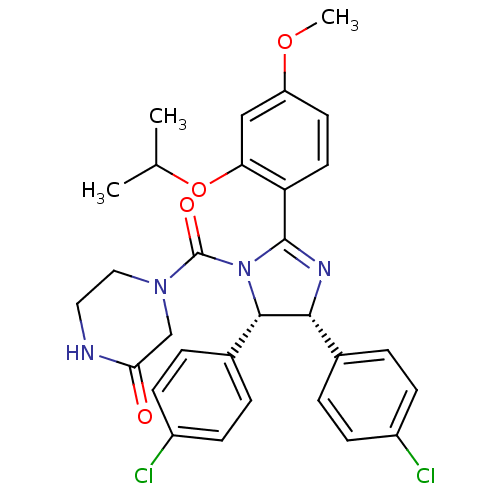
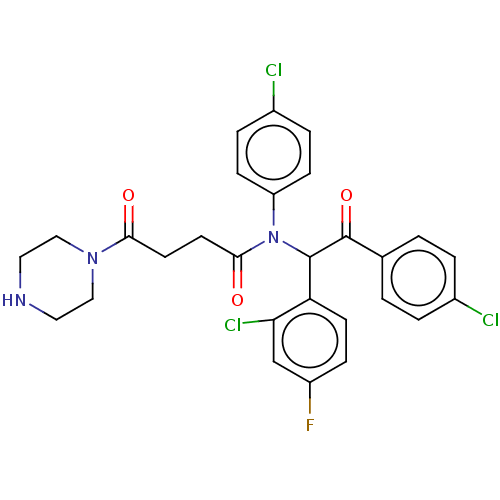
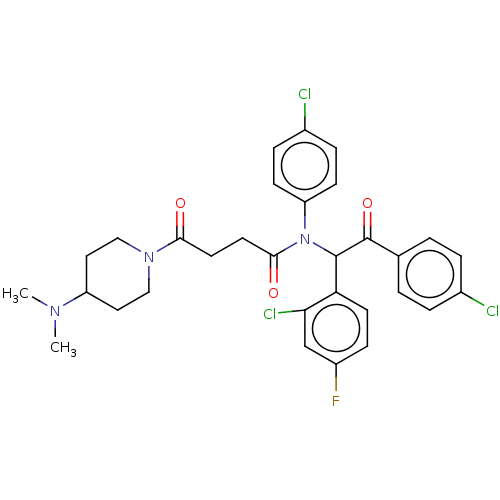
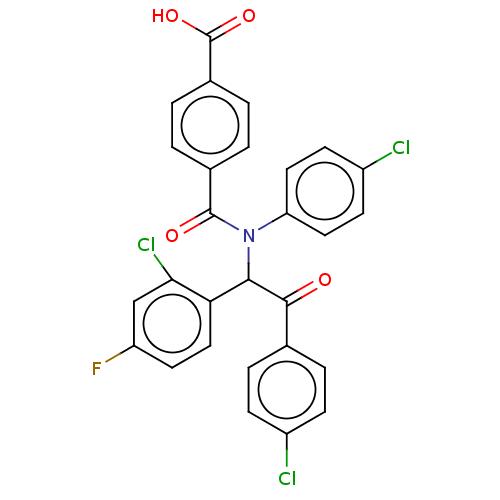
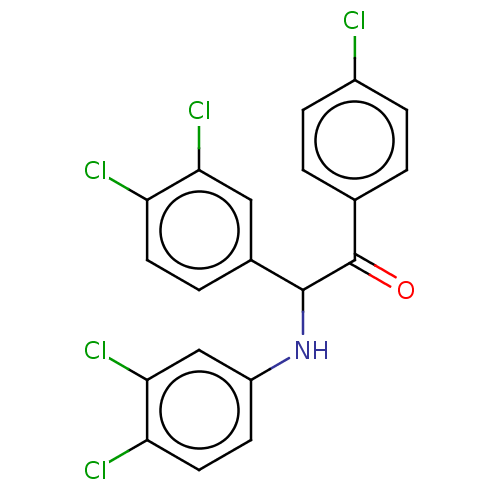
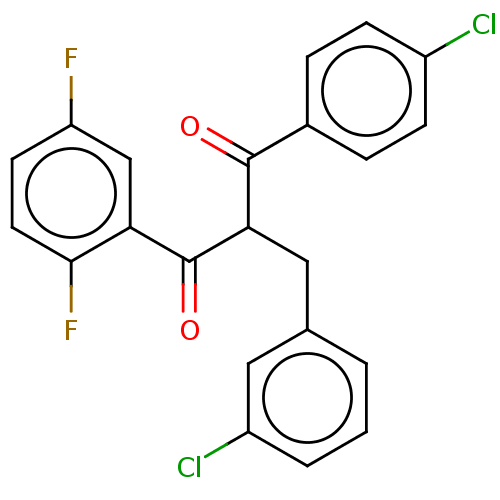
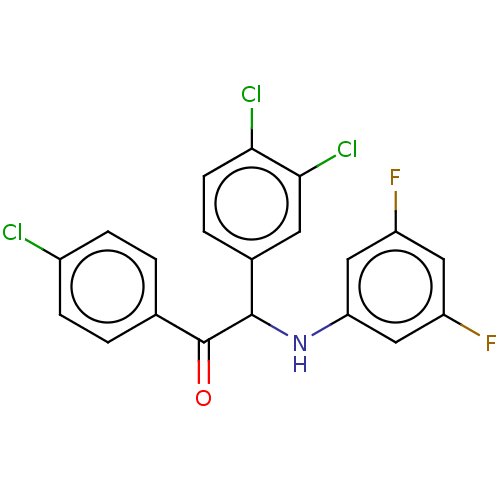
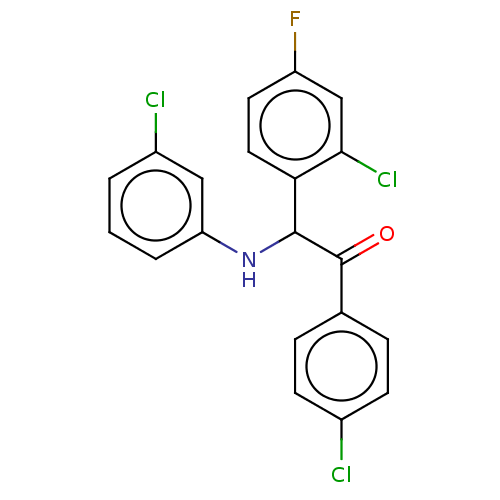
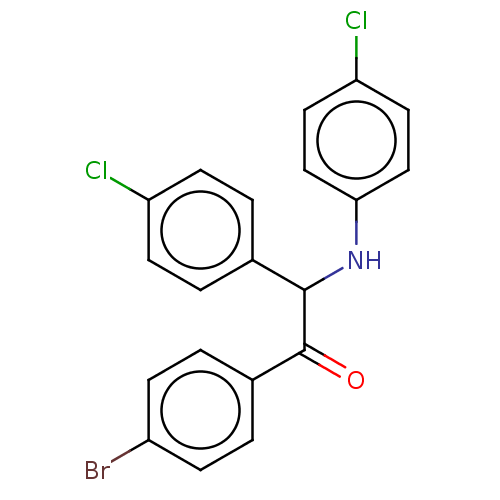
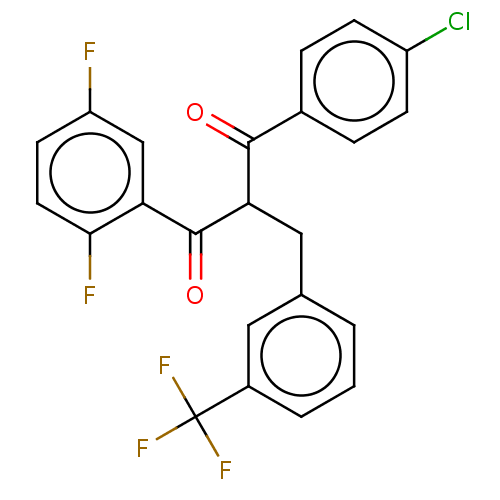
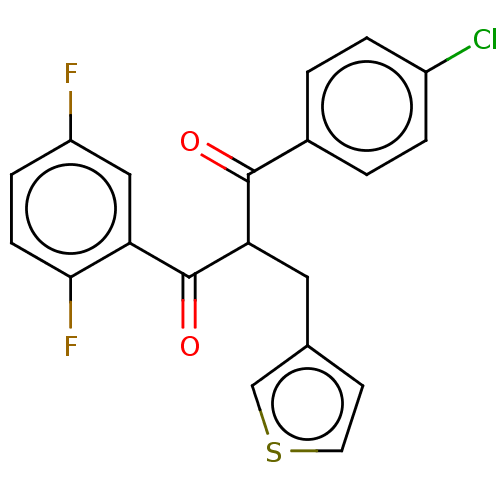
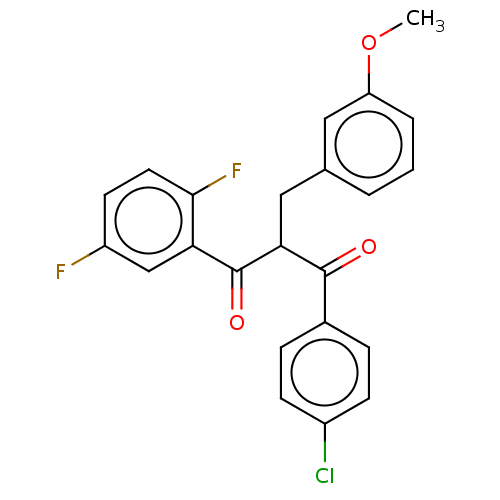
Report error Found 64 Enz. Inhib. hit(s) with all data for entry = 50020150
TargetE3 ubiquitin-protein ligase Mdm2(Human)
Nanjing University of Chinese Medicine
Curated by ChEMBL
Nanjing University of Chinese Medicine
Curated by ChEMBL
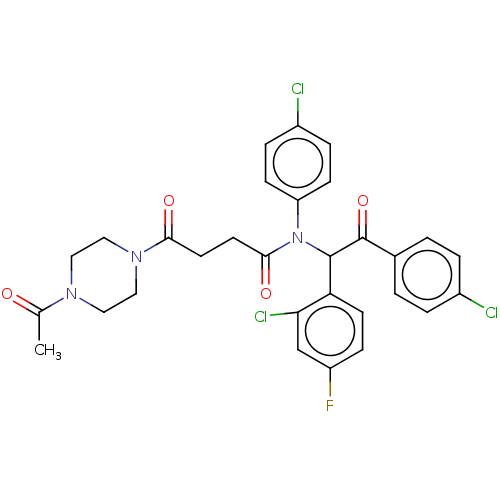
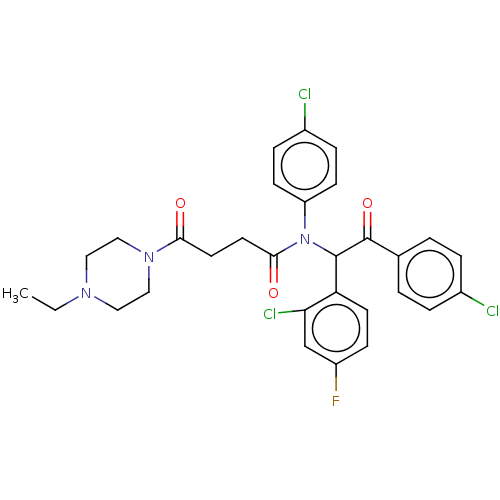
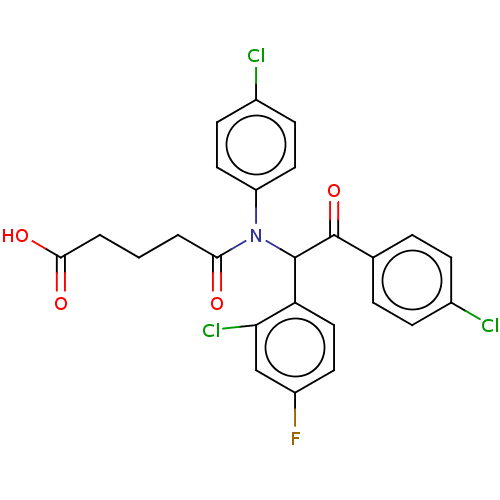
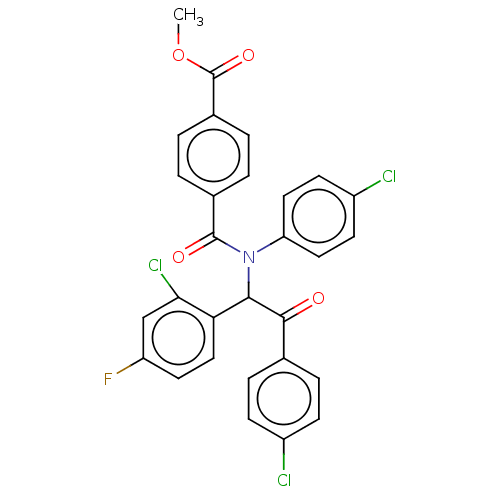
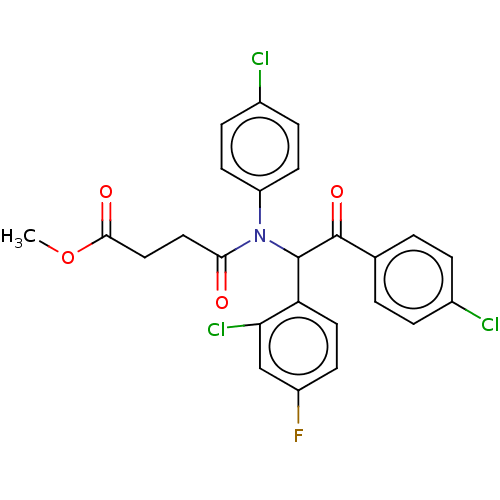
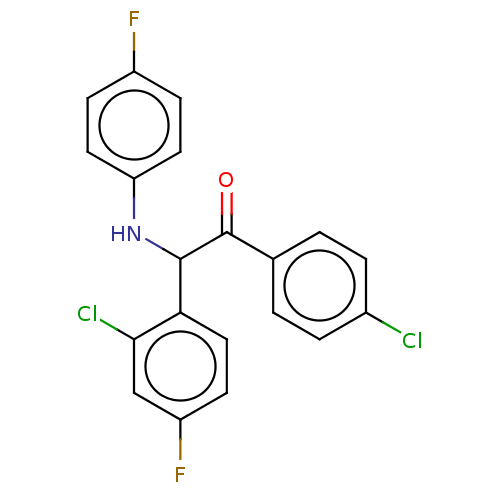
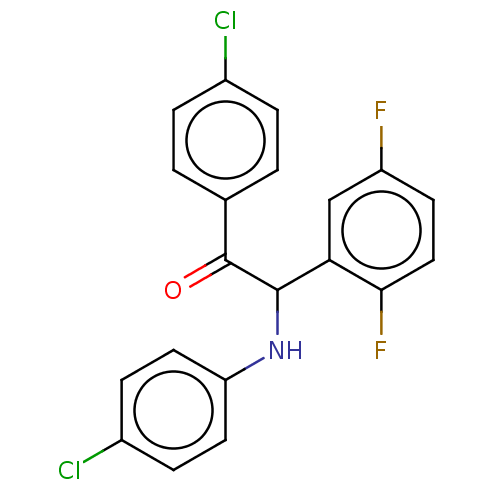
Affinity DataKi: 110nMAssay Description:Binding affinity to MDM2 (1 to 118 residues) (unknown origin) assessed as inhibition constant incubated for 1 hr measured by fluorescence polarizatio...More data for this Ligand-Target Pair
TargetE3 ubiquitin-protein ligase Mdm2(Human)
Nanjing University of Chinese Medicine
Curated by ChEMBL
Nanjing University of Chinese Medicine
Curated by ChEMBL
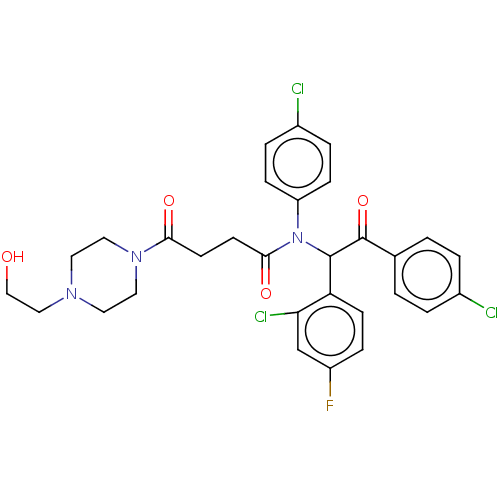
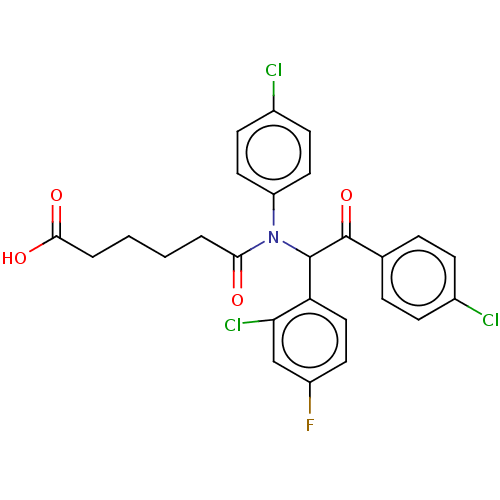
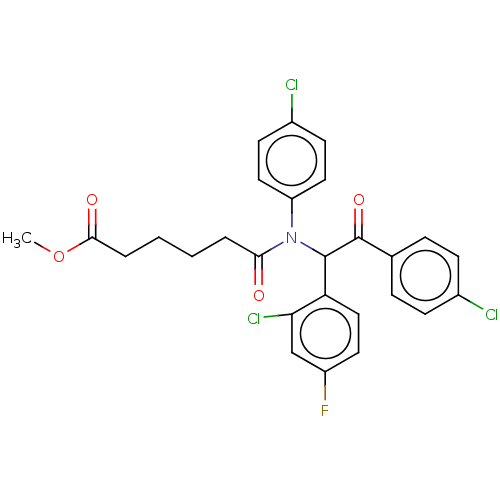
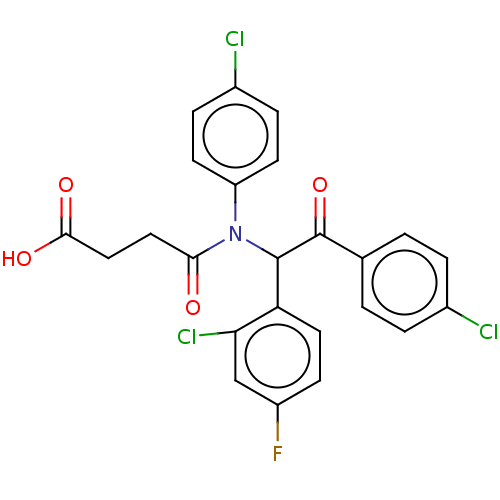
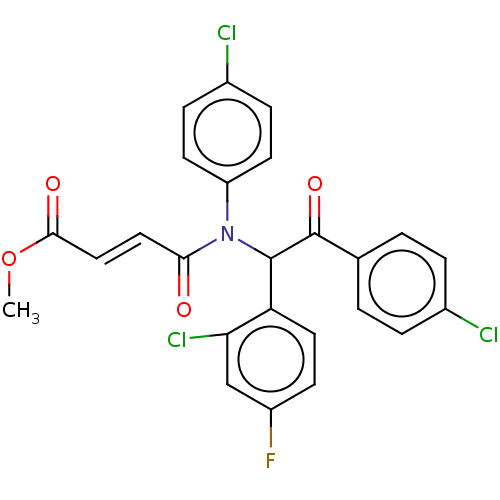
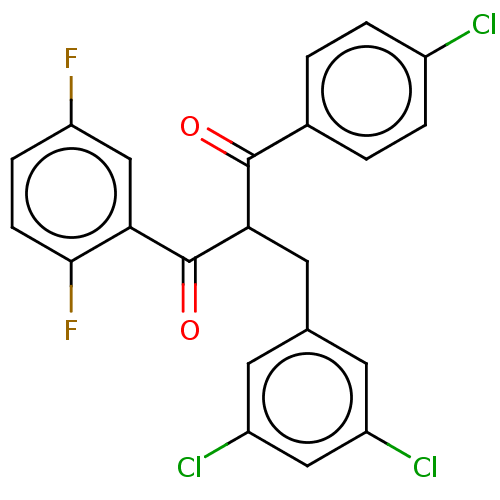
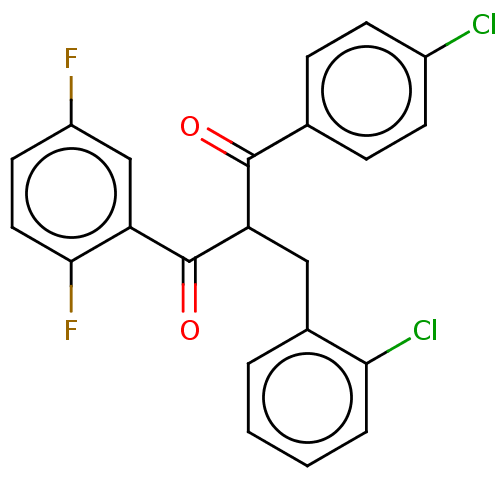
Affinity DataKi: 230nMAssay Description:Binding affinity to MDM2 (1 to 118 residues) (unknown origin) assessed as inhibition constant incubated for 1 hr measured by fluorescence polarizatio...More data for this Ligand-Target Pair
TargetE3 ubiquitin-protein ligase Mdm2(Human)
Nanjing University of Chinese Medicine
Curated by ChEMBL
Nanjing University of Chinese Medicine
Curated by ChEMBL
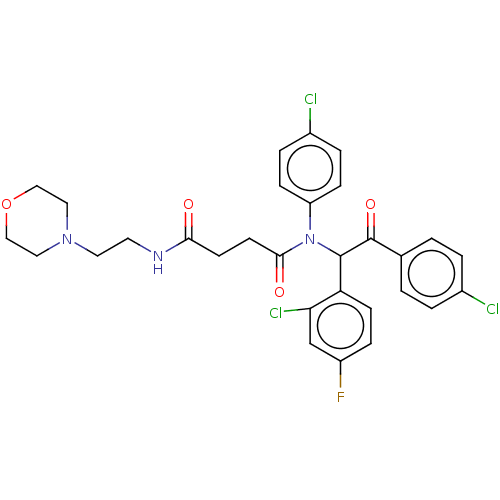
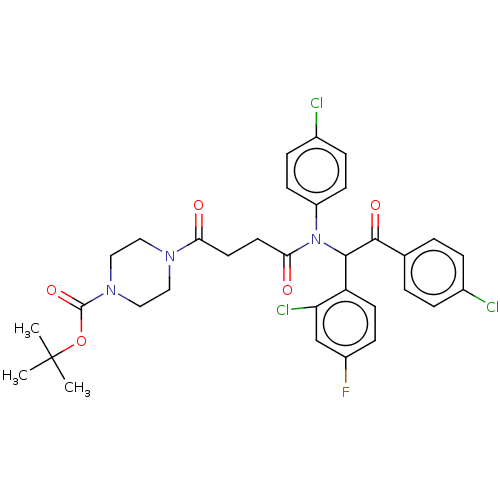
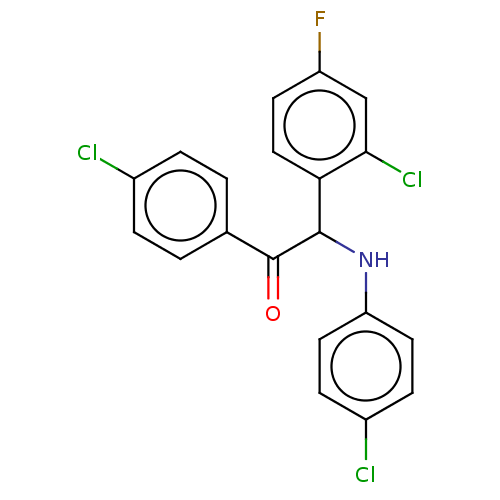
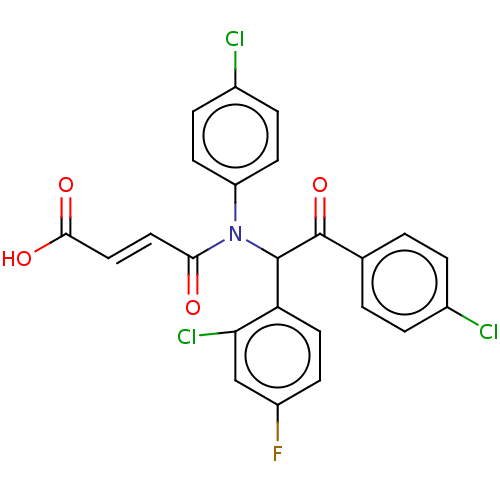
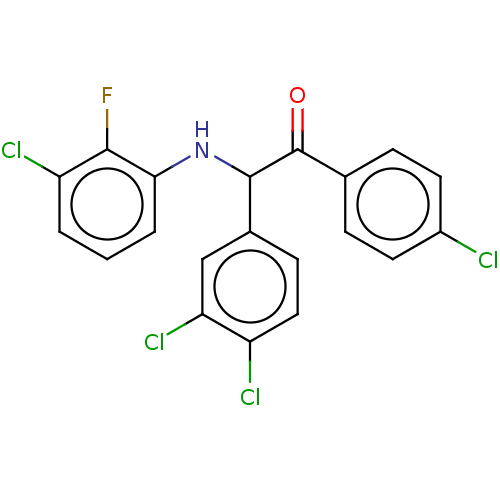
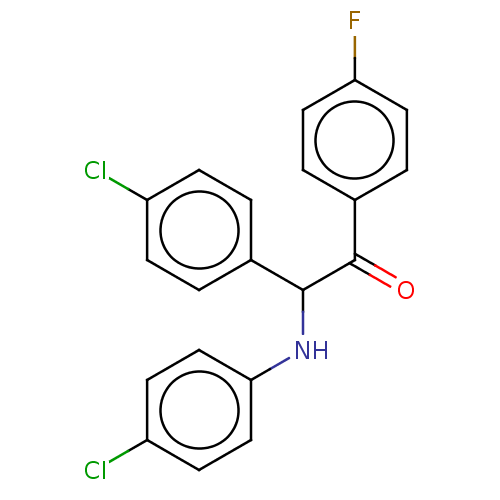
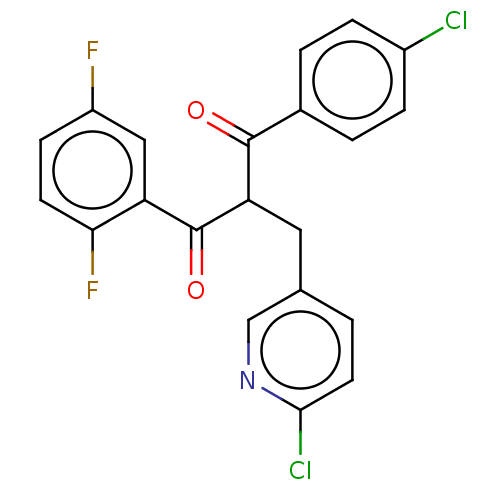
