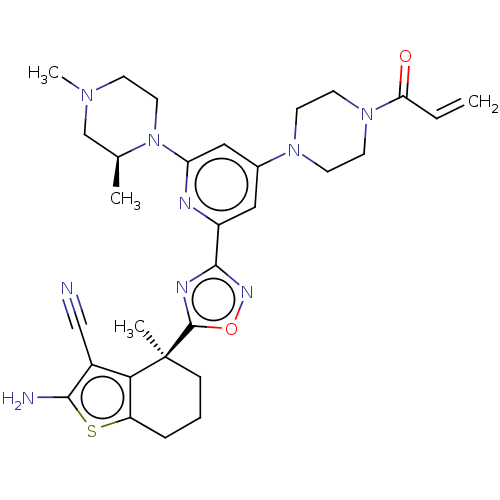
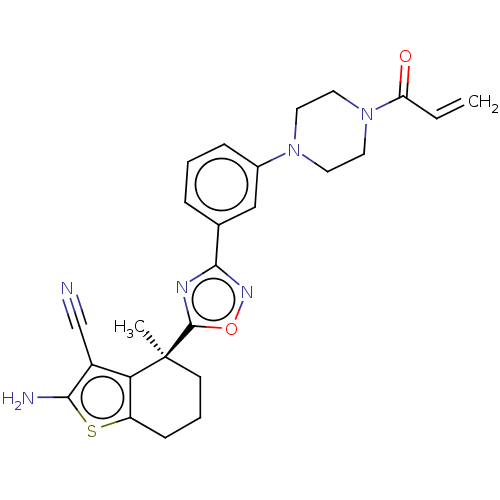
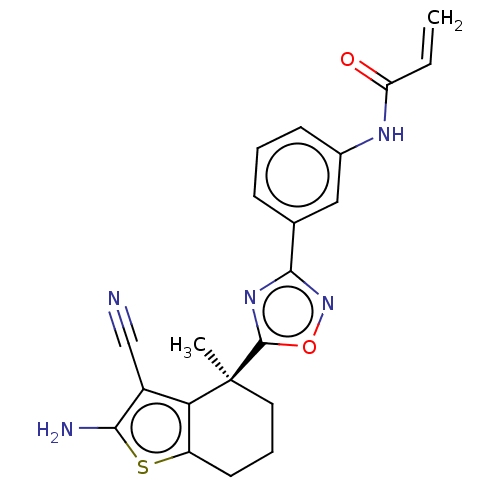
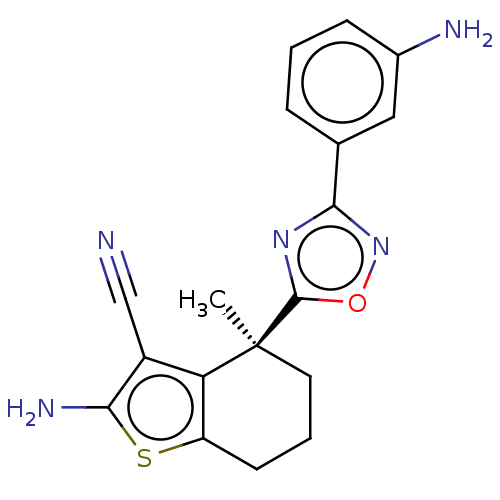
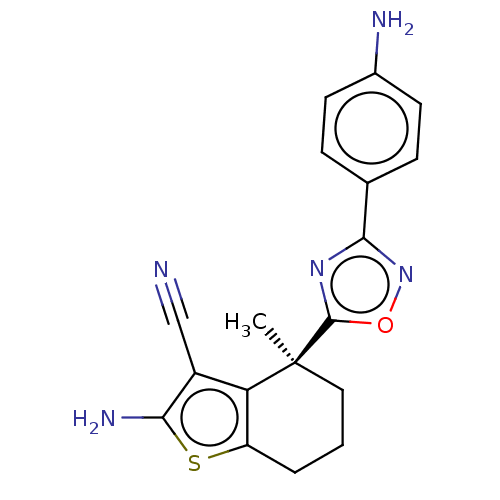
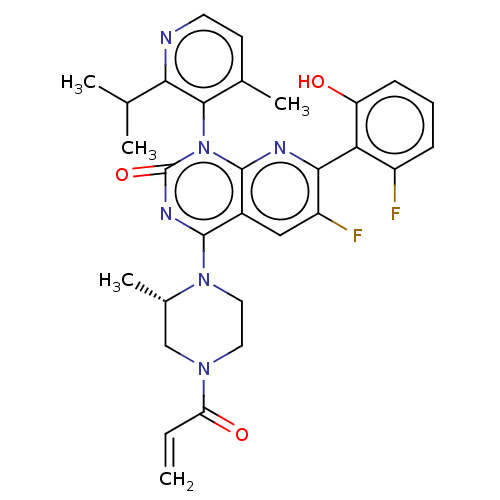
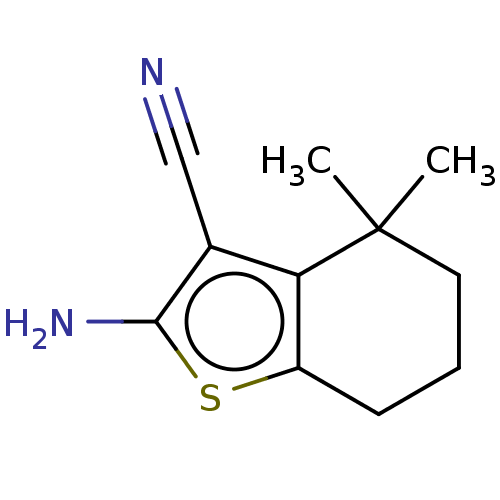
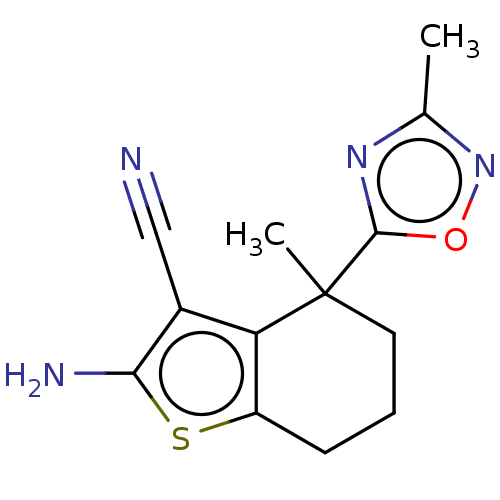
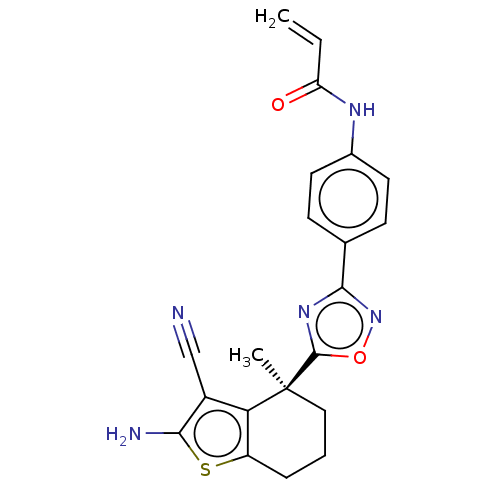
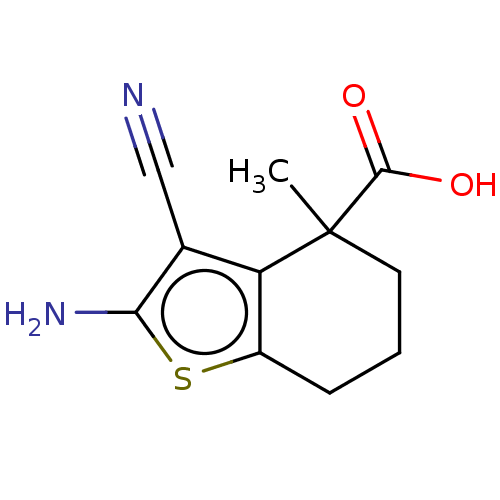
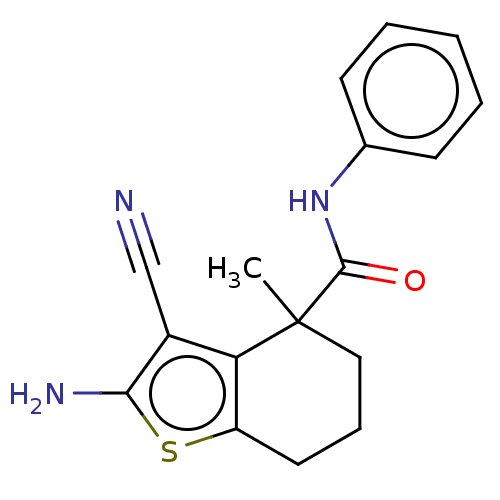
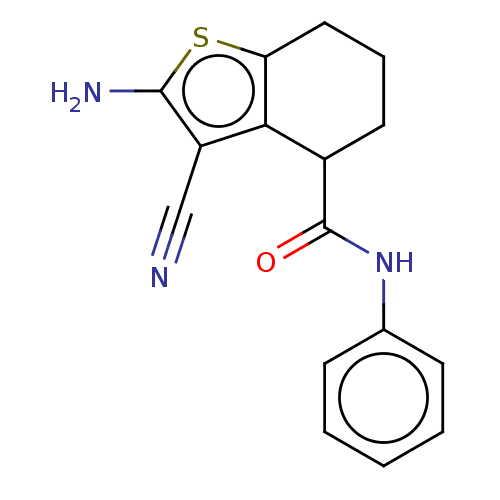
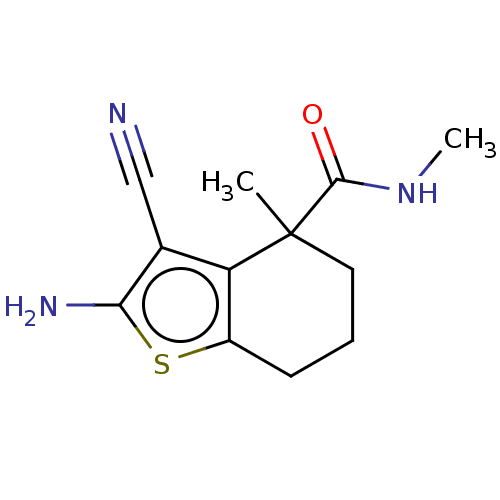
Report error Found 15 Enz. Inhib. hit(s) with all data for entry = 50018080
Affinity DataKi: 2.50E+3nMAssay Description:Binding affinity to KRAS G12C mutant (unknown origin) assessed as inhibition constant incubated for 24 hrs by LC-MS analysisMore data for this Ligand-Target Pair
Affinity DataKi: 7.50E+3nMAssay Description:Binding affinity to KRAS G12C mutant (unknown origin) assessed as inhibition constant incubated for 24 hrs by LC-MS analysisMore data for this Ligand-Target Pair
Affinity DataKi: 8.60E+3nMAssay Description:Binding affinity to KRAS G12C mutant (unknown origin) assessed as inhibition constant incubated for 24 hrs by LC-MS analysisMore data for this Ligand-Target Pair
