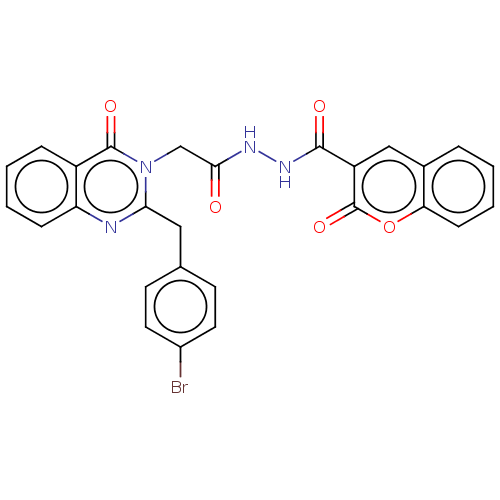
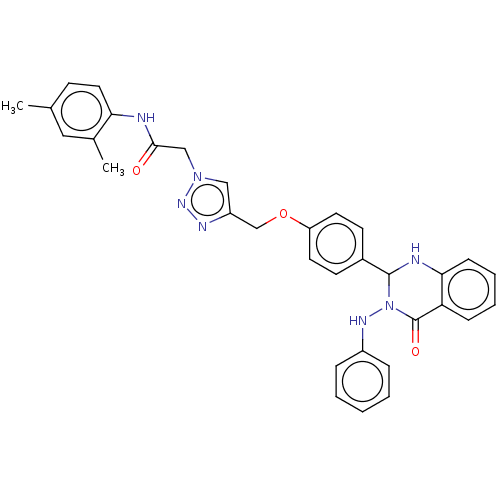
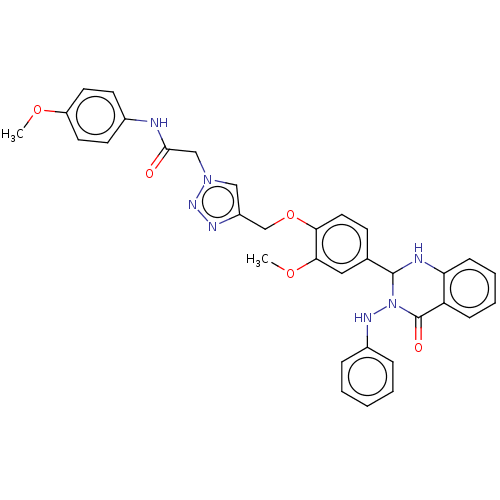
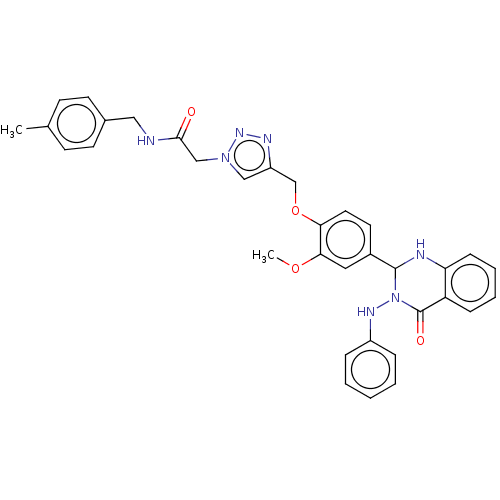
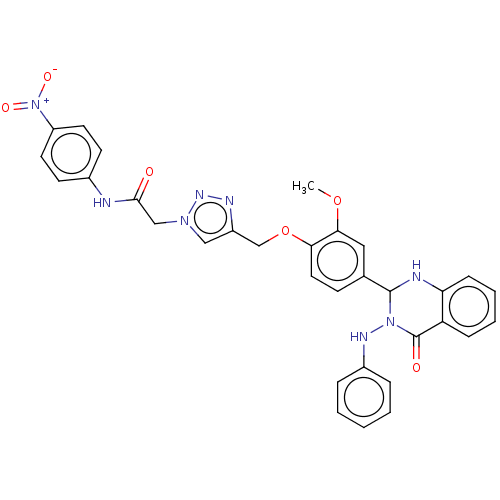
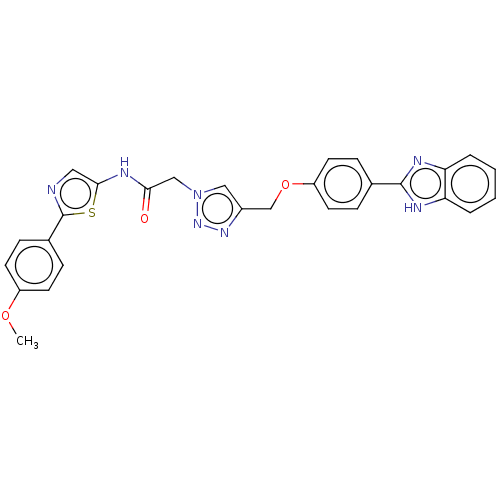
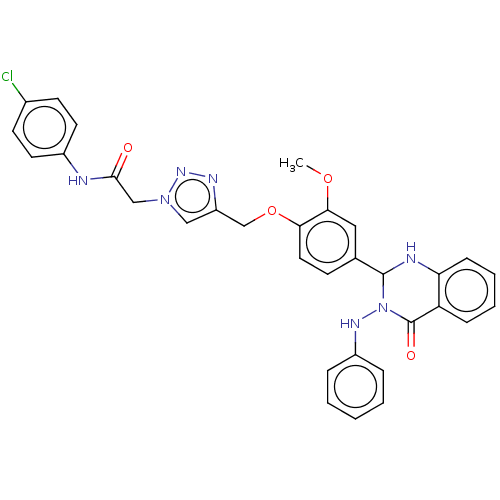
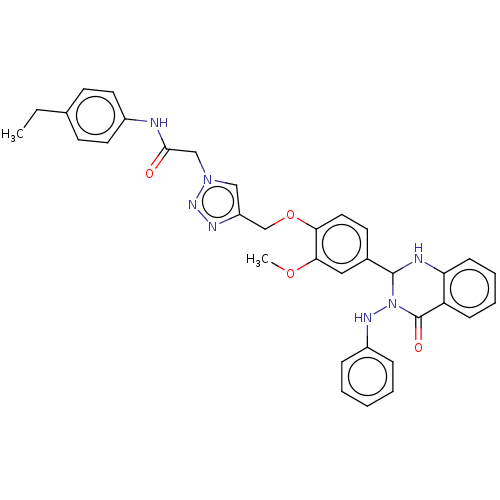
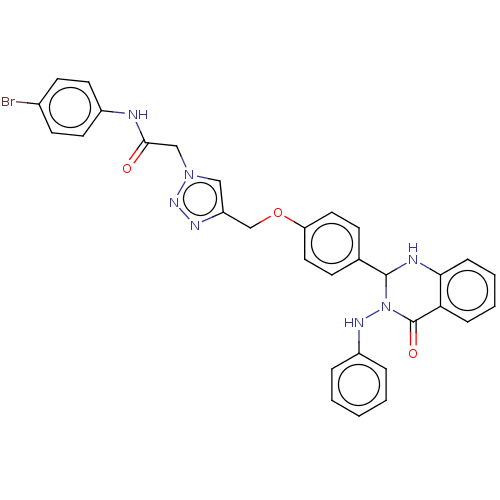
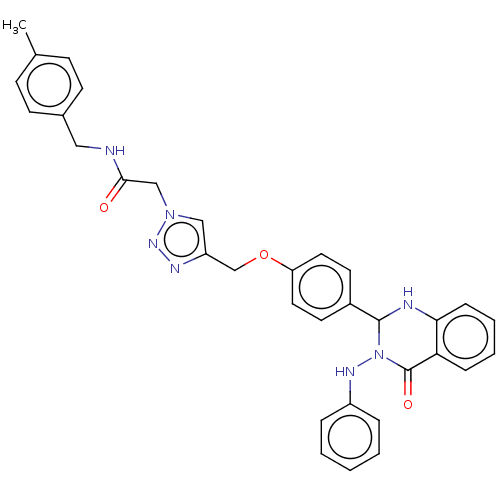
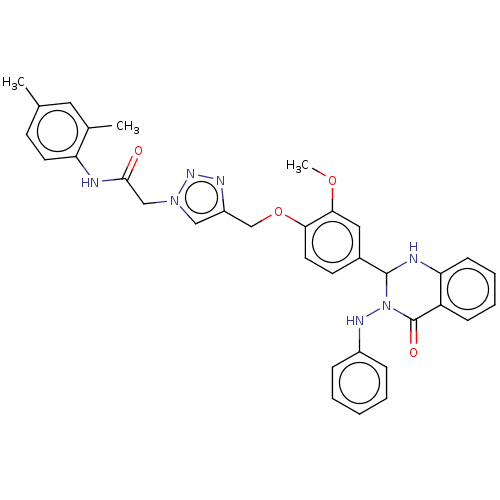
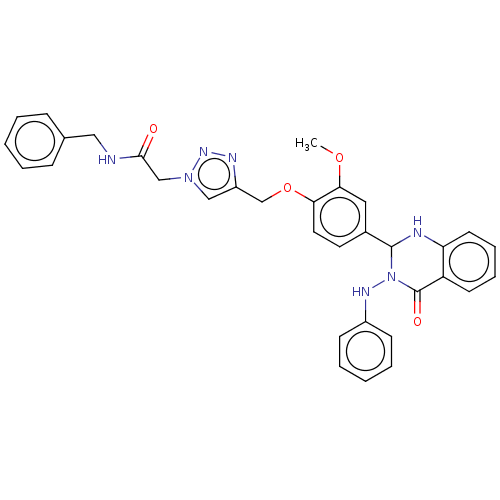
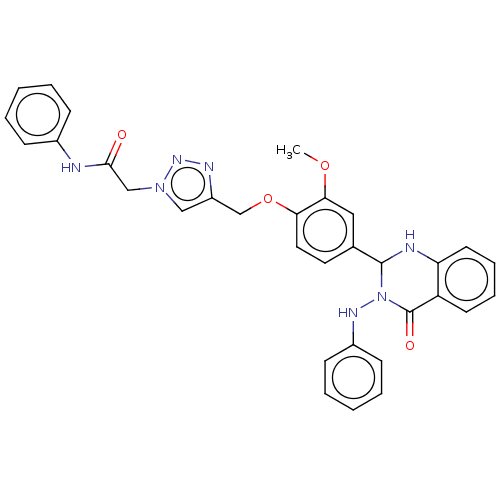
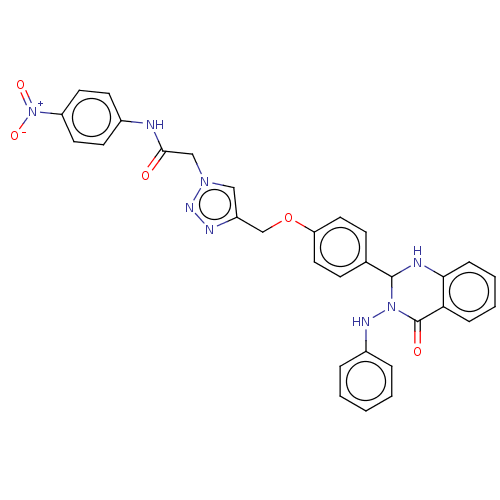
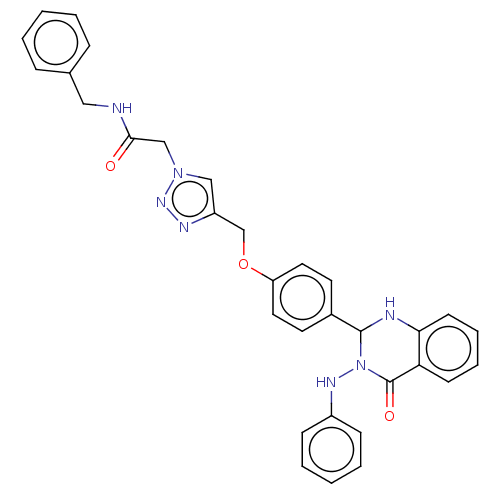
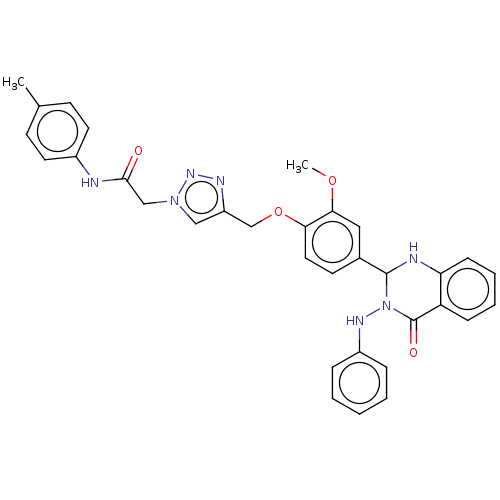
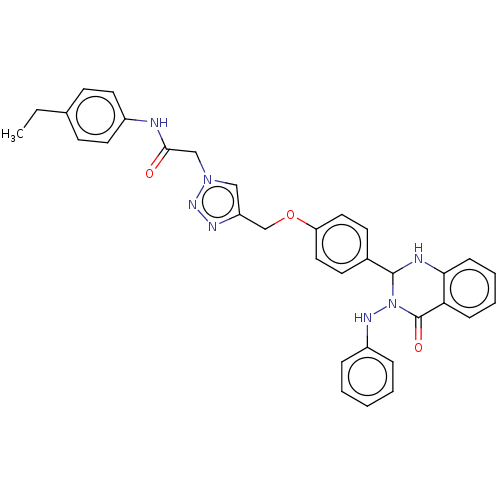
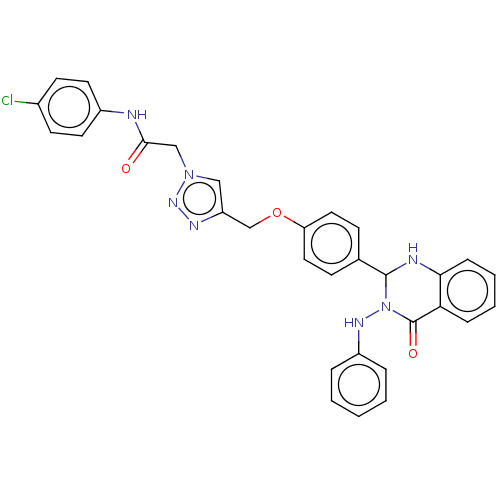
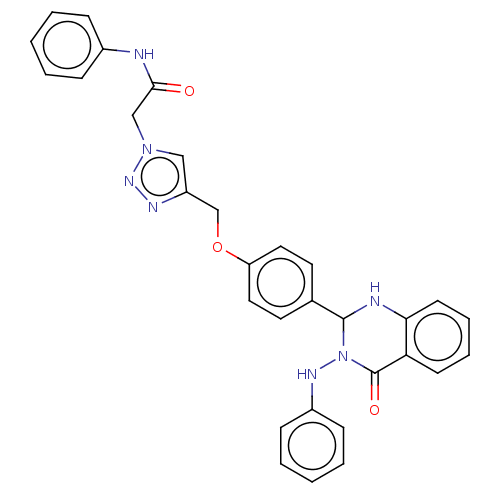
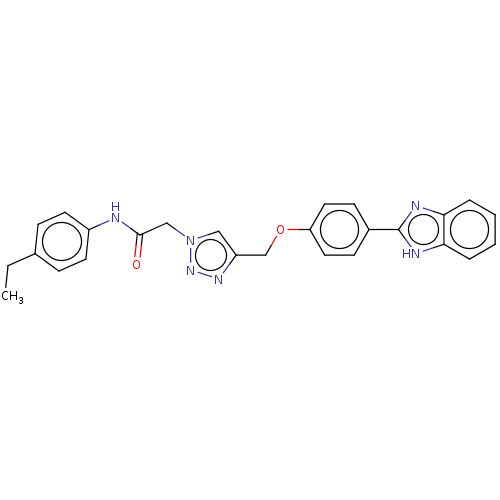
Report error Found 29 Enz. Inhib. hit(s) with all data for entry = 50018945
TargetPancreatic triacylglycerol lipase(Human)
Tehran University of Medical Sciences Tehran Iran
Curated by ChEMBL
Tehran University of Medical Sciences Tehran Iran
Curated by ChEMBL
Affinity DataIC50: 2.80E+3nMAssay Description:Inhibition of pancreatic lipase (unknown origin)More data for this Ligand-Target Pair
Affinity DataIC50: 4.80E+3nMAssay Description:Inhibition of alpha-glucosidase (unknown origin) using p-nitrophenyl glucopyranoside as substrate preincubated for 10 mins followed by substrate addi...More data for this Ligand-Target Pair
