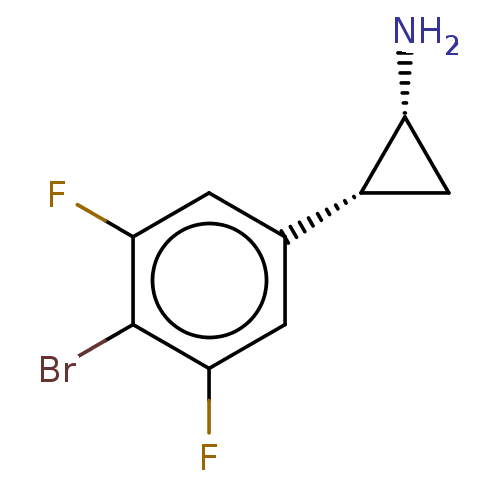
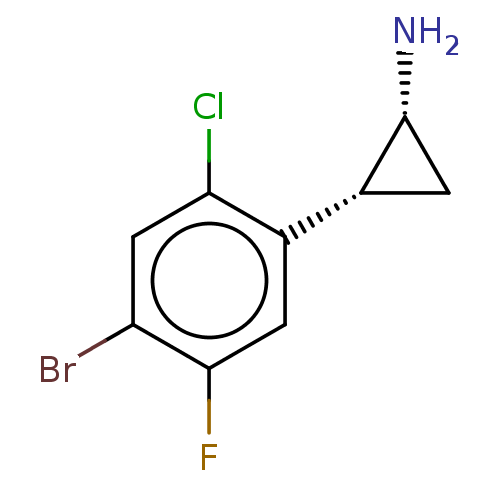
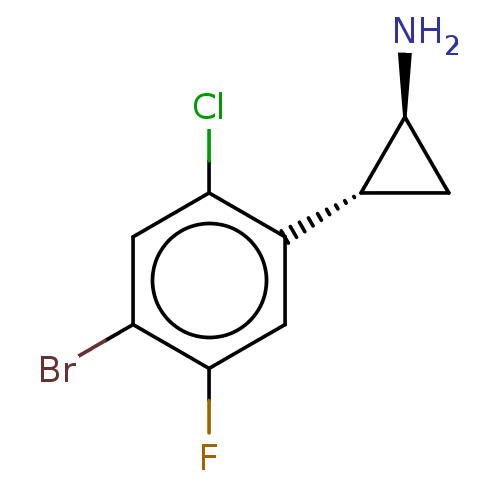
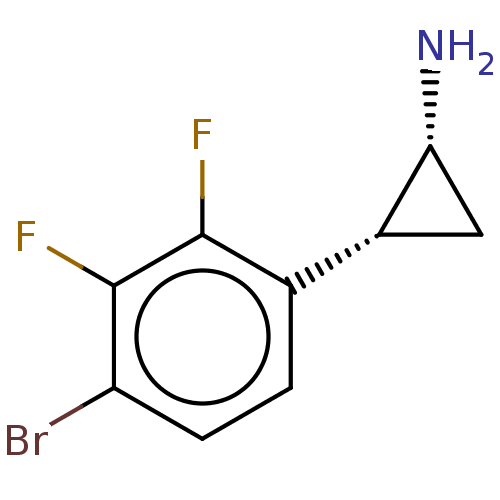
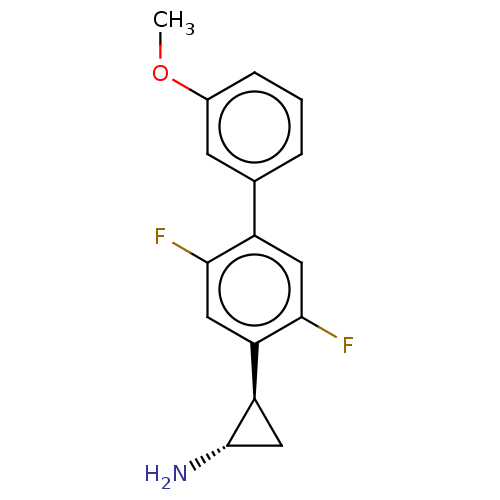
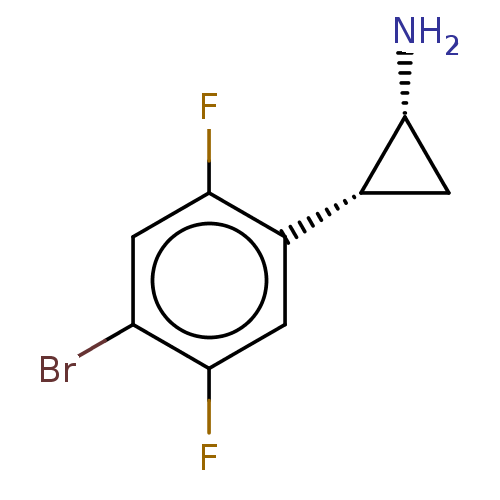
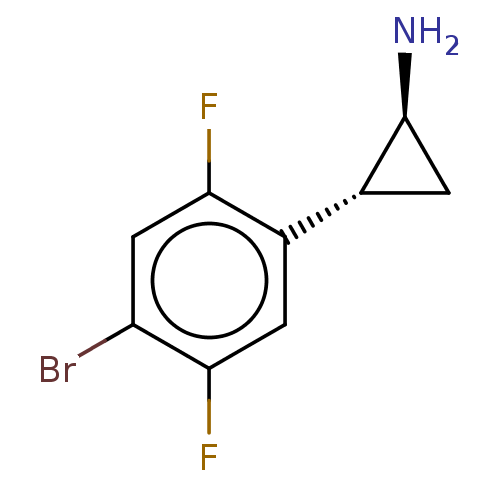
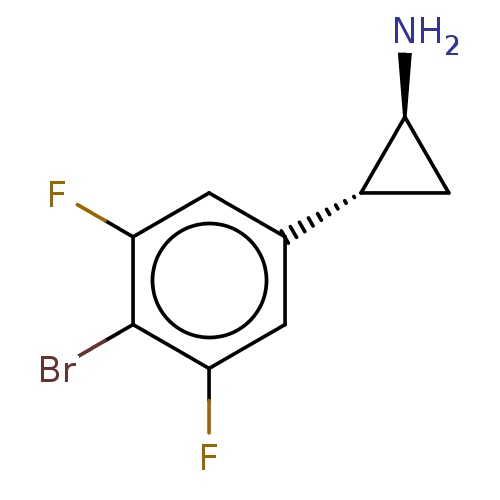
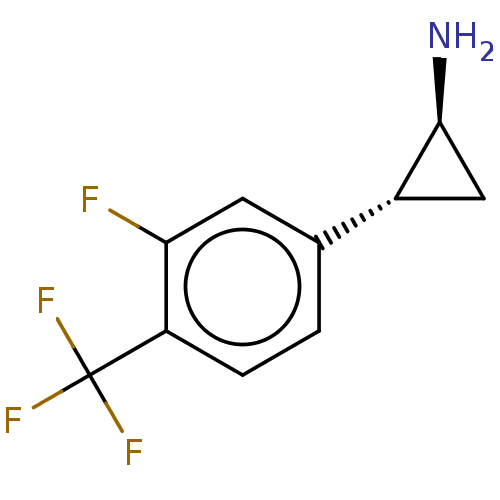
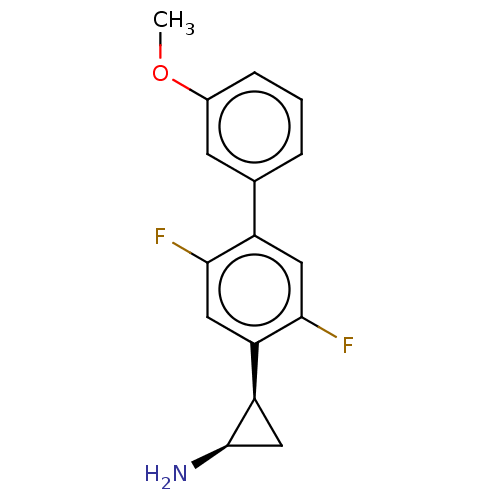
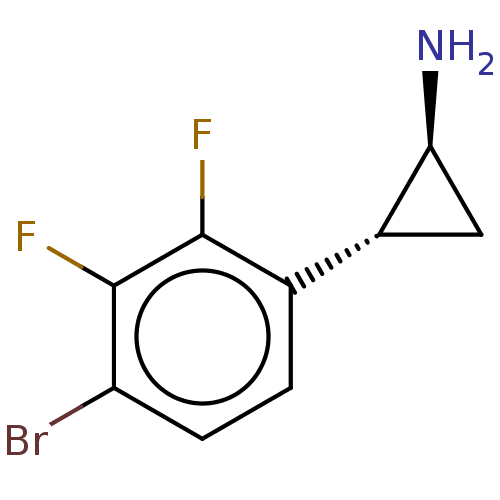
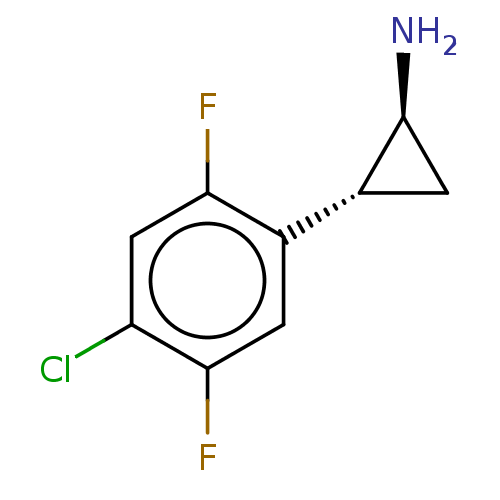
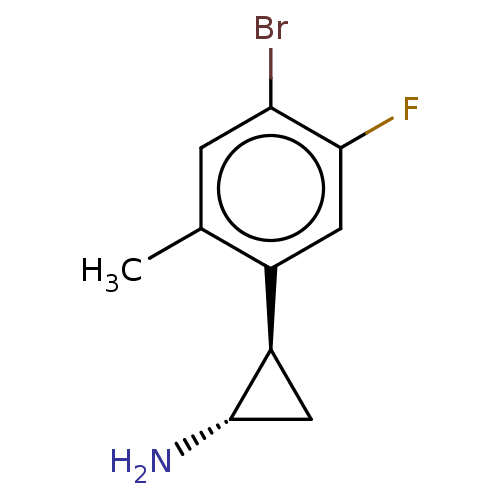
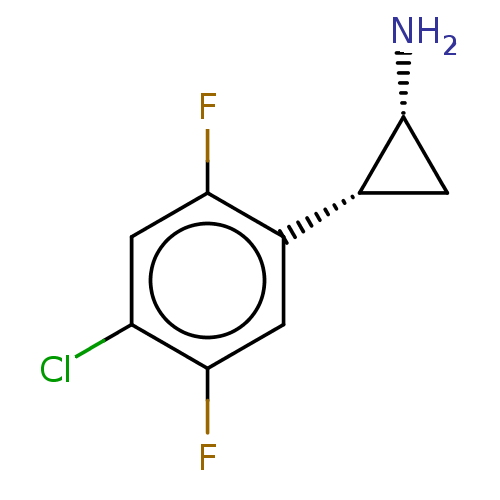
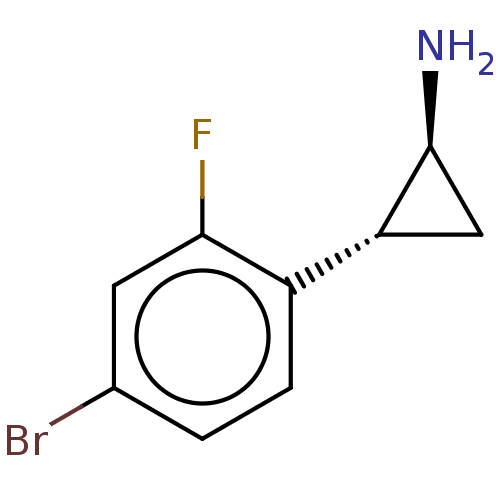
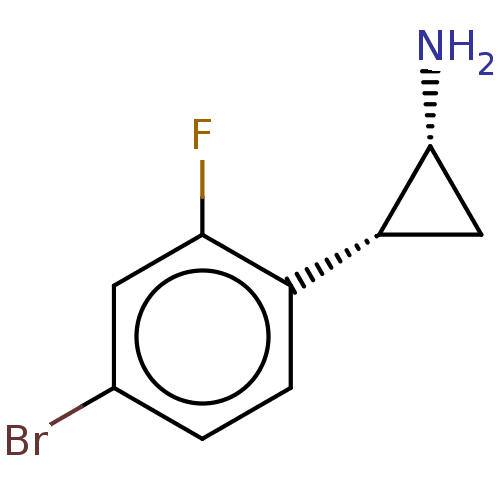
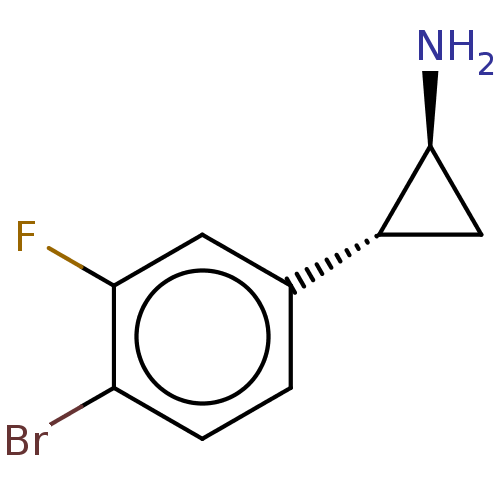
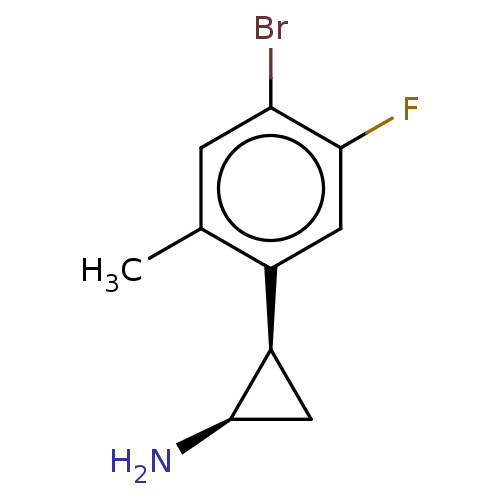
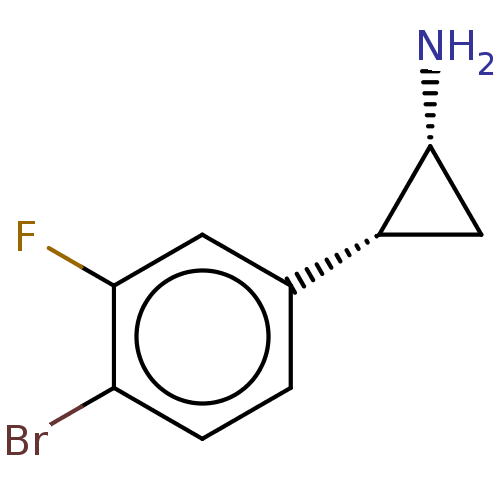
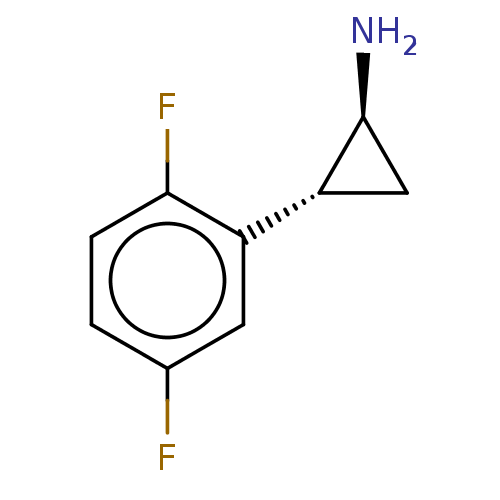
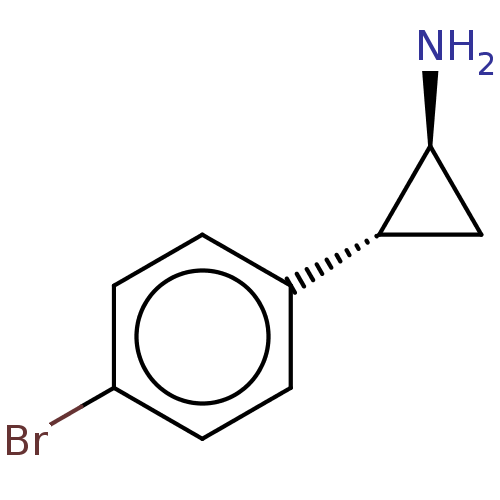
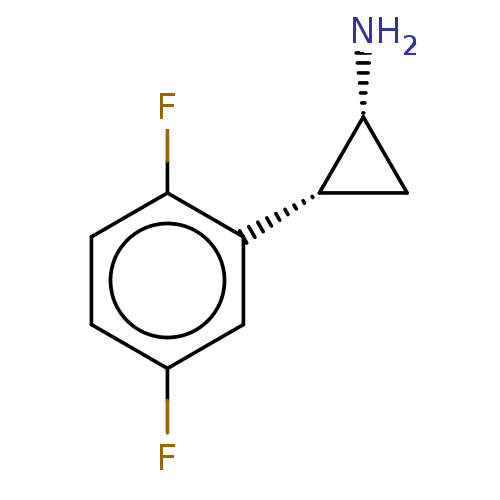
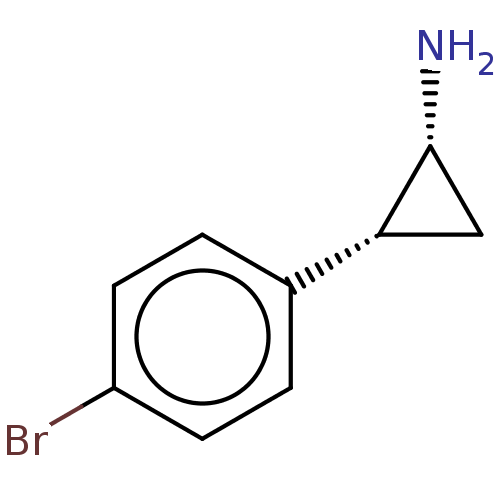
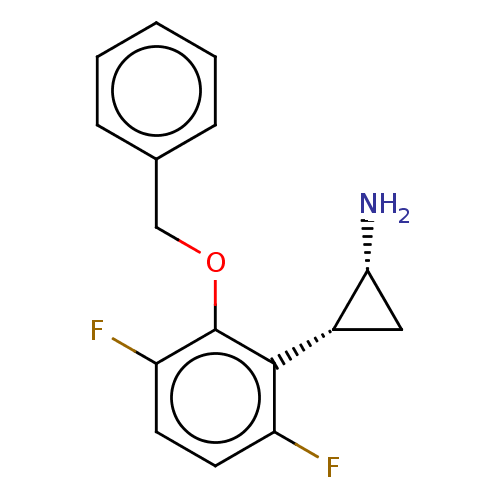
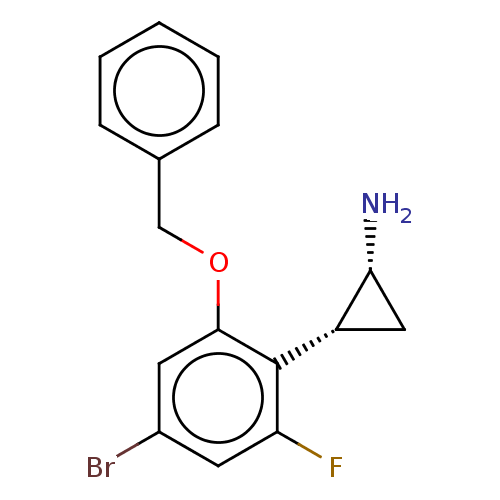
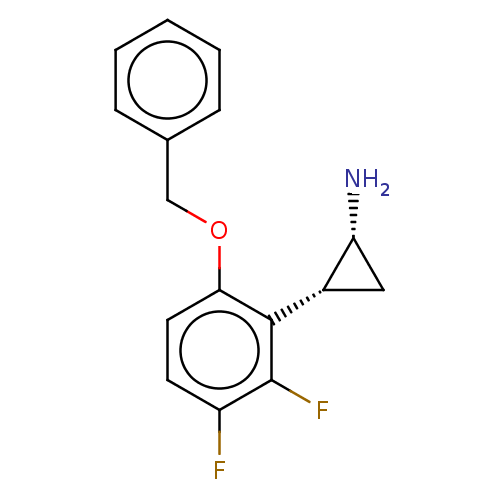
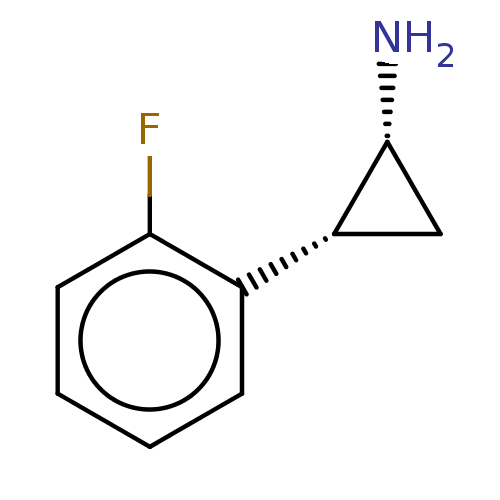
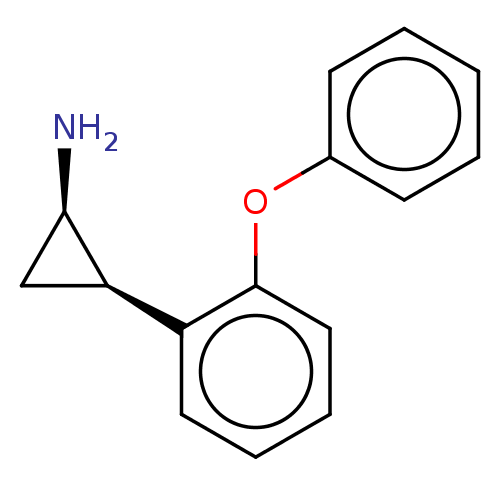
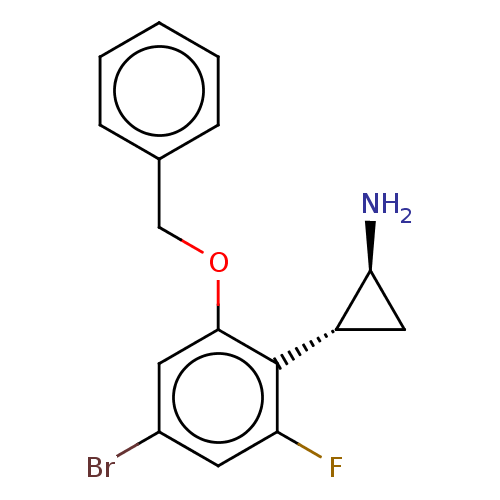
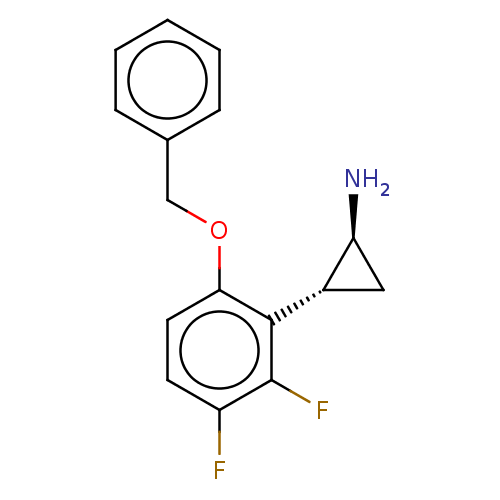
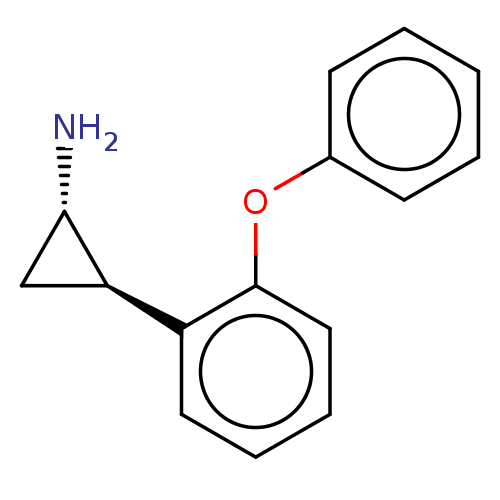
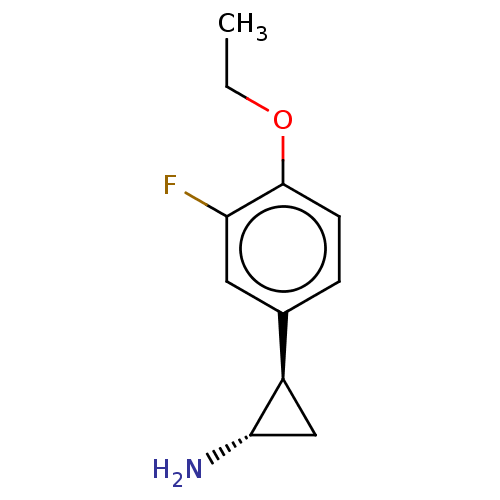
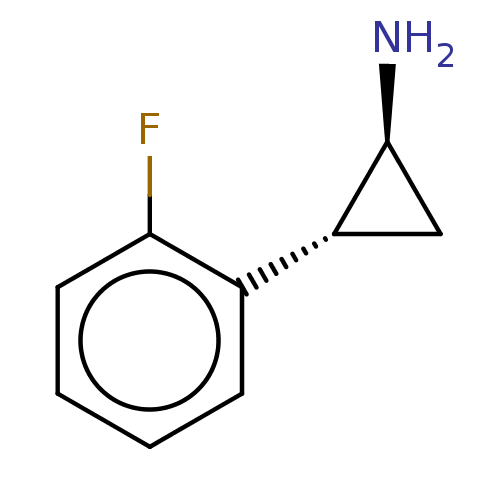
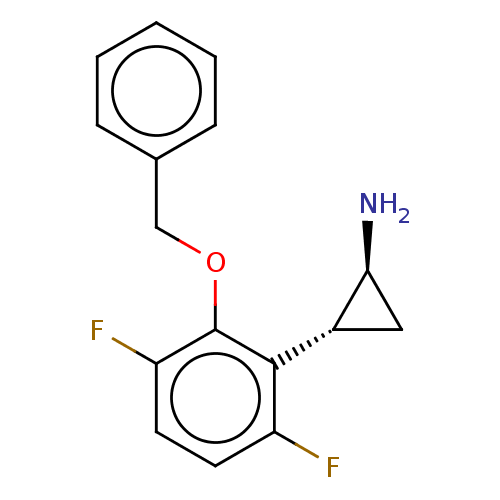
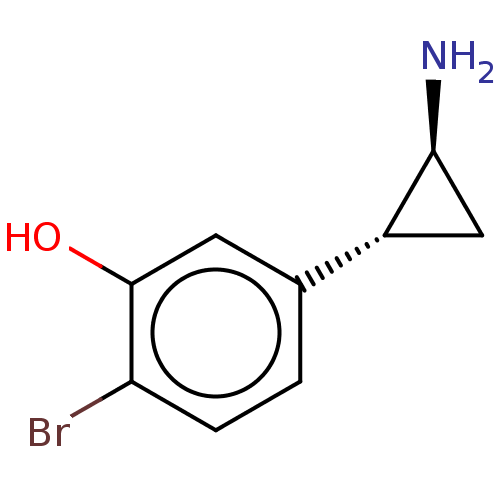
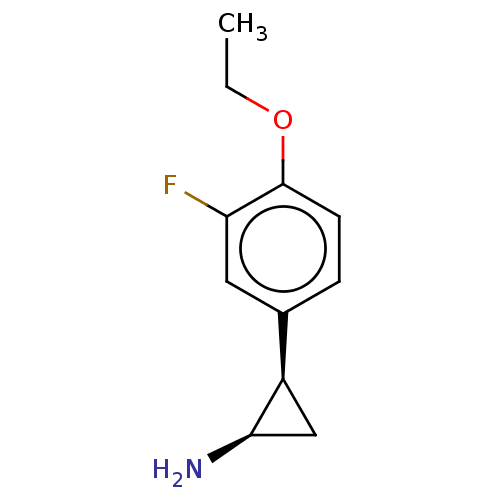
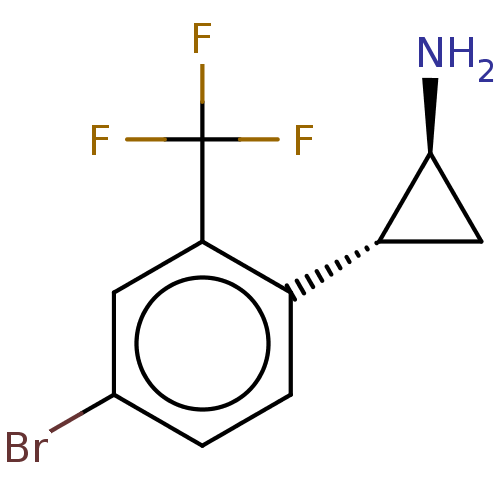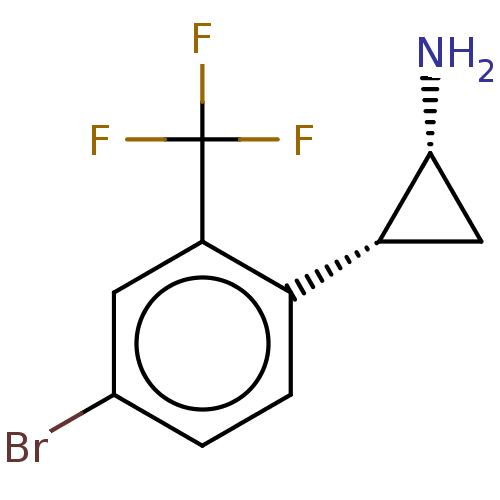Report error Found 94 Enz. Inhib. hit(s) with all data for entry = 50016822
TargetLysine-specific histone demethylase 1A(Human)
Riken Center For Biosystems Dynamics Research
Curated by ChEMBL
Riken Center For Biosystems Dynamics Research
Curated by ChEMBL
Affinity DataKi: 11nMAssay Description:Inhibition of LSD1 (unknown origin) (172 to 833 residues) expressed in baculovirus-infected Sf9 insect cells using K4-dimethylated H3 as substrate by...More data for this Ligand-Target Pair
TargetLysine-specific histone demethylase 1A(Human)
Riken Center For Biosystems Dynamics Research
Curated by ChEMBL
Riken Center For Biosystems Dynamics Research
Curated by ChEMBL
Affinity DataKi: 13nMAssay Description:Inhibition of LSD1 (unknown origin) (172 to 833 residues) expressed in baculovirus-infected Sf9 insect cells using K4-dimethylated H3 as substrate by...More data for this Ligand-Target Pair
TargetLysine-specific histone demethylase 1A(Human)
Riken Center For Biosystems Dynamics Research
Curated by ChEMBL
Riken Center For Biosystems Dynamics Research
Curated by ChEMBL
