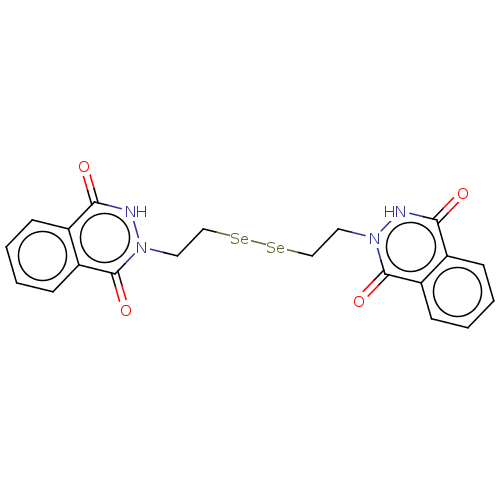
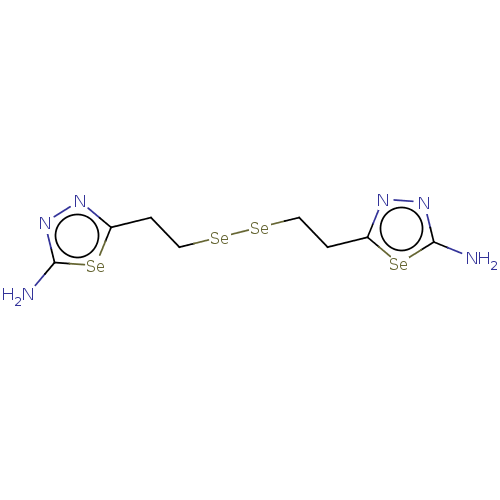
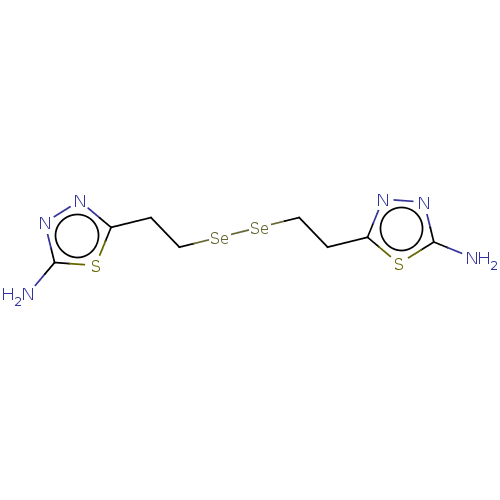
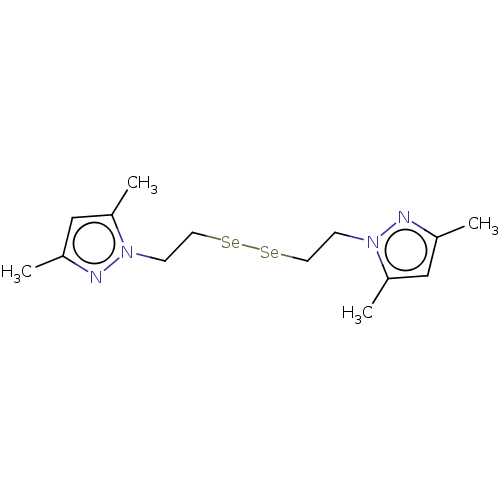
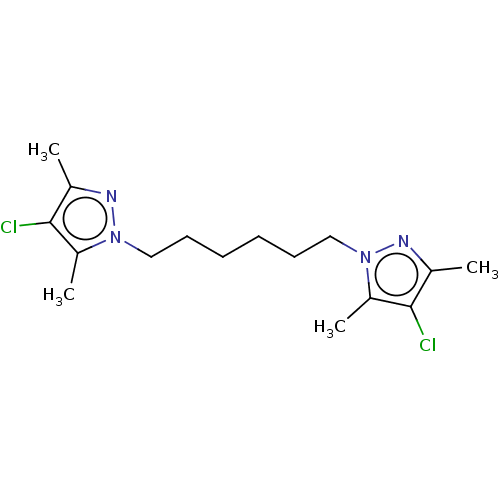
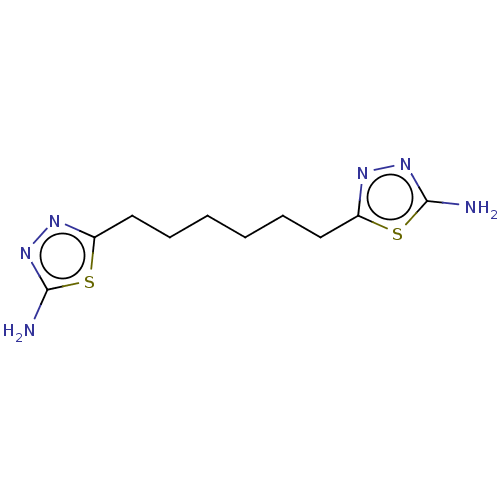
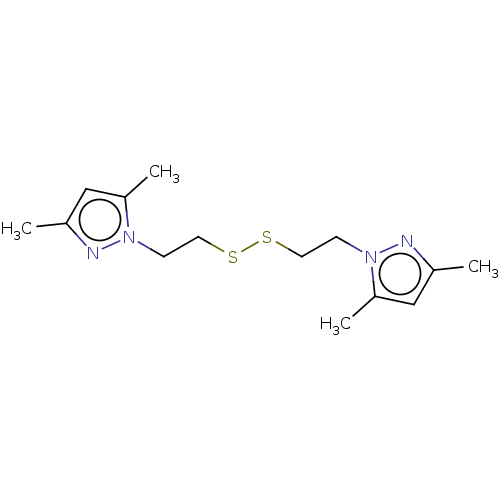
Report error Found 39 Enz. Inhib. hit(s) with all data for entry = 50020177
Affinity DataIC50: 2nMAssay Description:Negative allosteric modulator activity at human wildtype KGA incubated for 5 hrs in presence of GDH and NADP by EZMTT assayMore data for this Ligand-Target Pair
Affinity DataIC50: 17nMAssay Description:Negative allosteric modulator activity at human wildtype KGA using glutamine as substrate preincubated for 1 hr followed by substrate addition in pre...More data for this Ligand-Target Pair
Affinity DataIC50: 200nMAssay Description:Negative allosteric modulator activity at human wildtype KGA incubated for 5 hrs in presence of GDH and NADP by EZMTT assayMore data for this Ligand-Target Pair
Affinity DataIC50: 400nMAssay Description:Negative allosteric modulator activity at human wildtype KGA incubated for 5 hrs in presence of GDH and NADP by EZMTT assayMore data for this Ligand-Target Pair
Affinity DataIC50: 400nMAssay Description:Negative allosteric modulator activity at human wildtype KGA incubated for 5 hrs in presence of GDH and NADP by EZMTT assayMore data for this Ligand-Target Pair
Affinity DataIC50: 500nMAssay Description:Negative allosteric modulator activity at Escherichia coli TrxR using DTNB as substrate preincubated for 20 mins followed by substrate addition in pr...More data for this Ligand-Target Pair
Affinity DataIC50: 540nMAssay Description:Negative allosteric modulator activity at human wildtype KGA incubated for 5 hrs in presence of GDH and NADP by EZMTT assayMore data for this Ligand-Target Pair
Affinity DataIC50: 700nMAssay Description:Negative allosteric modulator activity at Escherichia coli TrxR using DTNB as substrate preincubated for 20 mins followed by substrate addition in pr...More data for this Ligand-Target Pair
Affinity DataIC50: 700nMAssay Description:Negative allosteric modulator activity at Escherichia coli TrxR using DTNB as substrate preincubated for 20 mins followed by substrate addition in pr...More data for this Ligand-Target Pair
Affinity DataIC50: 700nMAssay Description:Negative allosteric modulator activity at Escherichia coli TrxR using DTNB as substrate preincubated for 20 mins followed by substrate addition in pr...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+3nMAssay Description:Negative allosteric modulator activity at human wildtype KGA incubated for 5 hrs in presence of GDH and NADP by EZMTT assayMore data for this Ligand-Target Pair
Affinity DataIC50: 2.10E+3nMAssay Description:Negative allosteric modulator activity at Escherichia coli TrxR using DTNB as substrate preincubated for 20 mins followed by substrate addition in pr...More data for this Ligand-Target Pair
