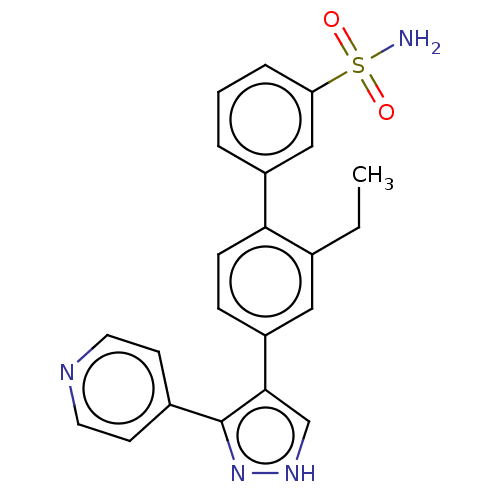
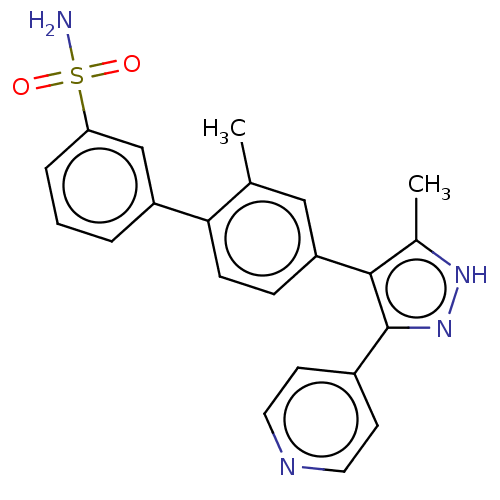
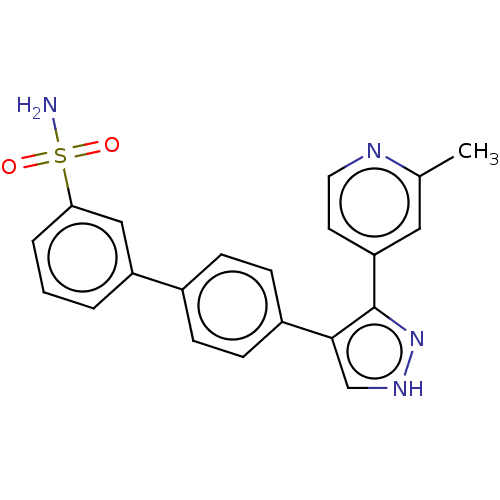
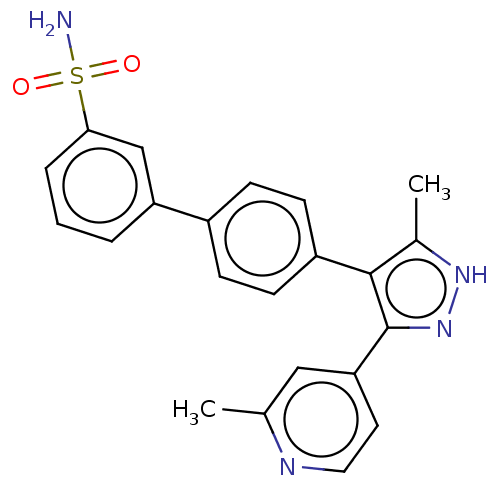
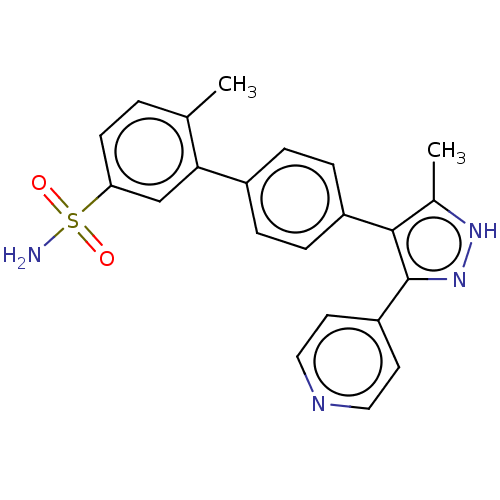
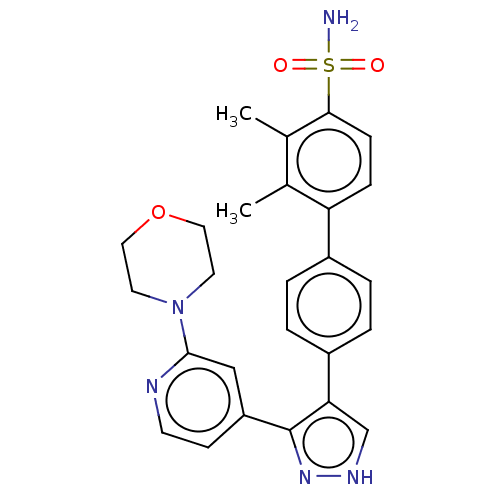
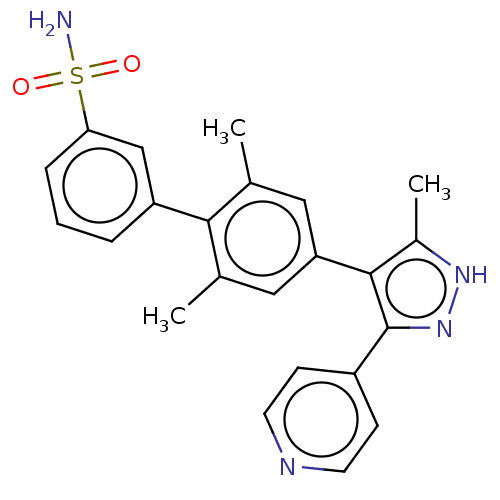
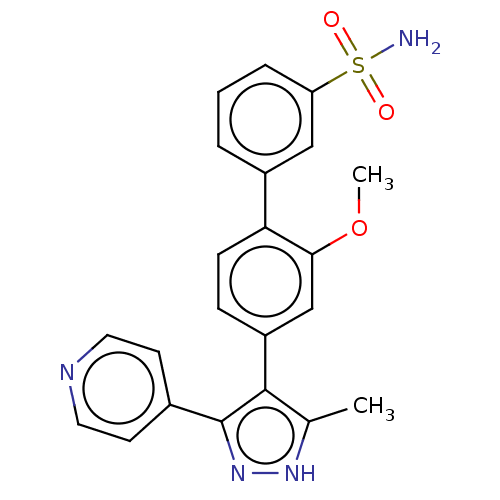
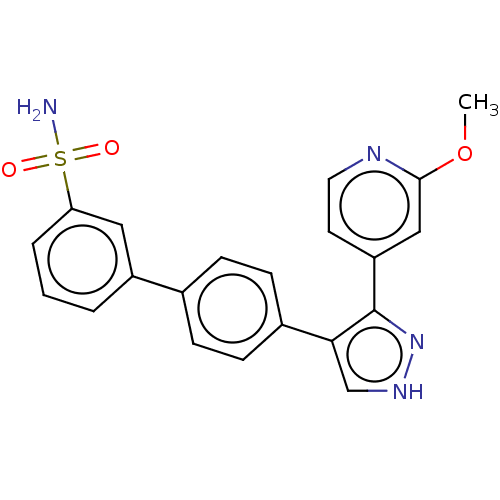
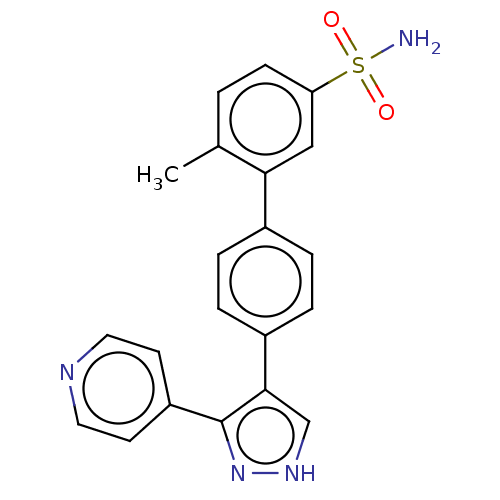
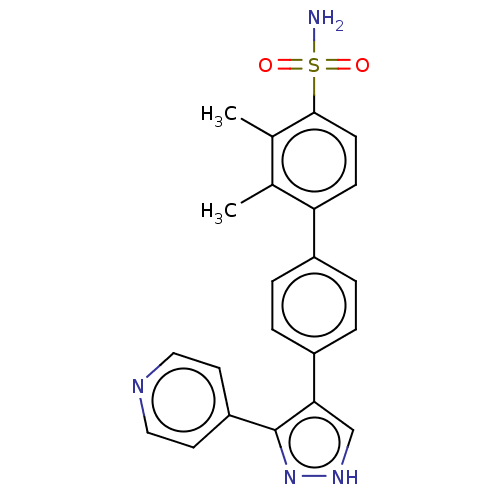
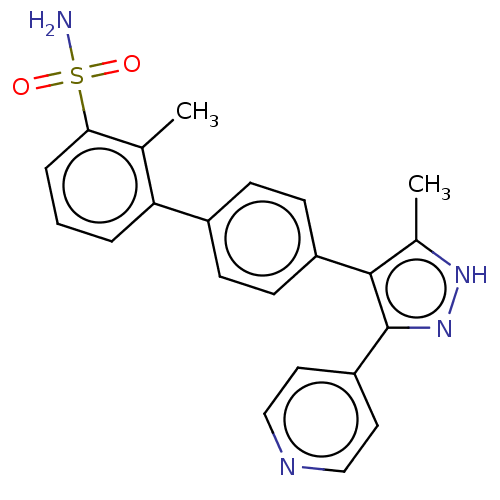
Report error Found 126 Enz. Inhib. hit(s) with all data for entry = 50016782
TargetLeucine-rich repeat serine/threonine-protein kinase 2(Human)
Stanford University
Curated by ChEMBL
Stanford University
Curated by ChEMBL
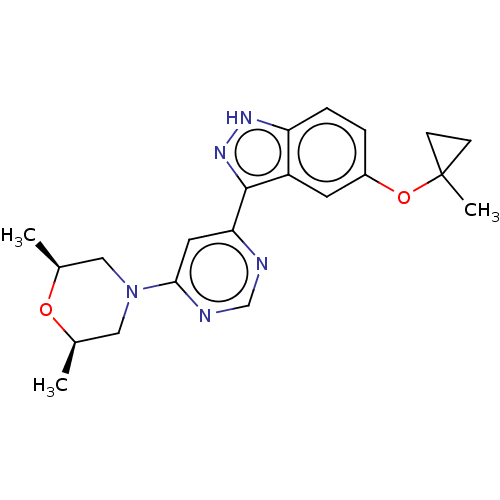
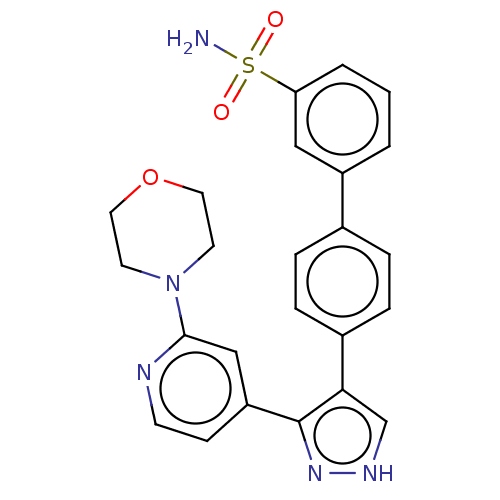
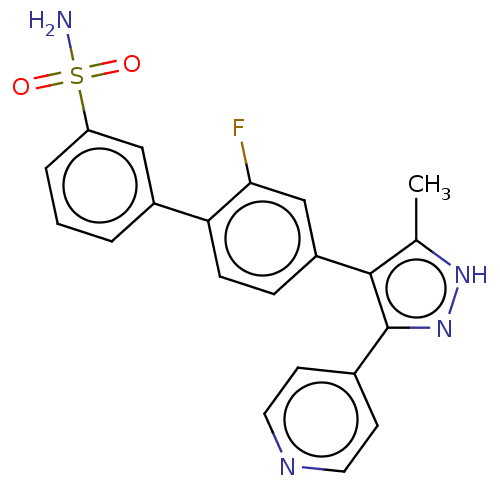