Affinity DataKi: 27nMAssay Description:Inhibition of LSD1 (unknown origin) (172 to 833 residues) expressed in baculovirus-infected Sf9 insect cells using K4-dimethylated H3 as substrate by...More data for this Ligand-Target Pair
TargetLysine-specific histone demethylase 1A(Human)
Riken Center For Biosystems Dynamics Research
Curated by ChEMBL
Riken Center For Biosystems Dynamics Research
Curated by ChEMBL
Affinity DataKi: 69nMAssay Description:Inhibition of LSD1 (unknown origin) (172 to 833 residues) expressed in baculovirus-infected Sf9 insect cells using K4-dimethylated H3 as substrate by...More data for this Ligand-Target Pair
TargetLysine-specific histone demethylase 1A(Human)
Riken Center For Biosystems Dynamics Research
Curated by ChEMBL
Riken Center For Biosystems Dynamics Research
Curated by ChEMBL
Affinity DataKi: 87nMAssay Description:Inhibition of LSD1 (unknown origin) (172 to 833 residues) expressed in baculovirus-infected Sf9 insect cells using K4-dimethylated H3 as substrate by...More data for this Ligand-Target Pair
TargetLysine-specific histone demethylase 1A(Human)
Riken Center For Biosystems Dynamics Research
Curated by ChEMBL
Riken Center For Biosystems Dynamics Research
Curated by ChEMBL
Affinity DataKi: 94nMAssay Description:Inhibition of LSD1 (unknown origin) (172 to 833 residues) expressed in baculovirus-infected Sf9 insect cells using K4-dimethylated H3 as substrate by...More data for this Ligand-Target Pair
TargetLysine-specific histone demethylase 1A(Human)
Riken Center For Biosystems Dynamics Research
Curated by ChEMBL
Riken Center For Biosystems Dynamics Research
Curated by ChEMBL
Affinity DataKi: 98nMAssay Description:Inhibition of LSD1 (unknown origin) (172 to 833 residues) expressed in baculovirus-infected Sf9 insect cells using K4-dimethylated H3 as substrate by...More data for this Ligand-Target Pair
TargetLysine-specific histone demethylase 1A(Human)
Riken Center For Biosystems Dynamics Research
Curated by ChEMBL
Riken Center For Biosystems Dynamics Research
Curated by ChEMBL
Affinity DataKi: 110nMAssay Description:Inhibition of LSD1 (unknown origin) (172 to 833 residues) expressed in baculovirus-infected Sf9 insect cells using K4-dimethylated H3 as substrate by...More data for this Ligand-Target Pair
TargetLysine-specific histone demethylase 1A(Human)
Riken Center For Biosystems Dynamics Research
Curated by ChEMBL
Riken Center For Biosystems Dynamics Research
Curated by ChEMBL
Affinity DataKi: 150nMAssay Description:Inhibition of LSD1 (unknown origin) (172 to 833 residues) expressed in baculovirus-infected Sf9 insect cells using K4-dimethylated H3 as substrate by...More data for this Ligand-Target Pair
TargetLysine-specific histone demethylase 1A(Human)
Riken Center For Biosystems Dynamics Research
Curated by ChEMBL
