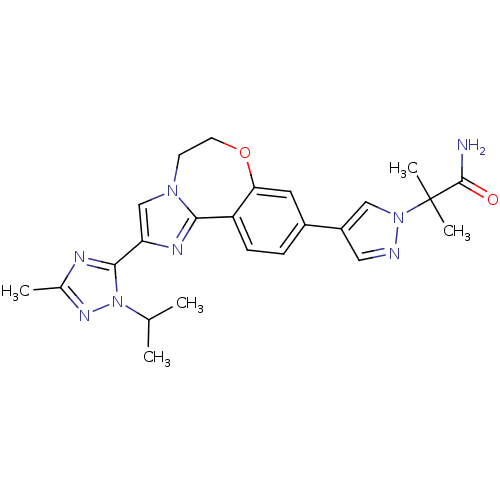
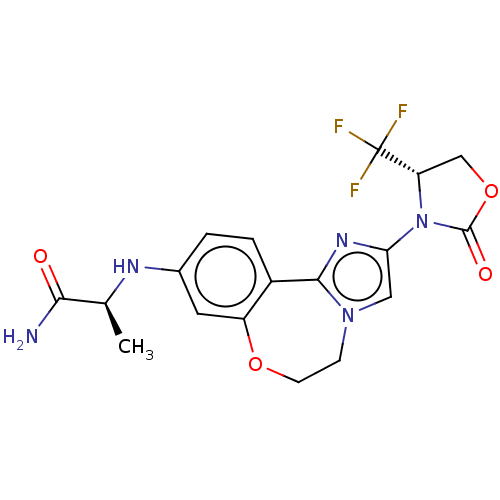
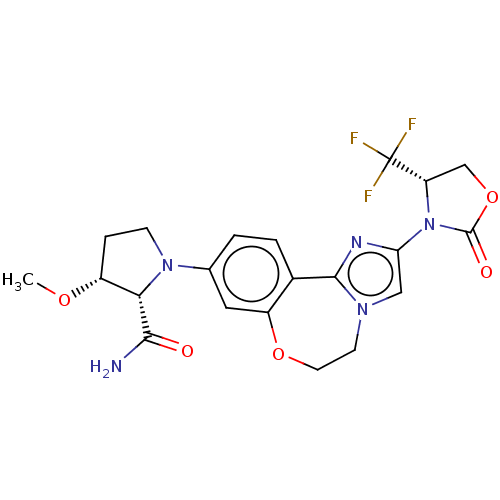
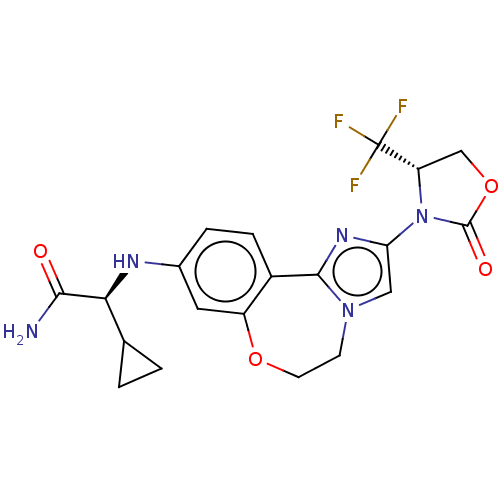
Report error Found 59 Enz. Inhib. hit(s) with all data for entry = 50017491
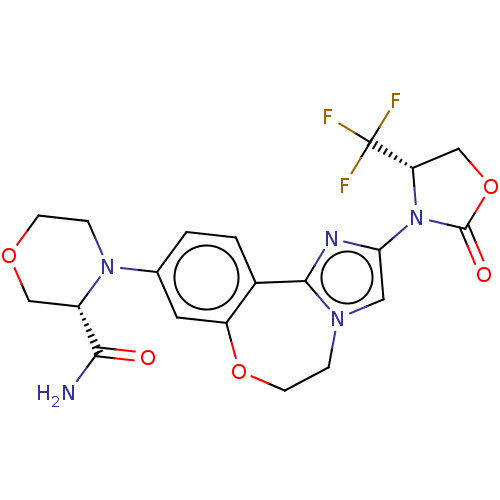
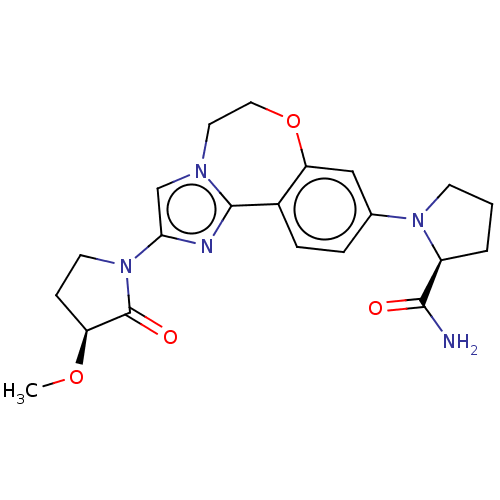
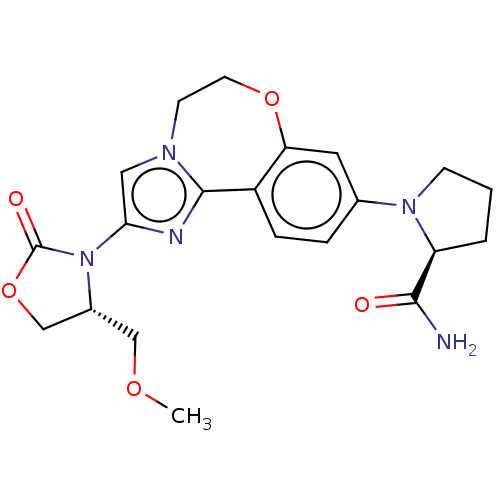
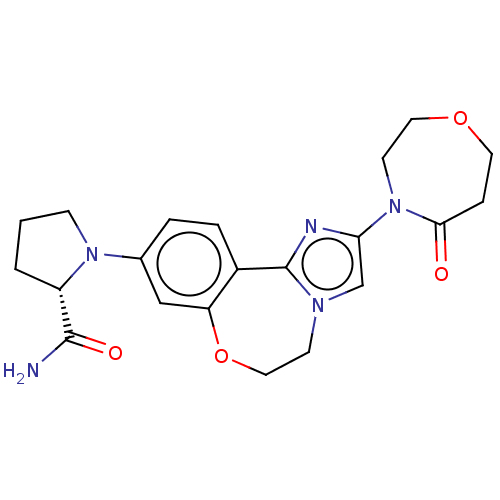
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit alpha isoform(Human)
Genentech
Curated by ChEMBL
Genentech
Curated by ChEMBL
Affinity DataKi: 0.0260nMAssay Description:Inhibition of PI3Kalpha (unknown origin) using PIP2:3PS as substrate in presence of ATP measured after 120 mins by ADP-Glo assayMore data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit alpha isoform(Human)
Genentech
Curated by ChEMBL
Genentech
Curated by ChEMBL
Affinity DataKi: 0.0340nMAssay Description:Inhibition of PI3Kalpha (unknown origin) using PIP2:3PS as substrate in presence of ATP measured after 120 mins by ADP-Glo assayMore data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit alpha isoform(Human)
Genentech
Curated by ChEMBL
Genentech
Curated by ChEMBL
Affinity DataKi: 0.0420nMAssay Description:Inhibition of PI3Kalpha (unknown origin) using PIP2:3PS as substrate in presence of ATP measured after 120 mins by ADP-Glo assayMore data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit alpha isoform(Human)
Genentech
Curated by ChEMBL
Genentech
Curated by ChEMBL
Affinity DataKi: 0.0430nMAssay Description:Inhibition of PI3Kalpha (unknown origin) using PIP2:3PS as substrate in presence of ATP measured after 120 mins by ADP-Glo assayMore data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit alpha isoform(Human)
Genentech
Curated by ChEMBL
Genentech
Curated by ChEMBL
Affinity DataKi: 0.0510nMAssay Description:Inhibition of PI3Kalpha (unknown origin) using PIP2:3PS as substrate in presence of ATP measured after 120 mins by ADP-Glo assayMore data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit alpha isoform(Human)
Genentech
Curated by ChEMBL
Genentech
Curated by ChEMBL
Affinity DataKi: 0.0530nMAssay Description:Inhibition of PI3Kalpha (unknown origin) using PIP2:3PS as substrate in presence of ATP measured after 120 mins by ADP-Glo assayMore data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit alpha isoform(Human)
Genentech
Curated by ChEMBL
Genentech
Curated by ChEMBL
Affinity DataKi: 0.0600nMAssay Description:Inhibition of PI3Kalpha (unknown origin) using PIP2:3PS as substrate in presence of ATP measured after 120 mins by ADP-Glo assayMore data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit alpha isoform(Human)
Genentech
Curated by ChEMBL
Genentech
Curated by ChEMBL
Affinity DataKi: 0.0620nMAssay Description:Inhibition of PI3Kalpha (unknown origin) using PIP2:3PS as substrate in presence of ATP measured after 120 mins by ADP-Glo assayMore data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit alpha isoform(Human)
Genentech
Curated by ChEMBL
Genentech
Curated by ChEMBL
Affinity DataKi: 0.0900nMAssay Description:Inhibition of PI3Kalpha (unknown origin) using PIP2:3PS as substrate in presence of ATP measured after 120 mins by ADP-Glo assayMore data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit alpha isoform(Human)