Affinity DataKd: <1.00E+4nMAssay Description:Binding affinity to KRAS G12V mutant (unknown origin) assessed as dissociation constant by HSQC NMR spectroscopy analysisMore data for this Ligand-Target Pair
Affinity DataKd: <1.00E+4nMAssay Description:Binding affinity to KRAS G12V mutant (unknown origin) assessed as dissociation constant by HSQC NMR spectroscopy analysisMore data for this Ligand-Target Pair
Affinity DataKi: 1.22E+4nMAssay Description:Binding affinity to KRAS G12C mutant (unknown origin) assessed as inhibition constant incubated for 24 hrs by LC-MS analysisMore data for this Ligand-Target Pair
Affinity DataKd: 1.50E+4nMAssay Description:Binding affinity to KRAS G12V mutant (unknown origin) assessed as dissociation constant by HSQC NMR spectroscopy analysisMore data for this Ligand-Target Pair
Affinity DataKd: 2.00E+4nMAssay Description:Binding affinity to KRAS G12V mutant (unknown origin) assessed as dissociation constant by HSQC NMR spectroscopy analysisMore data for this Ligand-Target Pair
Affinity DataKi: 2.41E+4nMAssay Description:Binding affinity to KRAS G12C mutant (unknown origin) assessed as inhibition constant incubated for 24 hrs by LC-MS analysisMore data for this Ligand-Target Pair