Riken Center For Biosystems Dynamics Research
Curated by ChEMBL
Affinity DataKi: 180nMAssay Description:Inhibition of LSD1 (unknown origin) (172 to 833 residues) expressed in baculovirus-infected Sf9 insect cells using K4-dimethylated H3 as substrate by...More data for this Ligand-Target Pair
TargetLysine-specific histone demethylase 1A(Human)
Riken Center For Biosystems Dynamics Research
Curated by ChEMBL
Riken Center For Biosystems Dynamics Research
Curated by ChEMBL
Affinity DataKi: 220nMAssay Description:Inhibition of LSD1 (unknown origin) (172 to 833 residues) expressed in baculovirus-infected Sf9 insect cells using K4-dimethylated H3 as substrate by...More data for this Ligand-Target Pair
TargetLysine-specific histone demethylase 1A(Human)
Riken Center For Biosystems Dynamics Research
Curated by ChEMBL
Riken Center For Biosystems Dynamics Research
Curated by ChEMBL
Affinity DataKi: 240nMAssay Description:Inhibition of LSD1 (unknown origin) (172 to 833 residues) expressed in baculovirus-infected Sf9 insect cells using K4-dimethylated H3 as substrate by...More data for this Ligand-Target Pair
TargetLysine-specific histone demethylase 1A(Human)
Riken Center For Biosystems Dynamics Research
Curated by ChEMBL
Riken Center For Biosystems Dynamics Research
Curated by ChEMBL
Affinity DataKi: 250nMAssay Description:Inhibition of LSD1 (unknown origin) (172 to 833 residues) expressed in baculovirus-infected Sf9 insect cells using K4-dimethylated H3 as substrate by...More data for this Ligand-Target Pair
TargetLysine-specific histone demethylase 1A(Human)
Riken Center For Biosystems Dynamics Research
Curated by ChEMBL
Riken Center For Biosystems Dynamics Research
Curated by ChEMBL
Affinity DataKi: 320nMAssay Description:Inhibition of LSD1 (unknown origin) (172 to 833 residues) expressed in baculovirus-infected Sf9 insect cells using K4-dimethylated H3 as substrate by...More data for this Ligand-Target Pair
TargetLysine-specific histone demethylase 1A(Human)
Riken Center For Biosystems Dynamics Research
Curated by ChEMBL
Riken Center For Biosystems Dynamics Research
Curated by ChEMBL
Affinity DataKi: 370nMAssay Description:Inhibition of LSD1 (unknown origin) (172 to 833 residues) expressed in baculovirus-infected Sf9 insect cells using K4-dimethylated H3 as substrate by...More data for this Ligand-Target Pair
TargetLysine-specific histone demethylase 1A(Human)
Riken Center For Biosystems Dynamics Research
Curated by ChEMBL
Riken Center For Biosystems Dynamics Research
Curated by ChEMBL
Affinity DataKi: 380nMAssay Description:Inhibition of LSD1 (unknown origin) (172 to 833 residues) expressed in baculovirus-infected Sf9 insect cells using K4-dimethylated H3 as substrate by...More data for this Ligand-Target Pair