Affinity DataKi: 4.80E+3nMAssay Description:Inhibition of alpha-glucosidase (unknown origin) assessed as inhibition constant using p-nitrophenyl-alpha D-glucopyranoside as substrate by Lineweav...More data for this Ligand-Target Pair
Affinity DataIC50: 5.00E+3nMAssay Description:Inhibition of alpha-glucosidase (unknown origin) using p-nitrophenyl glucopyranoside as substrate preincubated for 10 mins followed by substrate addi...More data for this Ligand-Target Pair
Affinity DataIC50: 5.40E+3nMAssay Description:Inhibition of alpha-glucosidase (unknown origin) using p-nitrophenyl glucopyranoside as substrate preincubated for 10 mins followed by substrate addi...More data for this Ligand-Target Pair
Affinity DataIC50: 6.00E+3nMAssay Description:Inhibition of alpha-glucosidase (unknown origin) using p-nitrophenyl glucopyranoside as substrate preincubated for 10 mins followed by substrate addi...More data for this Ligand-Target Pair
Affinity DataIC50: 6.31E+3nMAssay Description:Inhibition of alpha-glucosidase (unknown origin) using p-nitrophenyl glucopyranoside as substrate preincubated for 10 mins followed by substrate addi...More data for this Ligand-Target Pair
