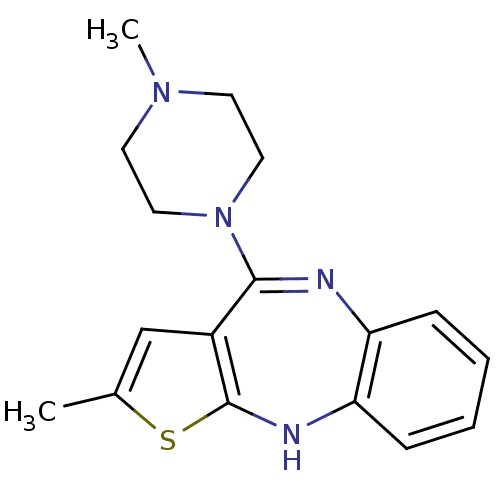
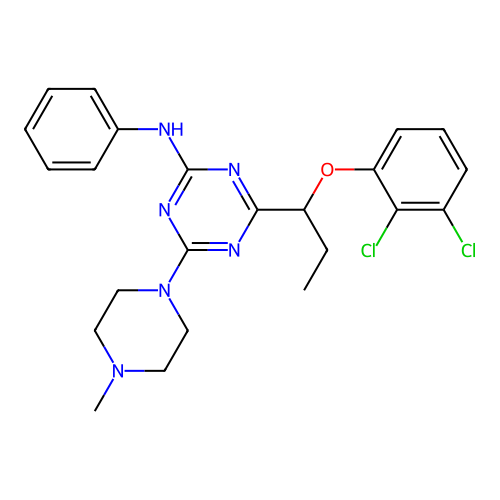
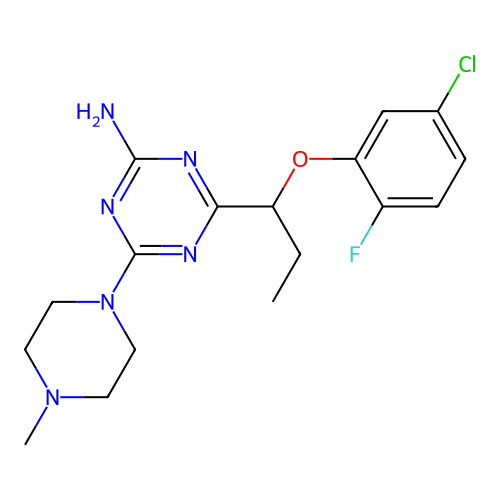
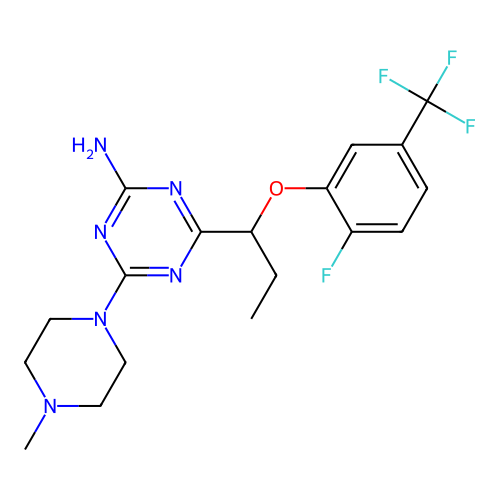
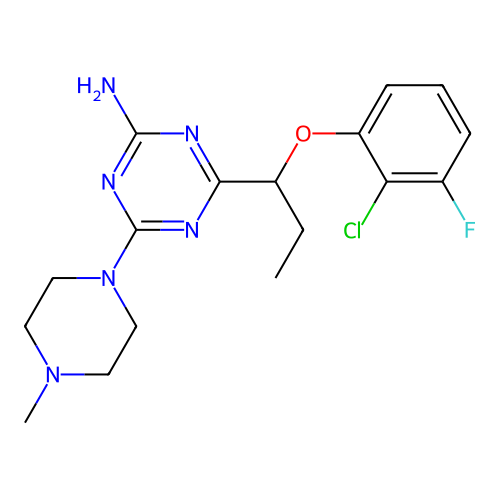
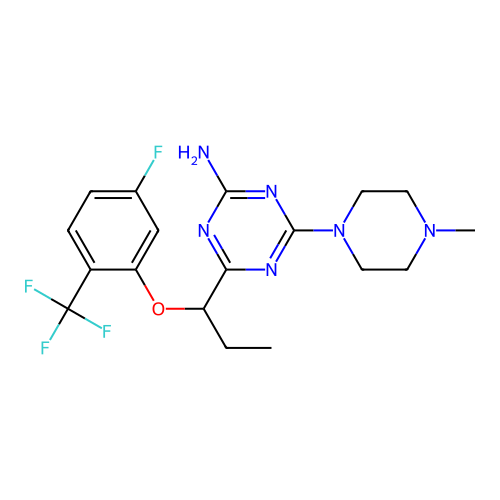
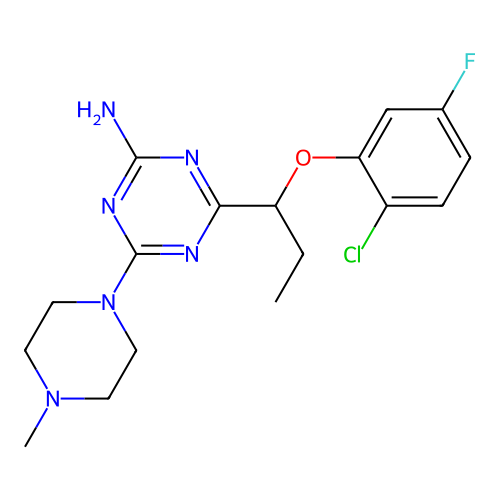
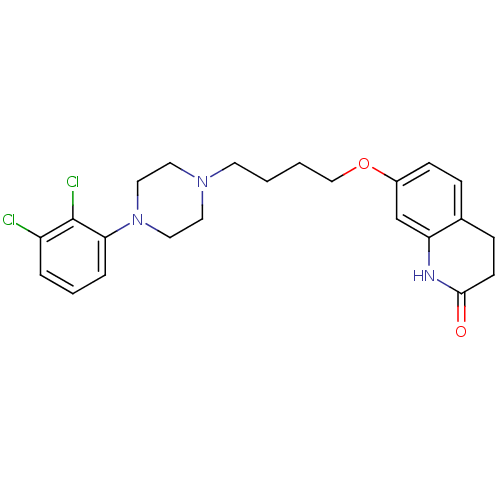