TargetLysine-specific histone demethylase 1A(Human)
Riken Center For Biosystems Dynamics Research
Curated by ChEMBL
Riken Center For Biosystems Dynamics Research
Curated by ChEMBL
Affinity DataKi: 420nMAssay Description:Inhibition of LSD1 (unknown origin) (172 to 833 residues) expressed in baculovirus-infected Sf9 insect cells using K4-dimethylated H3 as substrate by...More data for this Ligand-Target Pair
TargetLysine-specific histone demethylase 1A(Human)
Riken Center For Biosystems Dynamics Research
Curated by ChEMBL
Riken Center For Biosystems Dynamics Research
Curated by ChEMBL
Affinity DataKi: 470nMAssay Description:Inhibition of LSD1 (unknown origin) (172 to 833 residues) expressed in baculovirus-infected Sf9 insect cells using K4-dimethylated H3 as substrate by...More data for this Ligand-Target Pair
TargetLysine-specific histone demethylase 1A(Human)
Riken Center For Biosystems Dynamics Research
Curated by ChEMBL
Riken Center For Biosystems Dynamics Research
Curated by ChEMBL
Affinity DataKi: 500nMAssay Description:Inhibition of LSD1 (unknown origin) (172 to 833 residues) expressed in baculovirus-infected Sf9 insect cells using K4-dimethylated H3 as substrate by...More data for this Ligand-Target Pair
TargetLysine-specific histone demethylase 1A(Human)
Riken Center For Biosystems Dynamics Research
Curated by ChEMBL
Riken Center For Biosystems Dynamics Research
Curated by ChEMBL
Affinity DataKi: 560nMAssay Description:Inhibition of LSD1 (unknown origin) (172 to 833 residues) expressed in baculovirus-infected Sf9 insect cells using K4-dimethylated H3 as substrate by...More data for this Ligand-Target Pair
TargetLysine-specific histone demethylase 1A(Human)
Riken Center For Biosystems Dynamics Research
Curated by ChEMBL
Riken Center For Biosystems Dynamics Research
Curated by ChEMBL
Affinity DataKi: 660nMAssay Description:Inhibition of LSD1 (unknown origin) (172 to 833 residues) expressed in baculovirus-infected Sf9 insect cells using K4-dimethylated H3 as substrate by...More data for this Ligand-Target Pair
TargetLysine-specific histone demethylase 1A(Human)
Riken Center For Biosystems Dynamics Research
Curated by ChEMBL
Riken Center For Biosystems Dynamics Research
Curated by ChEMBL
Affinity DataKi: 1.90E+3nMAssay Description:Inhibition of LSD1 (unknown origin) (172 to 833 residues) expressed in baculovirus-infected Sf9 insect cells using K4-dimethylated H3 as substrate by...More data for this Ligand-Target Pair
TargetLysine-specific histone demethylase 1A(Human)
Riken Center For Biosystems Dynamics Research
Curated by ChEMBL
Riken Center For Biosystems Dynamics Research
Curated by ChEMBL