Affinity DataIC50: 6.70E+3nMAssay Description:Inhibition of alpha-glucosidase (unknown origin) using p-nitrophenyl glucopyranoside as substrate preincubated for 10 mins followed by substrate addi...More data for this Ligand-Target Pair
Affinity DataIC50: 7.20E+3nMAssay Description:Inhibition of alpha-glucosidase (unknown origin) using p-nitrophenyl glucopyranoside as substrate preincubated for 10 mins followed by substrate addi...More data for this Ligand-Target Pair
Affinity DataIC50: 7.30E+3nMAssay Description:Inhibition of alpha-glucosidase (unknown origin)More data for this Ligand-Target Pair
Affinity DataIC50: 8.00E+3nMAssay Description:Inhibition of alpha-glucosidase (unknown origin) using p-nitrophenyl glucopyranoside as substrate preincubated for 10 mins followed by substrate addi...More data for this Ligand-Target Pair
Affinity DataIC50: 8.30E+3nMAssay Description:Inhibition of alpha-glucosidase (unknown origin) using p-nitrophenyl glucopyranoside as substrate preincubated for 10 mins followed by substrate addi...More data for this Ligand-Target Pair
Affinity DataIC50: 1.06E+4nMAssay Description:Inhibition of alpha-glucosidase (unknown origin) using p-nitrophenyl glucopyranoside as substrate preincubated for 10 mins followed by substrate addi...More data for this Ligand-Target Pair