Affinity DataIC50: 2.10E+3nMAssay Description:Negative allosteric modulator activity at human wildtype KGA using glutamine as substrate preincubated for 1 hr followed by substrate addition in pre...More data for this Ligand-Target Pair
Affinity DataIC50: 2.20E+3nMAssay Description:Negative allosteric modulator activity at human wildtype KGA using glutamine as substrate preincubated for 1 hr followed by substrate addition in pre...More data for this Ligand-Target Pair
Affinity DataIC50: 3.00E+3nMAssay Description:Negative allosteric modulator activity at human wildtype KGA incubated for 5 hrs in presence of GDH and NADP by EZMTT assayMore data for this Ligand-Target Pair
Affinity DataIC50: 3.00E+3nMAssay Description:Negative allosteric modulator activity at Escherichia coli TrxR using DTNB as substrate preincubated for 20 mins followed by substrate addition in pr...More data for this Ligand-Target Pair
Affinity DataIC50: 3.60E+3nMAssay Description:Negative allosteric modulator activity at human wildtype KGA incubated for 5 hrs in presence of GDH and NADP by EZMTT assayMore data for this Ligand-Target Pair
Affinity DataIC50: 3.70E+3nMAssay Description:Negative allosteric modulator activity at Escherichia coli TrxR using DTNB as substrate preincubated for 20 mins followed by substrate addition in pr...More data for this Ligand-Target Pair
Affinity DataIC50: 4.00E+3nMAssay Description:Negative allosteric modulator activity at Escherichia coli TrxR using DTNB as substrate preincubated for 20 mins followed by substrate addition in pr...More data for this Ligand-Target Pair
Affinity DataIC50: 4.00E+3nMAssay Description:Negative allosteric modulator activity at human wildtype KGA incubated for 5 hrs in presence of GDH and NADP by EZMTT assayMore data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMAssay Description:Negative allosteric modulator activity at Escherichia coli TrxR using DTNB as substrate preincubated for 20 mins followed by substrate addition in pr...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMAssay Description:Negative allosteric modulator activity at Escherichia coli TrxR using DTNB as substrate preincubated for 20 mins followed by substrate addition in pr...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMAssay Description:Negative allosteric modulator activity at human wildtype KGA incubated for 5 hrs in presence of GDH and NADP by EZMTT assayMore data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMAssay Description:Negative allosteric modulator activity at Escherichia coli TrxR using DTNB as substrate preincubated for 20 mins followed by substrate addition in pr...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMAssay Description:Negative allosteric modulator activity at human wildtype KGA incubated for 5 hrs in presence of GDH and NADP by EZMTT assayMore data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMAssay Description:Negative allosteric modulator activity at Escherichia coli TrxR using DTNB as substrate preincubated for 20 mins followed by substrate addition in pr...