Affinity DataKi: 2.90E+3nMAssay Description:Inhibition of LSD1 (unknown origin) (172 to 833 residues) expressed in baculovirus-infected Sf9 insect cells using K4-dimethylated H3 as substrate by...More data for this Ligand-Target Pair
TargetLysine-specific histone demethylase 1A(Human)
Riken Center For Biosystems Dynamics Research
Curated by ChEMBL
Riken Center For Biosystems Dynamics Research
Curated by ChEMBL
Affinity DataKi: 3.40E+3nMAssay Description:Inhibition of LSD1 (unknown origin) (172 to 833 residues) expressed in baculovirus-infected Sf9 insect cells using K4-dimethylated H3 as substrate by...More data for this Ligand-Target Pair
TargetLysine-specific histone demethylase 1A(Human)
Riken Center For Biosystems Dynamics Research
Curated by ChEMBL
Riken Center For Biosystems Dynamics Research
Curated by ChEMBL
Affinity DataKi: 5.90E+3nMAssay Description:Inhibition of LSD1 (unknown origin) (172 to 833 residues) expressed in baculovirus-infected Sf9 insect cells using K4-dimethylated H3 as substrate by...More data for this Ligand-Target Pair
TargetLysine-specific histone demethylase 2(Human)
Riken Center For Biosystems Dynamics Research
Curated by ChEMBL
Riken Center For Biosystems Dynamics Research
Curated by ChEMBL
Affinity DataKi: 7.00E+3nMAssay Description:Inhibition of LSD2 (unknown origin) (26 to 822 residues) expressed in baculovirus-infected Sf9 insect cells using K4-dimethylated H3 as substrate by ...More data for this Ligand-Target Pair
TargetLysine-specific histone demethylase 2(Human)
Riken Center For Biosystems Dynamics Research
Curated by ChEMBL
Riken Center For Biosystems Dynamics Research
Curated by ChEMBL
Affinity DataKi: 7.00E+3nMAssay Description:Inhibition of LSD2 (unknown origin) (26 to 822 residues) expressed in baculovirus-infected Sf9 insect cells using K4-dimethylated H3 as substrate by ...More data for this Ligand-Target Pair
TargetPotassium voltage-gated channel subfamily H member 2(Human)
Riken Center For Biosystems Dynamics Research
Curated by ChEMBL
Riken Center For Biosystems Dynamics Research
Curated by ChEMBL
Affinity DataIC50: 7.30E+3nMAssay Description:Inhibition of human ERG by fluorescence polarization assayMore data for this Ligand-Target Pair
TargetLysine-specific histone demethylase 1A(Human)
Riken Center For Biosystems Dynamics Research
Curated by ChEMBL
Riken Center For Biosystems Dynamics Research
Curated by ChEMBL
Affinity DataKi: 7.30E+3nMAssay Description:Inhibition of LSD1 (unknown origin) (172 to 833 residues) expressed in baculovirus-infected Sf9 insect cells using K4-dimethylated H3 as substrate by...More data for this Ligand-Target Pair
TargetPotassium voltage-gated channel subfamily H member 2(Human)
Riken Center For Biosystems Dynamics Research
Curated by ChEMBL