Affinity DataIC50: 1.16E+4nMAssay Description:Inhibition of alpha-glucosidase (unknown origin) using p-nitrophenyl glucopyranoside as substrate preincubated for 10 mins followed by substrate addi...More data for this Ligand-Target Pair
Affinity DataIC50: 1.17E+4nMAssay Description:Inhibition of alpha-glucosidase (unknown origin) using p-nitrophenyl glucopyranoside as substrate preincubated for 10 mins followed by substrate addi...More data for this Ligand-Target Pair
Affinity DataIC50: 1.20E+4nMAssay Description:Inhibition of alpha-glucosidase (unknown origin) using p-nitrophenyl glucopyranoside as substrate preincubated for 10 mins followed by substrate addi...More data for this Ligand-Target Pair
Affinity DataIC50: 1.20E+4nMAssay Description:Inhibition of alpha-glucosidase (unknown origin) using p-nitrophenyl glucopyranoside as substrate preincubated for 10 mins followed by substrate addi...More data for this Ligand-Target Pair
Affinity DataIC50: 1.30E+4nMAssay Description:Inhibition of alpha-glucosidase (unknown origin) using p-nitrophenyl glucopyranoside as substrate preincubated for 10 mins followed by substrate addi...More data for this Ligand-Target Pair
Affinity DataIC50: 1.47E+4nMAssay Description:Inhibition of alpha-glucosidase (unknown origin) using p-nitrophenyl glucopyranoside as substrate preincubated for 10 mins followed by substrate addi...More data for this Ligand-Target Pair