Affinity DataKi: 280nMAssay Description:Binding affinity to MDM2 (1 to 118 residues) (unknown origin) assessed as inhibition constant incubated for 1 hr measured by fluorescence polarizatio...More data for this Ligand-Target Pair
TargetE3 ubiquitin-protein ligase Mdm2(Human)
Nanjing University of Chinese Medicine
Curated by ChEMBL
Nanjing University of Chinese Medicine
Curated by ChEMBL
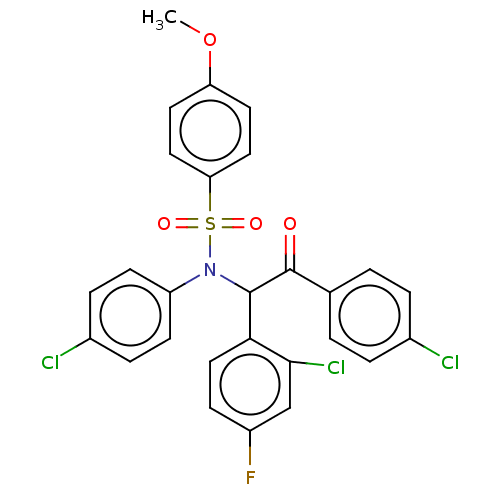
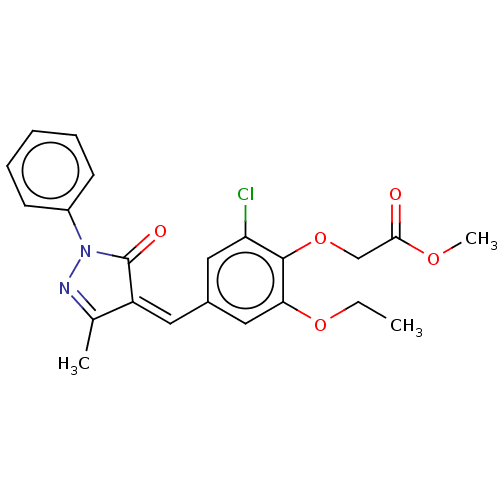
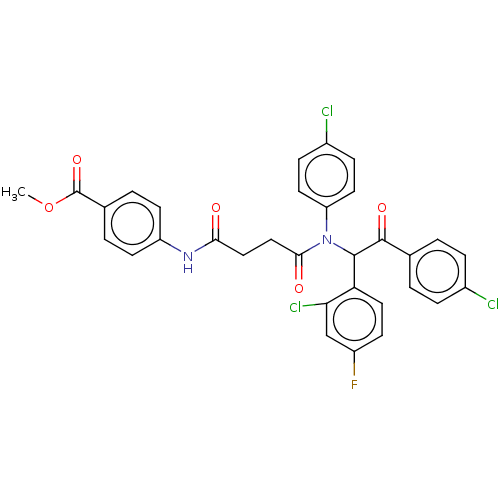
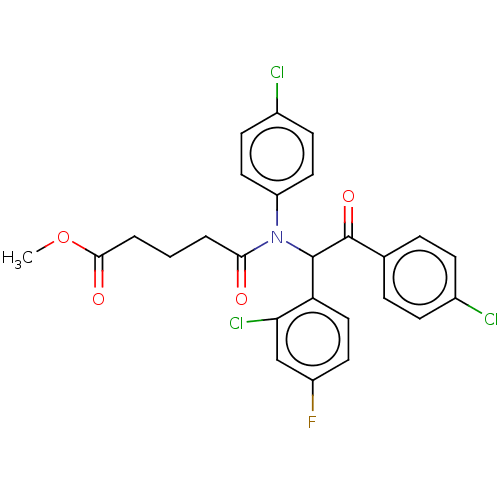
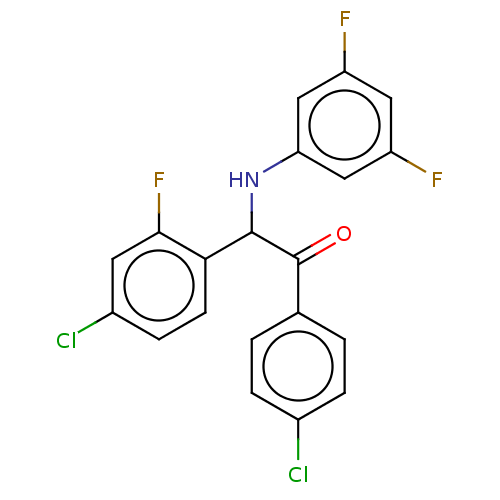
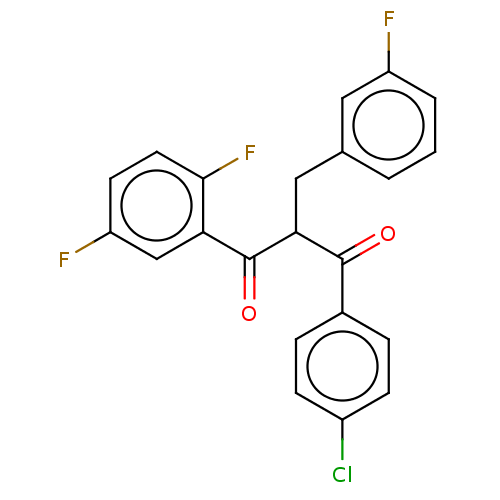
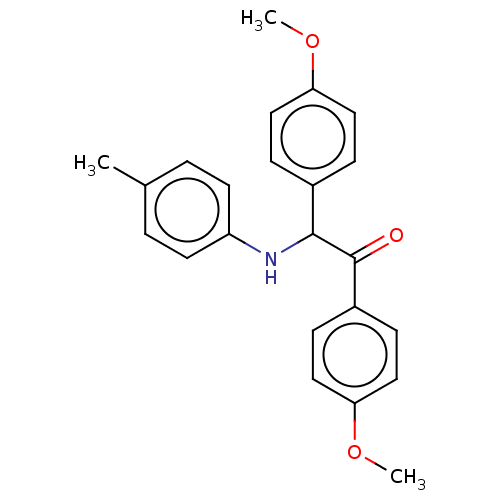
Affinity DataKi: 320nMAssay Description:Binding affinity to MDM2 (1 to 118 residues) (unknown origin) assessed as inhibition constant incubated for 1 hr measured by fluorescence polarizatio...More data for this Ligand-Target Pair
TargetE3 ubiquitin-protein ligase Mdm2(Human)
Nanjing University of Chinese Medicine
Curated by ChEMBL
Nanjing University of Chinese Medicine
Curated by ChEMBL
Affinity DataKi: 350nMAssay Description:Binding affinity to MDM2 (1 to 118 residues) (unknown origin) assessed as inhibition constant incubated for 1 hr measured by fluorescence polarizatio...More data for this Ligand-Target Pair
TargetE3 ubiquitin-protein ligase Mdm2(Human)
Nanjing University of Chinese Medicine
Curated by ChEMBL
Nanjing University of Chinese Medicine
Curated by ChEMBL
Affinity DataKi: 480nMAssay Description:Binding affinity to MDM2 (1 to 118 residues) (unknown origin) assessed as inhibition constant incubated for 1 hr measured by fluorescence polarizatio...More data for this Ligand-Target Pair
TargetE3 ubiquitin-protein ligase Mdm2(Human)
Nanjing University of Chinese Medicine
Curated by ChEMBL
Nanjing University of Chinese Medicine
Curated by ChEMBL
Affinity DataKi: 560nMAssay Description:Binding affinity to MDM2 (1 to 118 residues) (unknown origin) assessed as inhibition constant incubated for 1 hr measured by fluorescence polarizatio...More data for this Ligand-Target Pair
TargetE3 ubiquitin-protein ligase Mdm2(Human)
Nanjing University of Chinese Medicine
Curated by ChEMBL
Nanjing University of Chinese Medicine
Curated by ChEMBL
Affinity DataKi: 650nMAssay Description:Binding affinity to MDM2 (1 to 118 residues) (unknown origin) assessed as inhibition constant incubated for 1 hr measured by fluorescence polarizatio...More data for this Ligand-Target Pair
TargetE3 ubiquitin-protein ligase Mdm2(Human)
Nanjing University of Chinese Medicine
Curated by ChEMBL