Riken Center For Biosystems Dynamics Research
Curated by ChEMBL
Affinity DataIC50: 7.70E+3nMAssay Description:Inhibition of human ERG by fluorescence polarization assayMore data for this Ligand-Target Pair
TargetLysine-specific histone demethylase 1A(Human)
Riken Center For Biosystems Dynamics Research
Curated by ChEMBL
Riken Center For Biosystems Dynamics Research
Curated by ChEMBL
Affinity DataKi: 8.30E+3nMAssay Description:Inhibition of LSD1 (unknown origin) (172 to 833 residues) expressed in baculovirus-infected Sf9 insect cells using K4-dimethylated H3 as substrate by...More data for this Ligand-Target Pair
TargetLysine-specific histone demethylase 2(Human)
Riken Center For Biosystems Dynamics Research
Curated by ChEMBL
Riken Center For Biosystems Dynamics Research
Curated by ChEMBL
Affinity DataKi: 8.40E+3nMAssay Description:Inhibition of LSD2 (unknown origin) (26 to 822 residues) expressed in baculovirus-infected Sf9 insect cells using K4-dimethylated H3 as substrate by ...More data for this Ligand-Target Pair
TargetLysine-specific histone demethylase 1A(Human)
Riken Center For Biosystems Dynamics Research
Curated by ChEMBL
Riken Center For Biosystems Dynamics Research
Curated by ChEMBL
Affinity DataKi: 1.00E+4nMAssay Description:Inhibition of LSD1 (unknown origin) (172 to 833 residues) expressed in baculovirus-infected Sf9 insect cells using K4-dimethylated H3 as substrate by...More data for this Ligand-Target Pair
TargetLysine-specific histone demethylase 1A(Human)
Riken Center For Biosystems Dynamics Research
Curated by ChEMBL
Riken Center For Biosystems Dynamics Research
Curated by ChEMBL
Affinity DataKi: 1.10E+4nMAssay Description:Inhibition of LSD1 (unknown origin) (172 to 833 residues) expressed in baculovirus-infected Sf9 insect cells using K4-dimethylated H3 as substrate by...More data for this Ligand-Target Pair
TargetLysine-specific histone demethylase 1A(Human)
Riken Center For Biosystems Dynamics Research
Curated by ChEMBL
Riken Center For Biosystems Dynamics Research
Curated by ChEMBL
Affinity DataKi: 1.20E+4nMAssay Description:Inhibition of LSD1 (unknown origin) (172 to 833 residues) expressed in baculovirus-infected Sf9 insect cells using K4-dimethylated H3 as substrate by...More data for this Ligand-Target Pair
TargetLysine-specific histone demethylase 1A(Human)
Riken Center For Biosystems Dynamics Research
Curated by ChEMBL
Riken Center For Biosystems Dynamics Research
Curated by ChEMBL
Affinity DataKi: 1.20E+4nMAssay Description:Inhibition of LSD1 (unknown origin) (172 to 833 residues) expressed in baculovirus-infected Sf9 insect cells using K4-dimethylated H3 as substrate by...More data for this Ligand-Target Pair
TargetPotassium voltage-gated channel subfamily H member 2(Human)