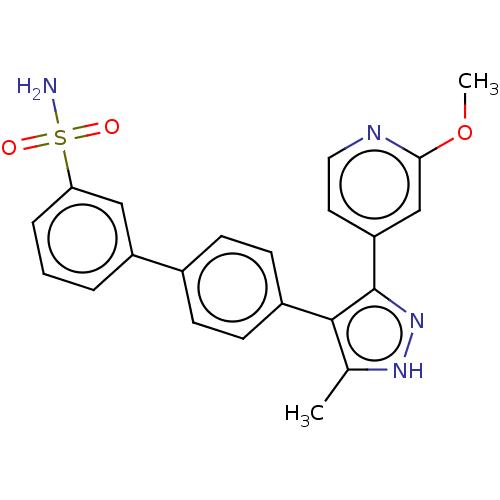
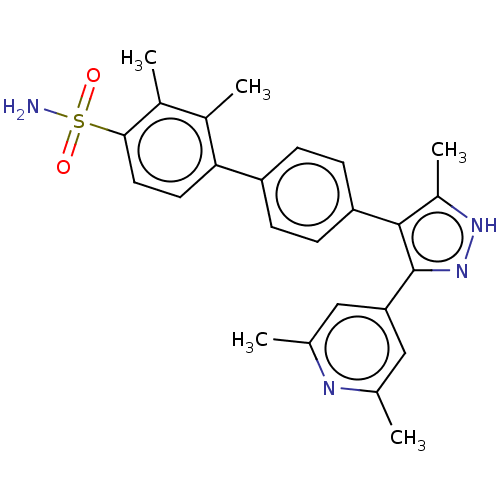
Affinity DataIC50: 0.400nMAssay Description:Inhibition of full-length LRRK2 G2019S mutant (unknown origin) using LRRKtide as substrate incubated for 1 hr in presence of ATP by TR-FRET based ADA...More data for this Ligand-Target Pair
TargetLeucine-rich repeat serine/threonine-protein kinase 2(Human)
Stanford University
Curated by ChEMBL
Stanford University
Curated by ChEMBL
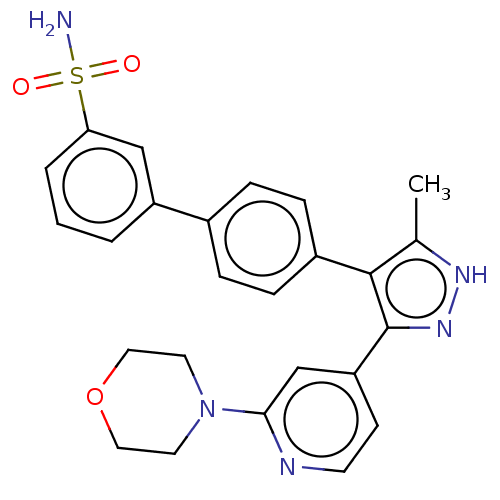
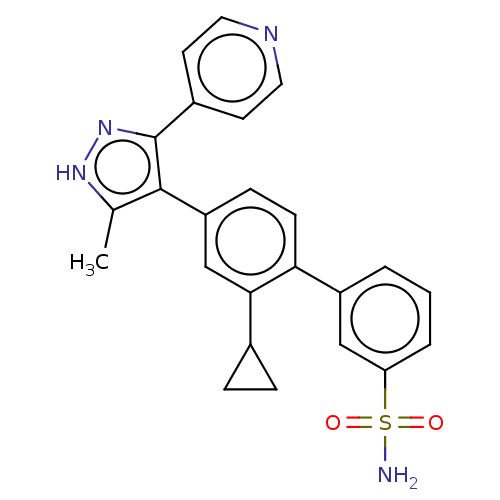
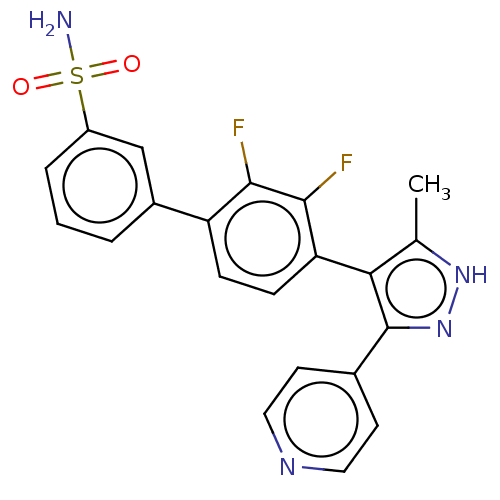
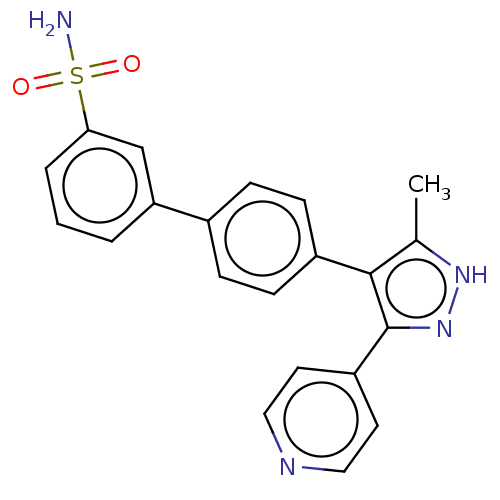
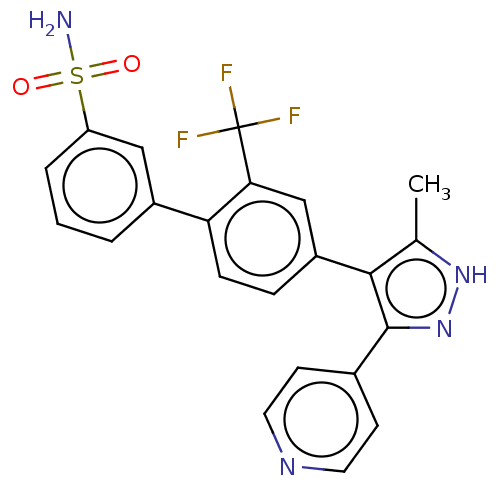
Affinity DataIC50: 1nMAssay Description:Inhibition of full-length wild type LRRK2 (unknown origin) using LRRKtide as substrate incubated for 1 hr in presence of ATP by TR-FRET based ADAPTA ...More data for this Ligand-Target Pair