Nanjing University of Chinese Medicine
Curated by ChEMBL
Affinity DataKi: 690nMAssay Description:Binding affinity to MDM2 (1 to 118 residues) (unknown origin) assessed as inhibition constant incubated for 1 hr measured by fluorescence polarizatio...More data for this Ligand-Target Pair
TargetE3 ubiquitin-protein ligase Mdm2(Human)
Nanjing University of Chinese Medicine
Curated by ChEMBL
Nanjing University of Chinese Medicine
Curated by ChEMBL
Affinity DataKi: 710nMAssay Description:Binding affinity to MDM2 (1 to 118 residues) (unknown origin) assessed as inhibition constant incubated for 1 hr measured by fluorescence polarizatio...More data for this Ligand-Target Pair
TargetE3 ubiquitin-protein ligase Mdm2(Human)
Nanjing University of Chinese Medicine
Curated by ChEMBL
Nanjing University of Chinese Medicine
Curated by ChEMBL
Affinity DataKi: 730nMAssay Description:Binding affinity to MDM2 (1 to 118 residues) (unknown origin) assessed as inhibition constant incubated for 1 hr measured by fluorescence polarizatio...More data for this Ligand-Target Pair
Affinity DataKi: 780nMAssay Description:Binding affinity to MDMX (1 to 118 residues) (unknown origin) assessed as inhibition constant incubated for 1 hr measured by fluorescence polarizatio...More data for this Ligand-Target Pair
TargetE3 ubiquitin-protein ligase Mdm2(Human)
Nanjing University of Chinese Medicine
Curated by ChEMBL
Nanjing University of Chinese Medicine
Curated by ChEMBL
Affinity DataKi: 780nMAssay Description:Binding affinity to MDM2 (1 to 118 residues) (unknown origin) assessed as inhibition constant incubated for 1 hr measured by fluorescence polarizatio...More data for this Ligand-Target Pair
TargetE3 ubiquitin-protein ligase Mdm2(Human)
Nanjing University of Chinese Medicine
Curated by ChEMBL
Nanjing University of Chinese Medicine
Curated by ChEMBL
Affinity DataKi: 820nMAssay Description:Binding affinity to MDM2 (1 to 118 residues) (unknown origin) assessed as inhibition constant incubated for 1 hr measured by fluorescence polarizatio...More data for this Ligand-Target Pair
TargetE3 ubiquitin-protein ligase Mdm2(Human)
Nanjing University of Chinese Medicine
Curated by ChEMBL
Nanjing University of Chinese Medicine
Curated by ChEMBL
Affinity DataKi: 910nMAssay Description:Binding affinity to MDM2 (1 to 118 residues) (unknown origin) assessed as inhibition constant incubated for 1 hr measured by fluorescence polarizatio...More data for this Ligand-Target Pair