Riken Center For Biosystems Dynamics Research
Curated by ChEMBL
Riken Center For Biosystems Dynamics Research
Curated by ChEMBL
Affinity DataIC50: 1.30E+4nMAssay Description:Inhibition of human ERG by fluorescence polarization assayMore data for this Ligand-Target Pair
TargetLysine-specific histone demethylase 1A(Human)
Riken Center For Biosystems Dynamics Research
Curated by ChEMBL
Riken Center For Biosystems Dynamics Research
Curated by ChEMBL
Affinity DataKi: 1.40E+4nMAssay Description:Inhibition of LSD1 (unknown origin) (172 to 833 residues) expressed in baculovirus-infected Sf9 insect cells using K4-dimethylated H3 as substrate by...More data for this Ligand-Target Pair
TargetPotassium voltage-gated channel subfamily H member 2(Human)
Riken Center For Biosystems Dynamics Research
Curated by ChEMBL
Riken Center For Biosystems Dynamics Research
Curated by ChEMBL
Affinity DataIC50: 1.50E+4nMAssay Description:Inhibition of human ERG by fluorescence polarization assayMore data for this Ligand-Target Pair
TargetPotassium voltage-gated channel subfamily H member 2(Human)
Riken Center For Biosystems Dynamics Research
Curated by ChEMBL
Riken Center For Biosystems Dynamics Research
Curated by ChEMBL
Affinity DataIC50: 1.60E+4nMAssay Description:Inhibition of human ERG by fluorescence polarization assayMore data for this Ligand-Target Pair
TargetLysine-specific histone demethylase 2(Human)
Riken Center For Biosystems Dynamics Research
Curated by ChEMBL
Riken Center For Biosystems Dynamics Research
Curated by ChEMBL
Affinity DataKi: 1.70E+4nMAssay Description:Inhibition of LSD2 (unknown origin) (26 to 822 residues) expressed in baculovirus-infected Sf9 insect cells using K4-dimethylated H3 as substrate by ...More data for this Ligand-Target Pair
TargetPotassium voltage-gated channel subfamily H member 2(Human)
Riken Center For Biosystems Dynamics Research
Curated by ChEMBL
Riken Center For Biosystems Dynamics Research
Curated by ChEMBL
Affinity DataIC50: 1.80E+4nMAssay Description:Inhibition of human ERG by fluorescence polarization assayMore data for this Ligand-Target Pair
TargetLysine-specific histone demethylase 1A(Human)
Riken Center For Biosystems Dynamics Research
Curated by ChEMBL
Riken Center For Biosystems Dynamics Research
Curated by ChEMBL
Affinity DataKi: 1.90E+4nMAssay Description:Inhibition of LSD1 (unknown origin) (172 to 833 residues) expressed in baculovirus-infected Sf9 insect cells using K4-dimethylated H3 as substrate by...More data for this Ligand-Target Pair
TargetLysine-specific histone demethylase 1A(Human)
Riken Center For Biosystems Dynamics Research
Curated by ChEMBL
Riken Center For Biosystems Dynamics Research
Curated by ChEMBL