TargetLeucine-rich repeat serine/threonine-protein kinase 2(Human)
Stanford University
Curated by ChEMBL
Stanford University
Curated by ChEMBL
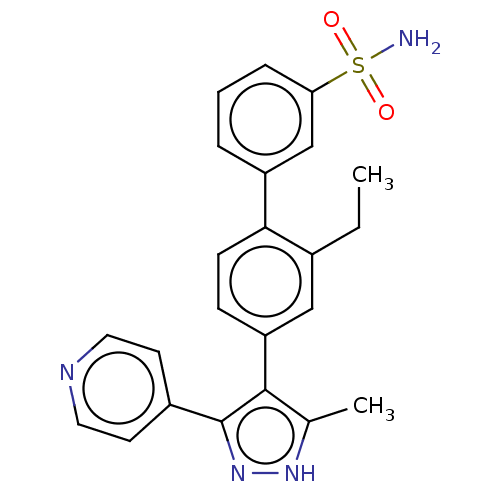
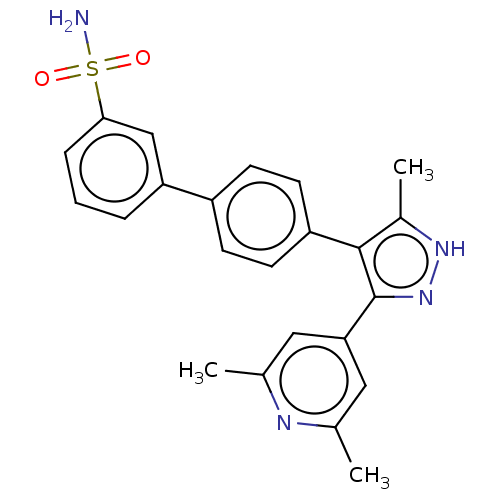
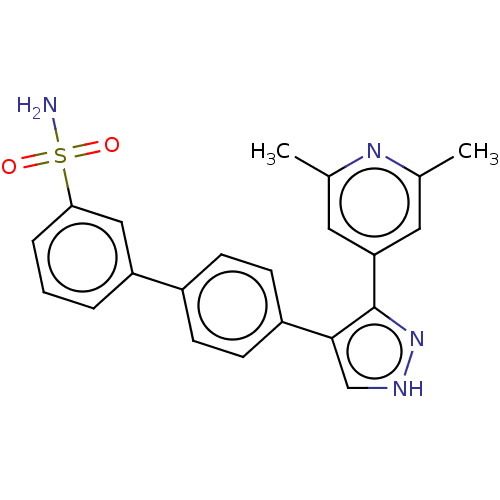
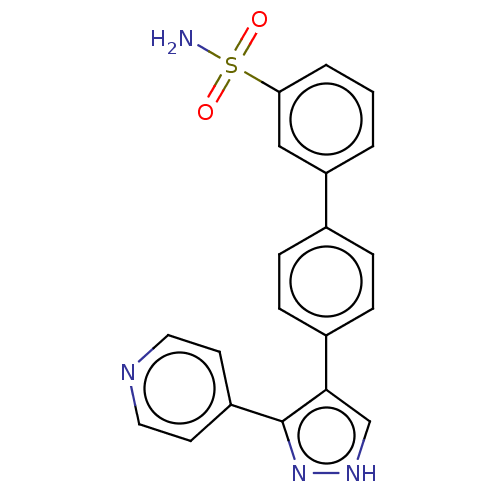
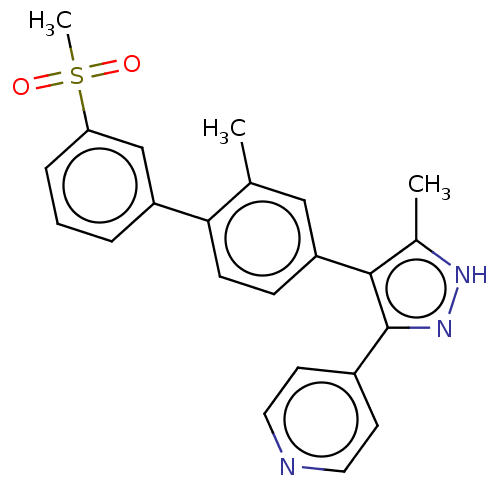
Affinity DataIC50: 1.10nMAssay Description:Inhibition of full-length LRRK2 G2019S mutant (unknown origin) using LRRKtide as substrate incubated for 1 hr in presence of ATP by TR-FRET based ADA...More data for this Ligand-Target Pair
TargetLeucine-rich repeat serine/threonine-protein kinase 2(Human)
Stanford University
Curated by ChEMBL
Stanford University
Curated by ChEMBL
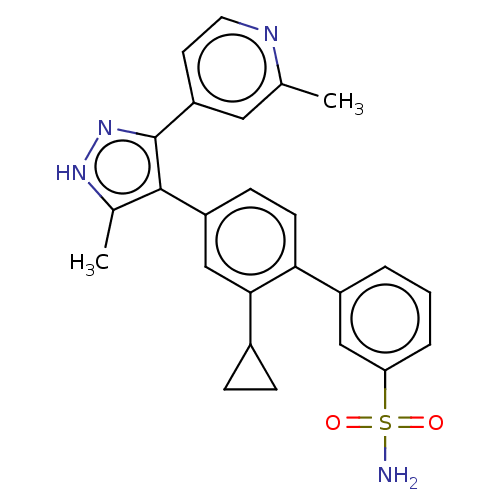