Genentech
Curated by ChEMBL
Genentech
Curated by ChEMBL
Affinity DataKi: 0.0950nMAssay Description:Inhibition of PI3Kalpha (unknown origin) using PIP2:3PS as substrate in presence of ATP measured after 120 mins by ADP-Glo assayMore data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit alpha isoform(Human)
Genentech
Curated by ChEMBL
Genentech
Curated by ChEMBL
Affinity DataKi: 0.0970nMAssay Description:Inhibition of PI3Kalpha (unknown origin) using PIP2:3PS as substrate in presence of ATP measured after 120 mins by ADP-Glo assayMore data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit alpha isoform(Human)
Genentech
Curated by ChEMBL
Genentech
Curated by ChEMBL
Affinity DataKi: 0.100nMAssay Description:Inhibition of PI3Kalpha (unknown origin) using PIP2:3PS as substrate in presence of ATP measured after 120 mins by ADP-Glo assayMore data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit alpha isoform(Human)
Genentech
Curated by ChEMBL
Genentech
Curated by ChEMBL
Affinity DataKi: 0.107nMAssay Description:Inhibition of PI3Kalpha (unknown origin) using PIP2:3PS as substrate in presence of ATP measured after 120 mins by ADP-Glo assayMore data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit alpha isoform(Human)
Genentech
Curated by ChEMBL
Genentech
Curated by ChEMBL
Affinity DataKi: 0.150nMAssay Description:Inhibition of PI3Kalpha (unknown origin) using PIP2:3PS as substrate in presence of ATP measured after 120 mins by ADP-Glo assayMore data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit alpha isoform(Human)
Genentech
Curated by ChEMBL
Genentech
Curated by ChEMBL
Affinity DataKi: 0.157nMAssay Description:Inhibition of PI3Kalpha (unknown origin) using PIP2:3PS as substrate in presence of ATP measured after 120 mins by ADP-Glo assayMore data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit alpha isoform(Human)
Genentech
Curated by ChEMBL
Genentech
Curated by ChEMBL
Affinity DataKi: 0.188nMAssay Description:Inhibition of PI3Kalpha (unknown origin) using PIP2:3PS as substrate in presence of ATP measured after 120 mins by ADP-Glo assayMore data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit alpha isoform(Human)
Genentech
Curated by ChEMBL
Genentech
Curated by ChEMBL
Affinity DataKi: 0.289nMAssay Description:Inhibition of PI3Kalpha (unknown origin) using PIP2:3PS as substrate in presence of ATP measured after 120 mins by ADP-Glo assayMore data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit alpha isoform(Human)
Genentech
Curated by ChEMBL
Genentech
Curated by ChEMBL
Affinity DataKi: 0.320nMAssay Description:Inhibition of PI3Kalpha (unknown origin) using PIP2:3PS as substrate in presence of ATP measured after 120 mins by ADP-Glo assayMore data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit alpha isoform(Human)
Genentech
Curated by ChEMBL
Genentech
Curated by ChEMBL
Affinity DataKi: 0.323nMAssay Description:Inhibition of PI3Kalpha (unknown origin) using PIP2:3PS as substrate in presence of ATP measured after 120 mins by ADP-Glo assayMore data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit alpha isoform(Human)
Genentech
Curated by ChEMBL
Genentech
Curated by ChEMBL
