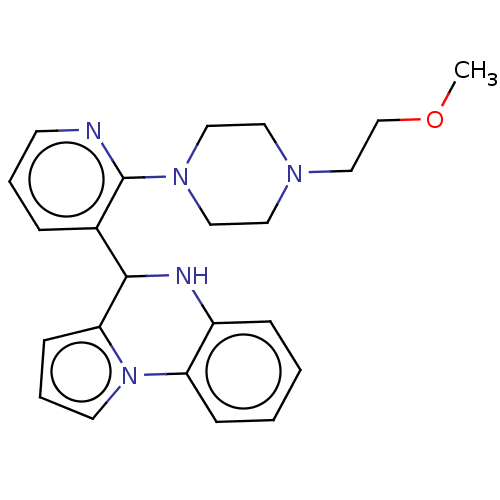
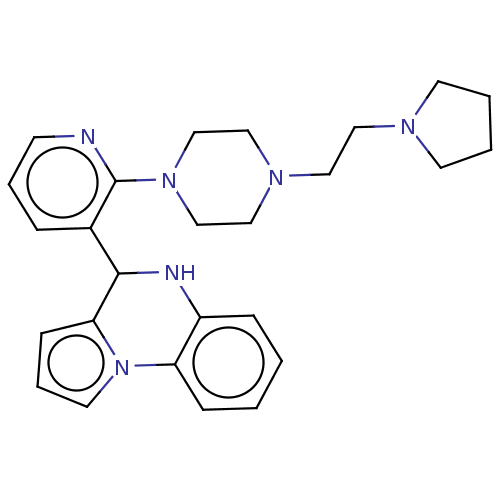
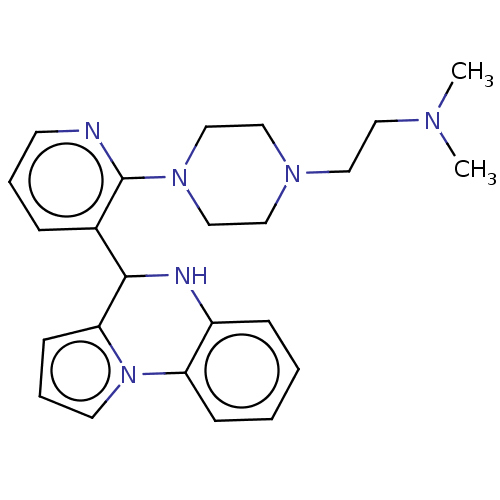
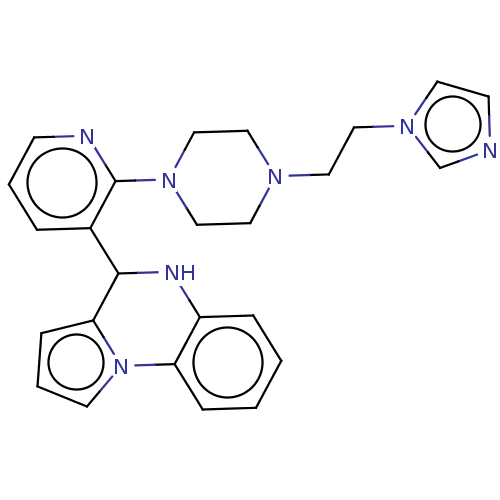
Report error Found 12 Enz. Inhib. hit(s) with all data for entry = 50019967
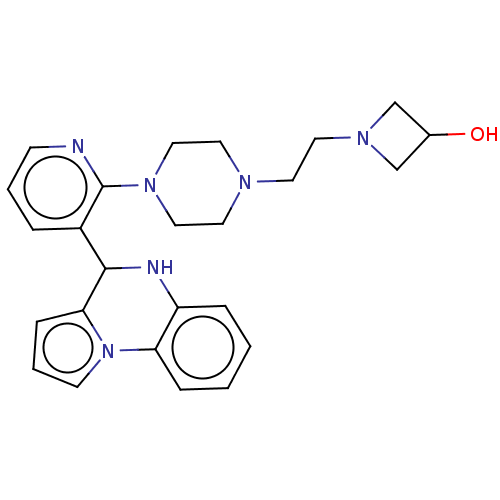
TargetNAD-dependent protein deacylase sirtuin-6(Human)
University of Texas Medical Branch
Curated by ChEMBL
University of Texas Medical Branch
Curated by ChEMBL
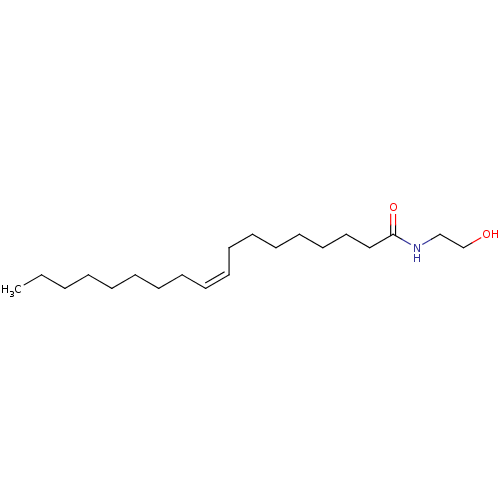
Affinity DataEC50: 3.10E+3nMAssay Description:Activation of his6-tagged SIRT6 (unknown origin) using H3K9Ac peptide substrate incubated for 30 minsMore data for this Ligand-Target Pair
TargetNAD-dependent protein deacylase sirtuin-6(Human)
University of Texas Medical Branch
Curated by ChEMBL
University of Texas Medical Branch
Curated by ChEMBL
Affinity DataEC50: 5.30E+3nMAssay Description:Activation of SIRT6 (unknown origin) deacetylase activityMore data for this Ligand-Target Pair
TargetNAD-dependent protein deacylase sirtuin-6(Human)
University of Texas Medical Branch
Curated by ChEMBL
University of Texas Medical Branch
Curated by ChEMBL
Affinity DataEC50: 1.03E+4nMAssay Description:Activation of SIRT6 (unknown origin) deacetylase activityMore data for this Ligand-Target Pair
TargetNAD-dependent protein deacylase sirtuin-6(Human)
University of Texas Medical Branch
Curated by ChEMBL
University of Texas Medical Branch
Curated by ChEMBL
Affinity DataEC50: 3.80E+4nMAssay Description:Activation of SIRT6 (unknown origin) deacetylase activityMore data for this Ligand-Target Pair
TargetNAD-dependent protein deacylase sirtuin-6(Human)
University of Texas Medical Branch
Curated by ChEMBL
University of Texas Medical Branch
Curated by ChEMBL
Affinity DataEC50: 3.88E+4nMAssay Description:Activation of recombinant SIRT6 (unknown origin) deacetylase activity using fluoro peptide as substrate by microplate reader analysisMore data for this Ligand-Target Pair
Ligand InfoSimilars
TargetNAD-dependent protein deacylase sirtuin-6(Human)
University of Texas Medical Branch
Curated by ChEMBL
University of Texas Medical Branch
Curated by ChEMBL
Affinity DataEC50: 4.86E+4nMAssay Description:Activation of recombinant SIRT6 (unknown origin) deacetylase activity using fluoro peptide as substrate by microplate reader analysisMore data for this Ligand-Target Pair