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMAssay Description:Negative allosteric modulator activity at human wildtype KGA incubated for 5 hrs in presence of GDH and NADP by EZMTT assayMore data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMAssay Description:Negative allosteric modulator activity at Escherichia coli TrxR using DTNB as substrate preincubated for 20 mins followed by substrate addition in pr...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMAssay Description:Negative allosteric modulator activity at human wildtype KGA incubated for 5 hrs in presence of GDH and NADP by EZMTT assayMore data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMAssay Description:Negative allosteric modulator activity at human wildtype KGA incubated for 5 hrs in presence of GDH and NADP by EZMTT assayMore data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMAssay Description:Negative allosteric modulator activity at human wildtype KGA incubated for 5 hrs in presence of GDH and NADP by EZMTT assayMore data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMAssay Description:Negative allosteric modulator activity at human wildtype KGA incubated for 5 hrs in presence of GDH and NADP by EZMTT assayMore data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMAssay Description:Negative allosteric modulator activity at human wildtype KGA incubated for 5 hrs in presence of GDH and NADP by EZMTT assayMore data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMAssay Description:Negative allosteric modulator activity at human wildtype KGA incubated for 5 hrs in presence of GDH and NADP by EZMTT assayMore data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMAssay Description:Negative allosteric modulator activity at Escherichia coli TrxR using DTNB as substrate preincubated for 20 mins followed by substrate addition in pr...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMAssay Description:Negative allosteric modulator activity at Escherichia coli TrxR using DTNB as substrate preincubated for 20 mins followed by substrate addition in pr...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMAssay Description:Negative allosteric modulator activity at Escherichia coli TrxR using DTNB as substrate preincubated for 20 mins followed by substrate addition in pr...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMAssay Description:Negative allosteric modulator activity at Escherichia coli TrxR using DTNB as substrate preincubated for 20 mins followed by substrate addition in pr...More data for this Ligand-Target Pair
Affinity DataIC50: 1.30E+4nMAssay Description:Negative allosteric modulator activity at Escherichia coli TrxR using DTNB as substrate preincubated for 20 mins followed by substrate addition in pr...More data for this Ligand-Target Pair
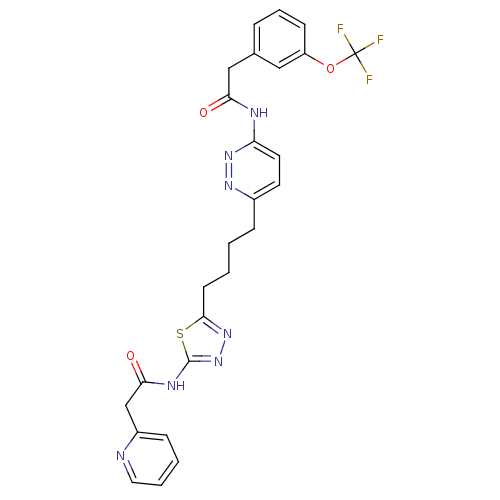
 3D Structure (crystal)
3D Structure (crystal)