Affinity DataKi: 0.346nMAssay Description:Inhibition of PI3Kalpha (unknown origin) using PIP2:3PS as substrate in presence of ATP measured after 120 mins by ADP-Glo assayMore data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit alpha isoform(Human)
Genentech
Curated by ChEMBL
Genentech
Curated by ChEMBL
Affinity DataKi: 0.355nMAssay Description:Inhibition of PI3Kalpha (unknown origin) using PIP2:3PS as substrate in presence of ATP measured after 120 mins by ADP-Glo assayMore data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit alpha isoform(Human)
Genentech
Curated by ChEMBL
Genentech
Curated by ChEMBL
Affinity DataKi: 0.440nMAssay Description:Inhibition of PI3Kalpha (unknown origin) using PIP2:3PS as substrate in presence of ATP measured after 120 mins by ADP-Glo assayMore data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit alpha isoform(Human)
Genentech
Curated by ChEMBL
Genentech
Curated by ChEMBL
Affinity DataKi: 0.560nMAssay Description:Inhibition of PI3Kalpha (unknown origin) using PIP2:3PS as substrate in presence of ATP measured after 120 mins by ADP-Glo assayMore data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit alpha isoform(Human)
Genentech
Curated by ChEMBL
Genentech
Curated by ChEMBL
Affinity DataKi: 0.681nMAssay Description:Inhibition of PI3Kalpha (unknown origin) using PIP2:3PS as substrate in presence of ATP measured after 120 mins by ADP-Glo assayMore data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit alpha isoform(Human)
Genentech
Curated by ChEMBL
Genentech
Curated by ChEMBL
Affinity DataKi: 0.692nMAssay Description:Inhibition of PI3Kalpha (unknown origin) using PIP2:3PS as substrate in presence of ATP measured after 120 mins by ADP-Glo assayMore data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit alpha isoform(Human)
Genentech
Curated by ChEMBL
Genentech
Curated by ChEMBL
Affinity DataKi: 0.800nMAssay Description:Inhibition of PI3Kalpha (unknown origin) using PIP2:3PS as substrate in presence of ATP measured after 120 mins by ADP-Glo assayMore data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit alpha isoform(Human)
Genentech
Curated by ChEMBL
Genentech
Curated by ChEMBL
Affinity DataKi: 0.853nMAssay Description:Inhibition of PI3Kalpha (unknown origin) using PIP2:3PS as substrate in presence of ATP measured after 120 mins by ADP-Glo assayMore data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit alpha isoform(Human)
Genentech
Curated by ChEMBL
Genentech
Curated by ChEMBL
Affinity DataKi: 2.5nMAssay Description:Inhibition of PI3Kalpha (unknown origin) using PIP2:3PS as substrate in presence of ATP measured after 120 mins by ADP-Glo assayMore data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit alpha isoform(Human)
Genentech
Curated by ChEMBL
Genentech
Curated by ChEMBL
Affinity DataKi: 3nMAssay Description:Inhibition of PI3Kalpha (unknown origin) using PIP2:3PS as substrate in presence of ATP measured after 120 mins by ADP-Glo assayMore data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit alpha isoform(Human)
Genentech
Curated by ChEMBL
Genentech
Curated by ChEMBL
Affinity DataKi: 3.20nMAssay Description:Inhibition of PI3Kalpha (unknown origin) using PIP2:3PS as substrate in presence of ATP measured after 120 mins by ADP-Glo assayMore data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit alpha isoform(Human)