Ligand InfoSimilars
TargetNAD-dependent protein deacylase sirtuin-6(Human)
University of Texas Medical Branch
Curated by ChEMBL
University of Texas Medical Branch
Curated by ChEMBL
Affinity DataEC50: 4.88E+4nMAssay Description:Activation of recombinant SIRT6 (unknown origin) deacetylase activity using fluoro peptide as substrate by microplate reader analysisMore data for this Ligand-Target Pair
Ligand InfoSimilars
TargetNAD-dependent protein deacylase sirtuin-6(Human)
University of Texas Medical Branch
Curated by ChEMBL
University of Texas Medical Branch
Curated by ChEMBL
Affinity DataEC50: 5.25E+4nMAssay Description:Activation of recombinant SIRT6 (unknown origin) deacetylase activity using fluoro peptide as substrate by microplate reader analysisMore data for this Ligand-Target Pair
Ligand InfoSimilars
TargetNAD-dependent protein deacylase sirtuin-6(Human)
University of Texas Medical Branch
Curated by ChEMBL
University of Texas Medical Branch
Curated by ChEMBL
Affinity DataEC50: 5.80E+4nMAssay Description:Activation of recombinant SIRT6 (unknown origin) deacetylase activity using fluoro peptide as substrate by microplate reader analysisMore data for this Ligand-Target Pair
Ligand InfoSimilars
TargetNAD-dependent protein deacylase sirtuin-6(Human)
University of Texas Medical Branch
Curated by ChEMBL
University of Texas Medical Branch
Curated by ChEMBL
Affinity DataEC50: 6.47E+4nMAssay Description:Activation of recombinant SIRT6 (unknown origin) deacetylase activity using fluoro peptide as substrate by microplate reader analysisMore data for this Ligand-Target Pair
Ligand InfoSimilars
TargetNAD-dependent protein deacylase sirtuin-6(Human)
University of Texas Medical Branch
Curated by ChEMBL
University of Texas Medical Branch
Curated by ChEMBL
Affinity DataEC50: 2.46E+5nMAssay Description:Activation of SIRT6 (unknown origin) deacetylase activity using H3K9Ac peptide substrate by reversed-phase HPLCMore data for this Ligand-Target Pair
TargetNAD-dependent protein deacylase sirtuin-6(Human)
University of Texas Medical Branch
Curated by ChEMBL
University of Texas Medical Branch
Curated by ChEMBL
Affinity DataEC50: 4.60E+5nMAssay Description:Activation of human SIRT6 deacetylase activity by FDL assayMore data for this Ligand-Target Pair