Affinity DataIC50: 1.53E+4nMAssay Description:Inhibition of alpha-glucosidase (unknown origin) using p-nitrophenyl glucopyranoside as substrate preincubated for 10 mins followed by substrate addi...More data for this Ligand-Target Pair
Affinity DataIC50: 1.66E+4nMAssay Description:Inhibition of alpha-glucosidase (unknown origin) using p-nitrophenyl glucopyranoside as substrate preincubated for 10 mins followed by substrate addi...More data for this Ligand-Target Pair
Affinity DataIC50: 2.52E+4nMAssay Description:Inhibition of alpha-glucosidase (unknown origin) using p-nitrophenyl glucopyranoside as substrate preincubated for 10 mins followed by substrate addi...More data for this Ligand-Target Pair
Affinity DataIC50: 2.61E+4nMAssay Description:Inhibition of alpha-glucosidase (unknown origin)More data for this Ligand-Target Pair
Affinity DataIC50: 4.26E+4nMAssay Description:Inhibition of alpha-glucosidase (unknown origin) using p-nitrophenyl glucopyranoside as substrate preincubated for 10 mins followed by substrate addi...More data for this Ligand-Target Pair
TargetPlatelet-derived growth factor receptor alpha/beta(Mouse)
Tehran University of Medical Sciences Tehran Iran
Curated by ChEMBL
Tehran University of Medical Sciences Tehran Iran
Curated by ChEMBL