Genentech
Curated by ChEMBL
Genentech
Curated by ChEMBL
Affinity DataKi: 3.20nMAssay Description:Inhibition of PI3Kalpha (unknown origin) using PIP2:3PS as substrate in presence of ATP measured after 120 mins by ADP-Glo assayMore data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit alpha isoform(Human)
Genentech
Curated by ChEMBL
Genentech
Curated by ChEMBL
Affinity DataKi: 3.80nMAssay Description:Inhibition of PI3Kalpha (unknown origin) using PIP2:3PS as substrate in presence of ATP measured after 120 mins by ADP-Glo assayMore data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit alpha isoform(Human)
Genentech
Curated by ChEMBL
Genentech
Curated by ChEMBL
Affinity DataKi: 5.80nMAssay Description:Inhibition of PI3Kalpha (unknown origin) using PIP2:3PS as substrate in presence of ATP measured after 120 mins by ADP-Glo assayMore data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit alpha isoform(Human)
Genentech
Curated by ChEMBL
Genentech
Curated by ChEMBL
Affinity DataKi: 6.5nMAssay Description:Inhibition of PI3Kalpha (unknown origin) using PIP2:3PS as substrate in presence of ATP measured after 120 mins by ADP-Glo assayMore data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit alpha isoform(Human)
Genentech
Curated by ChEMBL
Genentech
Curated by ChEMBL
Affinity DataKi: 12nMAssay Description:Inhibition of PI3Kalpha (unknown origin) using PIP2:3PS as substrate in presence of ATP measured after 120 mins by ADP-Glo assayMore data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit delta isoform(Human)
Genentech
Curated by ChEMBL
Genentech
Curated by ChEMBL
Affinity DataKi: 12nMAssay Description:Inhibition of PI3Kdelta (unknown origin) using PIP2:3PS as substrate in presence of ATP measured after 120 mins by ADP-Glo assayMore data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit gamma isoform(Human)
Genentech
Curated by ChEMBL
Genentech
Curated by ChEMBL
Affinity DataKi: 18nMAssay Description:Inhibition of PI3Kgamma (unknown origin) using PIP2:3PS as substrate in presence of ATP measured after 120 mins by ADP-Glo assayMore data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit alpha isoform(Human)
Genentech
Curated by ChEMBL
Genentech
Curated by ChEMBL
Affinity DataEC50: 19nMAssay Description:Inhibition of PI3Kalpha H1047R mutant in human HCC1954 cells assessed as reduction in PRAS40 phosphorylation after 24 hrs by electrochemiluminescent ...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit alpha isoform(Human)
Genentech
Curated by ChEMBL
Genentech
Curated by ChEMBL
Affinity DataEC50: 24nMAssay Description:Inhibition of PI3Kalpha H1047R mutant in human HCC1954 cells assessed as reduction in PRAS40 phosphorylation after 24 hrs by electrochemiluminescent ...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit alpha isoform(Human)
Genentech
Curated by ChEMBL
Genentech
Curated by ChEMBL
Affinity DataEC50: 27nMAssay Description:Inhibition of PI3Kalpha H1047R mutant in human HCC1954 cells assessed as reduction in PRAS40 phosphorylation after 24 hrs by electrochemiluminescent ...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit alpha isoform(Human)