Affinity DataKi: 1.90E+4nMAssay Description:Inhibition of LSD1 (unknown origin) (172 to 833 residues) expressed in baculovirus-infected Sf9 insect cells using K4-dimethylated H3 as substrate by...More data for this Ligand-Target Pair
TargetLysine-specific histone demethylase 2(Human)
Riken Center For Biosystems Dynamics Research
Curated by ChEMBL
Riken Center For Biosystems Dynamics Research
Curated by ChEMBL
Affinity DataKi: 2.00E+4nMAssay Description:Inhibition of LSD2 (unknown origin) (26 to 822 residues) expressed in baculovirus-infected Sf9 insect cells using K4-dimethylated H3 as substrate by ...More data for this Ligand-Target Pair
TargetLysine-specific histone demethylase 1A(Human)
Riken Center For Biosystems Dynamics Research
Curated by ChEMBL
Riken Center For Biosystems Dynamics Research
Curated by ChEMBL
Affinity DataKi: 2.00E+4nMAssay Description:Inhibition of LSD1 (unknown origin) (172 to 833 residues) expressed in baculovirus-infected Sf9 insect cells using K4-dimethylated H3 as substrate by...More data for this Ligand-Target Pair
TargetLysine-specific histone demethylase 1A(Human)
Riken Center For Biosystems Dynamics Research
Curated by ChEMBL
Riken Center For Biosystems Dynamics Research
Curated by ChEMBL
Affinity DataKi: 2.60E+4nMAssay Description:Inhibition of LSD1 (unknown origin) (172 to 833 residues) expressed in baculovirus-infected Sf9 insect cells using K4-dimethylated H3 as substrate by...More data for this Ligand-Target Pair
TargetLysine-specific histone demethylase 1A(Human)
Riken Center For Biosystems Dynamics Research
Curated by ChEMBL
Riken Center For Biosystems Dynamics Research
Curated by ChEMBL
Affinity DataKi: 2.80E+4nMAssay Description:Inhibition of LSD1 (unknown origin) (172 to 833 residues) expressed in baculovirus-infected Sf9 insect cells using K4-dimethylated H3 as substrate by...More data for this Ligand-Target Pair
TargetLysine-specific histone demethylase 2(Human)
Riken Center For Biosystems Dynamics Research
Curated by ChEMBL
Riken Center For Biosystems Dynamics Research
Curated by ChEMBL
Affinity DataKi: 2.90E+4nMAssay Description:Inhibition of LSD2 (unknown origin) (26 to 822 residues) expressed in baculovirus-infected Sf9 insect cells using K4-dimethylated H3 as substrate by ...More data for this Ligand-Target Pair
TargetLysine-specific histone demethylase 1A(Human)
Riken Center For Biosystems Dynamics Research
Curated by ChEMBL
Riken Center For Biosystems Dynamics Research
Curated by ChEMBL
Affinity DataKi: 3.10E+4nMAssay Description:Inhibition of LSD1 (unknown origin) (172 to 833 residues) expressed in baculovirus-infected Sf9 insect cells using K4-dimethylated H3 as substrate by...More data for this Ligand-Target Pair