TargetE3 ubiquitin-protein ligase Mdm2(Human)
Nanjing University of Chinese Medicine
Curated by ChEMBL
Nanjing University of Chinese Medicine
Curated by ChEMBL
Affinity DataKi: 960nMAssay Description:Binding affinity to MDM2 (1 to 118 residues) (unknown origin) assessed as inhibition constant incubated for 1 hr measured by fluorescence polarizatio...More data for this Ligand-Target Pair
TargetE3 ubiquitin-protein ligase Mdm2(Human)
Nanjing University of Chinese Medicine
Curated by ChEMBL
Nanjing University of Chinese Medicine
Curated by ChEMBL
Affinity DataKi: 1.06E+3nMAssay Description:Binding affinity to MDM2 (1 to 118 residues) (unknown origin) assessed as inhibition constant incubated for 1 hr measured by fluorescence polarizatio...More data for this Ligand-Target Pair
TargetE3 ubiquitin-protein ligase Mdm2(Human)
Nanjing University of Chinese Medicine
Curated by ChEMBL
Nanjing University of Chinese Medicine
Curated by ChEMBL
Affinity DataKi: 1.07E+3nMAssay Description:Binding affinity to MDM2 (1 to 118 residues) (unknown origin) assessed as inhibition constant incubated for 1 hr measured by fluorescence polarizatio...More data for this Ligand-Target Pair
TargetE3 ubiquitin-protein ligase Mdm2(Human)
Nanjing University of Chinese Medicine
Curated by ChEMBL
Nanjing University of Chinese Medicine
Curated by ChEMBL
Affinity DataKi: 1.10E+3nMAssay Description:Binding affinity to MDM2 (1 to 118 residues) (unknown origin) assessed as inhibition constant incubated for 1 hr measured by fluorescence polarizatio...More data for this Ligand-Target Pair
TargetE3 ubiquitin-protein ligase Mdm2(Human)
Nanjing University of Chinese Medicine
Curated by ChEMBL
Nanjing University of Chinese Medicine
Curated by ChEMBL
Affinity DataKi: 1.15E+3nMAssay Description:Binding affinity to MDM2 (1 to 118 residues) (unknown origin) assessed as inhibition constant incubated for 1 hr measured by fluorescence polarizatio...More data for this Ligand-Target Pair
TargetE3 ubiquitin-protein ligase Mdm2(Human)
Nanjing University of Chinese Medicine
Curated by ChEMBL
Nanjing University of Chinese Medicine
Curated by ChEMBL
Affinity DataKi: 1.39E+3nMAssay Description:Binding affinity to MDM2 (1 to 118 residues) (unknown origin) assessed as inhibition constant incubated for 1 hr measured by fluorescence polarizatio...More data for this Ligand-Target Pair
TargetE3 ubiquitin-protein ligase Mdm2(Human)
Nanjing University of Chinese Medicine
Curated by ChEMBL