Affinity DataKi: 5.60E+4nMAssay Description:Inhibition of Saccharomyces cerevisiae alpha-glucosidase assessed as inhibition constant using p-nitrophenyl-alpha D-glucopyranoside as substrate pre...More data for this Ligand-Target Pair
TargetPlatelet-derived growth factor receptor alpha/beta(Mouse)
Tehran University of Medical Sciences Tehran Iran
Curated by ChEMBL
Tehran University of Medical Sciences Tehran Iran
Curated by ChEMBL
Affinity DataIC50: 6.05E+4nMAssay Description:Inhibition of Saccharomyces cerevisiae alpha-glucosidase using p-nitrophenyl-alpha D-glucopyranoside as substrate preincubated for 15 mins followed b...More data for this Ligand-Target Pair
Affinity DataIC50: 9.56E+4nMAssay Description:Inhibition of alpha-glucosidase (unknown origin) using p-nitrophenyl glucopyranoside as substrate preincubated for 10 mins followed by substrate addi...More data for this Ligand-Target Pair
Affinity DataIC50: 1.40E+5nMAssay Description:Inhibition of alpha-glucosidase (unknown origin) using p-nitrophenyl glucopyranoside as substrate preincubated for 10 mins followed by substrate addi...More data for this Ligand-Target Pair
Affinity DataIC50: 7.50E+5nMAssay Description:Inhibition of alpha-glucosidase (unknown origin) using p-nitrophenyl glucopyranoside as substrate preincubated for 10 mins followed by substrate addi...More data for this Ligand-Target Pair