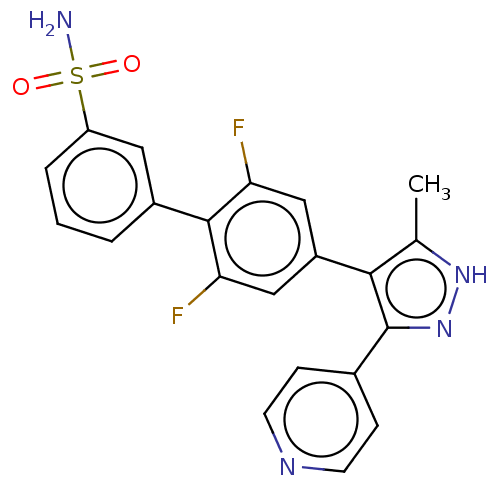
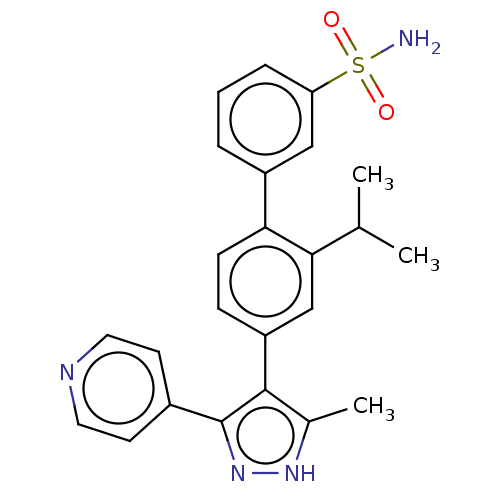
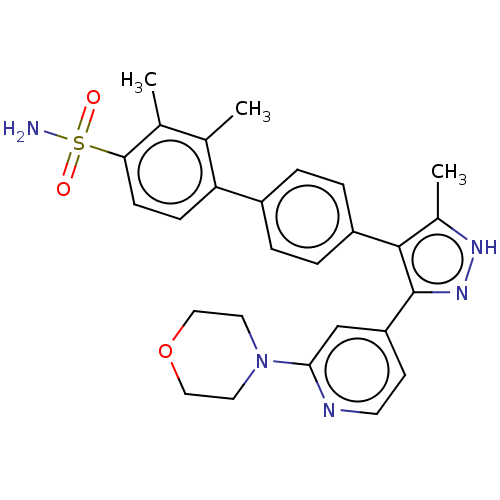
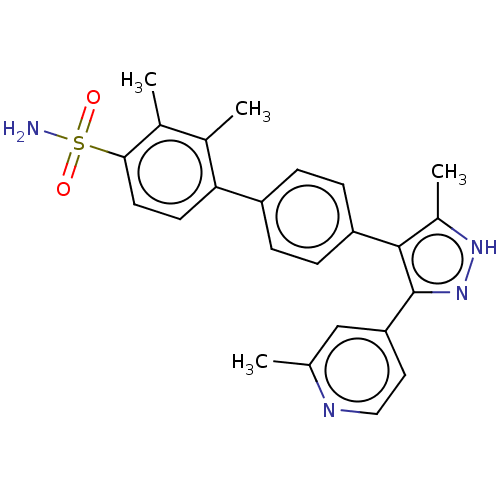
Affinity DataIC50: 2.20nMAssay Description:Inhibition of full-length LRRK2 G2019S mutant (unknown origin) using LRRKtide as substrate incubated for 1 hr in presence of ATP by TR-FRET based ADA...More data for this Ligand-Target Pair
TargetLeucine-rich repeat serine/threonine-protein kinase 2(Human)
Stanford University
Curated by ChEMBL
Stanford University
Curated by ChEMBL
Affinity DataIC50: 2.40nMAssay Description:Inhibition of full-length LRRK2 G2019S mutant (unknown origin) using LRRKtide as substrate incubated for 1 hr in presence of ATP by TR-FRET based ADA...More data for this Ligand-Target Pair
TargetLeucine-rich repeat serine/threonine-protein kinase 2(Human)