Nanjing University of Chinese Medicine
Curated by ChEMBL
Affinity DataKi: 1.46E+3nMAssay Description:Binding affinity to MDM2 (1 to 118 residues) (unknown origin) assessed as inhibition constant incubated for 1 hr measured by fluorescence polarizatio...More data for this Ligand-Target Pair
TargetE3 ubiquitin-protein ligase Mdm2(Human)
Nanjing University of Chinese Medicine
Curated by ChEMBL
Nanjing University of Chinese Medicine
Curated by ChEMBL
Affinity DataKi: 2.13E+3nMAssay Description:Binding affinity to MDM2 (1 to 118 residues) (unknown origin) assessed as inhibition constant incubated for 1 hr measured by fluorescence polarizatio...More data for this Ligand-Target Pair
Affinity DataKi: 2.45E+3nMAssay Description:Binding affinity to MDMX (1 to 118 residues) (unknown origin) assessed as inhibition constant incubated for 1 hr measured by fluorescence polarizatio...More data for this Ligand-Target Pair
TargetE3 ubiquitin-protein ligase Mdm2(Human)
Nanjing University of Chinese Medicine
Curated by ChEMBL
Nanjing University of Chinese Medicine
Curated by ChEMBL
Affinity DataKi: 2.45E+3nMAssay Description:Binding affinity to MDM2 (1 to 118 residues) (unknown origin) assessed as inhibition constant incubated for 1 hr measured by fluorescence polarizatio...More data for this Ligand-Target Pair
Affinity DataKi: 3.46E+3nMAssay Description:Binding affinity to MDMX (1 to 118 residues) (unknown origin) assessed as inhibition constant incubated for 1 hr measured by fluorescence polarizatio...More data for this Ligand-Target Pair
Affinity DataKi: 4.22E+3nMAssay Description:Binding affinity to MDMX (1 to 118 residues) (unknown origin) assessed as inhibition constant incubated for 1 hr measured by fluorescence polarizatio...More data for this Ligand-Target Pair
TargetE3 ubiquitin-protein ligase Mdm2(Human)
Nanjing University of Chinese Medicine
Curated by ChEMBL
Nanjing University of Chinese Medicine
Curated by ChEMBL
Affinity DataKi: 4.29E+3nMAssay Description:Binding affinity to MDM2 (1 to 118 residues) (unknown origin) assessed as inhibition constant incubated for 1 hr measured by fluorescence polarizatio...More data for this Ligand-Target Pair
TargetE3 ubiquitin-protein ligase Mdm2(Human)
Nanjing University of Chinese Medicine
Curated by ChEMBL
Nanjing University of Chinese Medicine
Curated by ChEMBL
Affinity DataKi: 6.02E+3nMAssay Description:Binding affinity to MDM2 (1 to 118 residues) (unknown origin) assessed as inhibition constant incubated for 1 hr measured by fluorescence polarizatio...More data for this Ligand-Target Pair