Genentech
Curated by ChEMBL
Genentech
Curated by ChEMBL
Affinity DataEC50: 28nMAssay Description:Inhibition of PI3Kalpha H1047R mutant in human HCC1954 cells assessed as reduction in PRAS40 phosphorylation after 24 hrs by electrochemiluminescent ...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit alpha isoform(Human)
Genentech
Curated by ChEMBL
Genentech
Curated by ChEMBL
Affinity DataKi: 30nMAssay Description:Inhibition of PI3Kalpha (unknown origin) using PIP2:3PS as substrate in presence of ATP measured after 120 mins by ADP-Glo assayMore data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit alpha isoform(Human)
Genentech
Curated by ChEMBL
Genentech
Curated by ChEMBL
Affinity DataEC50: 39nMAssay Description:Inhibition of PI3Kalpha H1047R mutant in human HCC1954 cells assessed as reduction in PRAS40 phosphorylation after 24 hrs by electrochemiluminescent ...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit alpha isoform(Human)
Genentech
Curated by ChEMBL
Genentech
Curated by ChEMBL
Affinity DataEC50: 40nMAssay Description:Inhibition of PI3Kalpha H1047R mutant in human HCC1954 cells assessed as reduction in PRAS40 phosphorylation after 24 hrs by electrochemiluminescent ...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit alpha isoform(Human)
Genentech
Curated by ChEMBL
Genentech
Curated by ChEMBL
Affinity DataEC50: 49nMAssay Description:Inhibition of PI3Kalpha H1047R mutant in human HCC1954 cells assessed as reduction in PRAS40 phosphorylation after 24 hrs by electrochemiluminescent ...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit alpha isoform(Human)
Genentech
Curated by ChEMBL
Genentech
Curated by ChEMBL
Affinity DataEC50: 52nMAssay Description:Inhibition of PI3Kalpha H1047R mutant in human HCC1954 cells assessed as reduction in PRAS40 phosphorylation after 24 hrs by electrochemiluminescent ...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit alpha isoform(Human)
Genentech
Curated by ChEMBL
Genentech
Curated by ChEMBL
Affinity DataEC50: 56nMAssay Description:Inhibition of PI3Kalpha H1047R mutant in human HCC1954 cells assessed as reduction in PRAS40 phosphorylation after 24 hrs by electrochemiluminescent ...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit alpha isoform(Human)
Genentech
Curated by ChEMBL
Genentech
Curated by ChEMBL
Affinity DataEC50: 63nMAssay Description:Inhibition of PI3Kalpha H1047R mutant in human HCC1954 cells assessed as reduction in PRAS40 phosphorylation after 24 hrs by electrochemiluminescent ...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit alpha isoform(Human)
Genentech
Curated by ChEMBL
Genentech
Curated by ChEMBL
Affinity DataEC50: 65nMAssay Description:Inhibition of PI3Kalpha H1047R mutant in human HCC1954 cells assessed as reduction in PRAS40 phosphorylation after 24 hrs by electrochemiluminescent ...More data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit alpha isoform(Human)
Genentech
Curated by ChEMBL
Genentech
Curated by ChEMBL
Affinity DataEC50: 89nMAssay Description:Inhibition of PI3Kalpha H1047R mutant in human HCC1954 cells assessed as reduction in PRAS40 phosphorylation after 24 hrs by electrochemiluminescent ...More data for this Ligand-Target Pair