Stanford University
Curated by ChEMBL
Stanford University
Curated by ChEMBL
Affinity DataIC50: 2.90nMAssay Description:Inhibition of full-length LRRK2 G2019S mutant (unknown origin) using LRRKtide as substrate incubated for 1 hr in presence of ATP by TR-FRET based ADA...More data for this Ligand-Target Pair
TargetLeucine-rich repeat serine/threonine-protein kinase 2(Human)
Stanford University
Curated by ChEMBL
Stanford University
Curated by ChEMBL
Affinity DataIC50: 3.10nMAssay Description:Inhibition of full-length LRRK2 G2019S mutant (unknown origin) using LRRKtide as substrate incubated for 1 hr in presence of ATP by TR-FRET based ADA...More data for this Ligand-Target Pair
TargetLeucine-rich repeat serine/threonine-protein kinase 2(Human)
Stanford University
Curated by ChEMBL
Stanford University
Curated by ChEMBL
Affinity DataIC50: 4nMAssay Description:Inhibition of full-length LRRK2 G2019S mutant (unknown origin) using LRRKtide as substrate incubated for 1 hr in presence of ATP by TR-FRET based ADA...More data for this Ligand-Target Pair
TargetLeucine-rich repeat serine/threonine-protein kinase 2(Human)
Stanford University
Curated by ChEMBL
Stanford University
Curated by ChEMBL
Affinity DataIC50: 4.30nMAssay Description:Inhibition of full-length LRRK2 G2019S mutant (unknown origin) using LRRKtide as substrate incubated for 1 hr in presence of ATP by TR-FRET based ADA...More data for this Ligand-Target Pair
TargetLeucine-rich repeat serine/threonine-protein kinase 2(Human)
Stanford University
Curated by ChEMBL
Stanford University
Curated by ChEMBL