Affinity DataKd: 1.16E+6nMAssay Description:Binding affinity to KRAS G12V mutant (unknown origin) assessed as dissociation constant by HSQC NMR spectroscopy analysisMore data for this Ligand-Target Pair
Affinity DataKd: 1.18E+6nMAssay Description:Binding affinity to KRAS G12V mutant (unknown origin) assessed as dissociation constant by HSQC NMR spectroscopy analysisMore data for this Ligand-Target Pair
Affinity DataKd: 1.27E+6nMAssay Description:Binding affinity to KRAS G12V mutant (unknown origin) assessed as dissociation constant by HSQC NMR spectroscopy analysisMore data for this Ligand-Target Pair
Affinity DataKd: >2.00E+6nMAssay Description:Binding affinity to KRAS G12D mutant (unknown origin) assessed as dissociation constant by HSQC NMR spectroscopy analysisMore data for this Ligand-Target Pair
Affinity DataKd: >2.00E+6nMAssay Description:Binding affinity to KRAS G12D mutant (unknown origin) assessed as dissociation constant by HSQC NMR spectroscopy analysisMore data for this Ligand-Target Pair
Affinity DataKd: >2.00E+6nMAssay Description:Binding affinity to KRAS G12V mutant (unknown origin) assessed as dissociation constant by HSQC NMR spectroscopy analysisMore data for this Ligand-Target Pair