TargetE3 ubiquitin-protein ligase Mdm2(Human)
Nanjing University of Chinese Medicine
Curated by ChEMBL
Nanjing University of Chinese Medicine
Curated by ChEMBL
Affinity DataKi: 6.86E+3nMAssay Description:Binding affinity to MDM2 (1 to 118 residues) (unknown origin) assessed as inhibition constant incubated for 1 hr measured by fluorescence polarizatio...More data for this Ligand-Target Pair
TargetE3 ubiquitin-protein ligase Mdm2(Human)
Nanjing University of Chinese Medicine
Curated by ChEMBL
Nanjing University of Chinese Medicine
Curated by ChEMBL
Affinity DataKi: 6.92E+3nMAssay Description:Binding affinity to MDM2 (1 to 118 residues) (unknown origin) assessed as inhibition constant incubated for 1 hr measured by fluorescence polarizatio...More data for this Ligand-Target Pair
TargetE3 ubiquitin-protein ligase Mdm2(Human)
Nanjing University of Chinese Medicine
Curated by ChEMBL
Nanjing University of Chinese Medicine
Curated by ChEMBL
Affinity DataKi: 7.34E+3nMAssay Description:Binding affinity to MDM2 (1 to 118 residues) (unknown origin) assessed as inhibition constant incubated for 1 hr measured by fluorescence polarizatio...More data for this Ligand-Target Pair
Affinity DataKi: 7.43E+3nMAssay Description:Binding affinity to MDMX (1 to 118 residues) (unknown origin) assessed as inhibition constant incubated for 1 hr measured by fluorescence polarizatio...More data for this Ligand-Target Pair
TargetE3 ubiquitin-protein ligase Mdm2(Human)
Nanjing University of Chinese Medicine
Curated by ChEMBL
Nanjing University of Chinese Medicine
Curated by ChEMBL
Affinity DataKi: 7.69E+3nMAssay Description:Binding affinity to MDM2 (1 to 118 residues) (unknown origin) assessed as inhibition constant incubated for 1 hr measured by fluorescence polarizatio...More data for this Ligand-Target Pair
Affinity DataKi: 9.37E+3nMAssay Description:Binding affinity to MDMX (1 to 118 residues) (unknown origin) assessed as inhibition constant incubated for 1 hr measured by fluorescence polarizatio...More data for this Ligand-Target Pair
Affinity DataKi: 1.16E+4nMAssay Description:Binding affinity to MDMX (1 to 118 residues) (unknown origin) assessed as inhibition constant incubated for 1 hr measured by fluorescence polarizatio...More data for this Ligand-Target Pair
TargetE3 ubiquitin-protein ligase Mdm2(Human)
Nanjing University of Chinese Medicine
Curated by ChEMBL