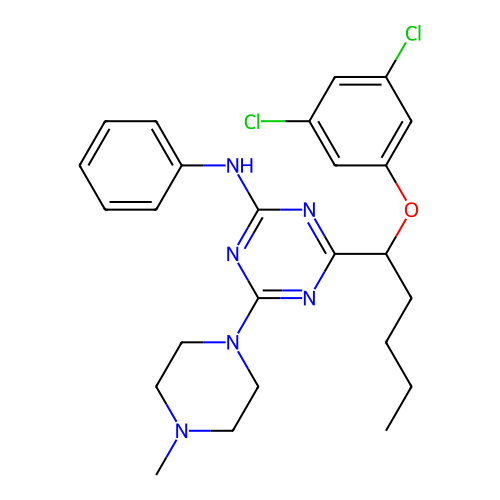
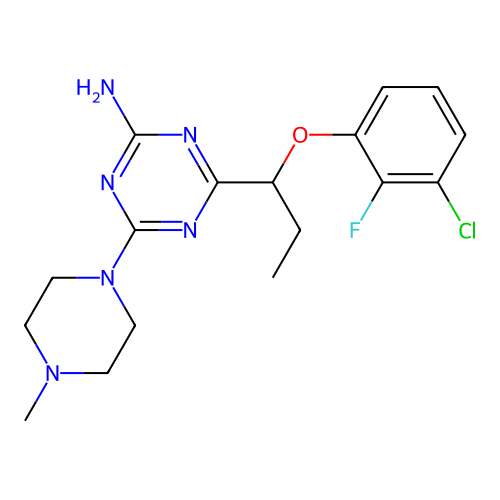
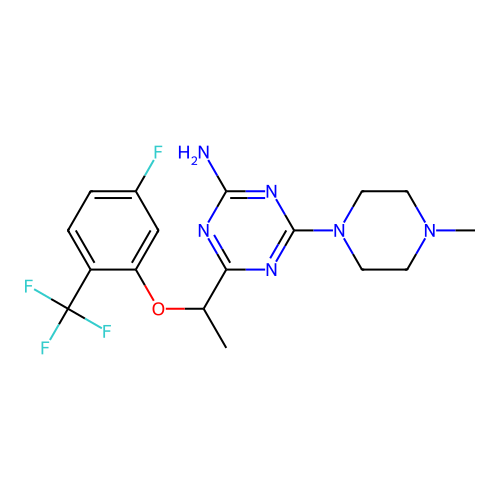
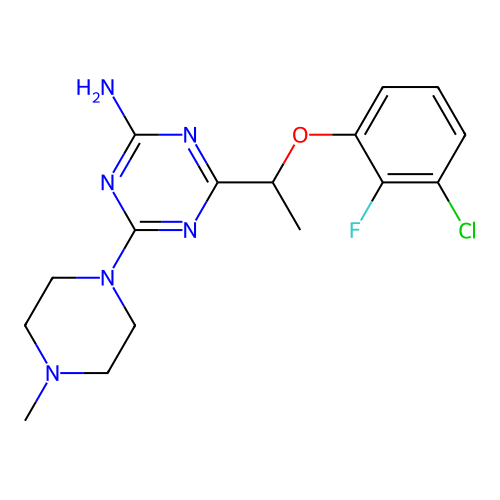
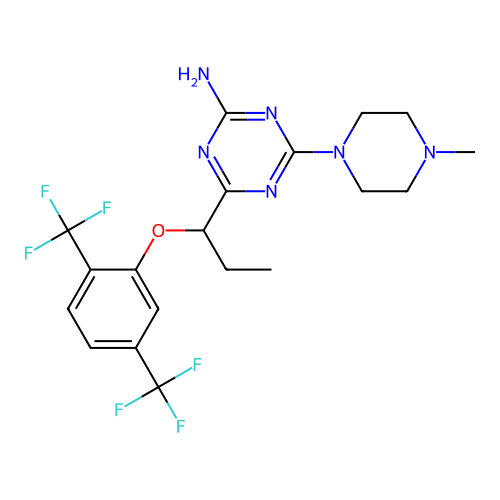
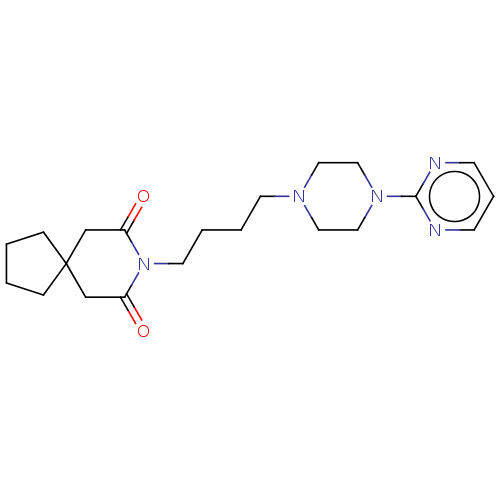
 3D Structure (crystal)
3D Structure (crystal)