Nanjing University of Chinese Medicine
Curated by ChEMBL
Affinity DataKi: 1.23E+4nMAssay Description:Binding affinity to MDM2 (1 to 118 residues) (unknown origin) assessed as inhibition constant incubated for 1 hr measured by fluorescence polarizatio...More data for this Ligand-Target Pair
TargetE3 ubiquitin-protein ligase Mdm2(Human)
Nanjing University of Chinese Medicine
Curated by ChEMBL
Nanjing University of Chinese Medicine
Curated by ChEMBL
Affinity DataKi: 1.28E+4nMAssay Description:Binding affinity to MDM2 (1 to 118 residues) (unknown origin) assessed as inhibition constant incubated for 1 hr measured by fluorescence polarizatio...More data for this Ligand-Target Pair
Affinity DataKi: 1.29E+4nMAssay Description:Binding affinity to MDMX (1 to 118 residues) (unknown origin) assessed as inhibition constant incubated for 1 hr measured by fluorescence polarizatio...More data for this Ligand-Target Pair
Affinity DataKi: 1.30E+4nMAssay Description:Binding affinity to MDMX (1 to 118 residues) (unknown origin) assessed as inhibition constant incubated for 1 hr measured by fluorescence polarizatio...More data for this Ligand-Target Pair
TargetE3 ubiquitin-protein ligase Mdm2(Human)
Nanjing University of Chinese Medicine
Curated by ChEMBL
Nanjing University of Chinese Medicine
Curated by ChEMBL
Affinity DataKi: 1.32E+4nMAssay Description:Binding affinity to MDM2 (1 to 118 residues) (unknown origin) assessed as inhibition constant incubated for 1 hr measured by fluorescence polarizatio...More data for this Ligand-Target Pair
TargetE3 ubiquitin-protein ligase Mdm2(Human)
Nanjing University of Chinese Medicine
Curated by ChEMBL
Nanjing University of Chinese Medicine
Curated by ChEMBL
Affinity DataKi: 1.34E+4nMAssay Description:Binding affinity to MDM2 (1 to 118 residues) (unknown origin) assessed as inhibition constant incubated for 1 hr measured by fluorescence polarizatio...More data for this Ligand-Target Pair
TargetE3 ubiquitin-protein ligase Mdm2(Human)
Nanjing University of Chinese Medicine
Curated by ChEMBL
Nanjing University of Chinese Medicine
Curated by ChEMBL
Affinity DataKi: 1.40E+4nMAssay Description:Binding affinity to MDM2 (1 to 118 residues) (unknown origin) assessed as inhibition constant incubated for 1 hr measured by fluorescence polarizatio...More data for this Ligand-Target Pair
TargetE3 ubiquitin-protein ligase Mdm2(Human)
Nanjing University of Chinese Medicine
Curated by ChEMBL
Nanjing University of Chinese Medicine
Curated by ChEMBL
Affinity DataKi: 1.46E+4nMAssay Description:Binding affinity to MDM2 (1 to 118 residues) (unknown origin) assessed as inhibition constant incubated for 1 hr measured by fluorescence polarizatio...More data for this Ligand-Target Pair
TargetE3 ubiquitin-protein ligase Mdm2(Human)
Nanjing University of Chinese Medicine
Curated by ChEMBL
Nanjing University of Chinese Medicine
Curated by ChEMBL
Affinity DataKi: 1.49E+4nMAssay Description:Binding affinity to MDM2 (1 to 118 residues) (unknown origin) assessed as inhibition constant incubated for 1 hr measured by fluorescence polarizatio...More data for this Ligand-Target Pair
TargetE3 ubiquitin-protein ligase Mdm2(Human)
Nanjing University of Chinese Medicine
Curated by ChEMBL
Nanjing University of Chinese Medicine
Curated by ChEMBL
Affinity DataKi: 1.51E+4nMAssay Description:Binding affinity to MDM2 (1 to 118 residues) (unknown origin) assessed as inhibition constant incubated for 1 hr measured by fluorescence polarizatio...More data for this Ligand-Target Pair
TargetE3 ubiquitin-protein ligase Mdm2(Human)
Nanjing University of Chinese Medicine
Curated by ChEMBL
Nanjing University of Chinese Medicine
Curated by ChEMBL
Affinity DataKi: 1.53E+4nMAssay Description:Binding affinity to MDM2 (1 to 118 residues) (unknown origin) assessed as inhibition constant incubated for 1 hr measured by fluorescence polarizatio...More data for this Ligand-Target Pair
Affinity DataKi: 1.56E+4nMAssay Description:Binding affinity to MDMX (1 to 118 residues) (unknown origin) assessed as inhibition constant incubated for 1 hr measured by fluorescence polarizatio...More data for this Ligand-Target Pair
TargetE3 ubiquitin-protein ligase Mdm2(Human)
Nanjing University of Chinese Medicine
Curated by ChEMBL
Nanjing University of Chinese Medicine
Curated by ChEMBL
Affinity DataKi: 1.65E+4nMAssay Description:Binding affinity to MDM2 (1 to 118 residues) (unknown origin) assessed as inhibition constant incubated for 1 hr measured by fluorescence polarizatio...More data for this Ligand-Target Pair
TargetE3 ubiquitin-protein ligase Mdm2(Human)
Nanjing University of Chinese Medicine
Curated by ChEMBL
Nanjing University of Chinese Medicine
Curated by ChEMBL
Affinity DataKi: 1.74E+4nMAssay Description:Binding affinity to MDM2 (1 to 118 residues) (unknown origin) assessed as inhibition constant incubated for 1 hr measured by fluorescence polarizatio...More data for this Ligand-Target Pair