Affinity DataIC50: 6.10nMAssay Description:Inhibition of full-length LRRK2 G2019S mutant (unknown origin) using LRRKtide as substrate incubated for 1 hr in presence of ATP by TR-FRET based ADA...More data for this Ligand-Target Pair
TargetLeucine-rich repeat serine/threonine-protein kinase 2(Human)
Stanford University
Curated by ChEMBL
Stanford University
Curated by ChEMBL
Affinity DataIC50: 6.20nMAssay Description:Inhibition of full-length LRRK2 G2019S mutant (unknown origin) using LRRKtide as substrate incubated for 1 hr in presence of ATP by TR-FRET based ADA...More data for this Ligand-Target Pair
TargetLeucine-rich repeat serine/threonine-protein kinase 2(Human)
Stanford University
Curated by ChEMBL
Stanford University
Curated by ChEMBL
Affinity DataIC50: 6.20nMAssay Description:Inhibition of full-length LRRK2 G2019S mutant (unknown origin) using LRRKtide as substrate incubated for 1 hr in presence of ATP by TR-FRET based ADA...More data for this Ligand-Target Pair
TargetLeucine-rich repeat serine/threonine-protein kinase 2(Human)
Stanford University
Curated by ChEMBL
Stanford University
Curated by ChEMBL
Affinity DataIC50: 6.90nMAssay Description:Inhibition of full-length LRRK2 G2019S mutant (unknown origin) using LRRKtide as substrate incubated for 1 hr in presence of ATP by TR-FRET based ADA...More data for this Ligand-Target Pair
TargetLeucine-rich repeat serine/threonine-protein kinase 2(Human)
Stanford University
Curated by ChEMBL
Stanford University
Curated by ChEMBL
Affinity DataIC50: 7.60nMAssay Description:Inhibition of full-length LRRK2 G2019S mutant (unknown origin) using LRRKtide as substrate incubated for 1 hr in presence of ATP by TR-FRET based ADA...More data for this Ligand-Target Pair
TargetLeucine-rich repeat serine/threonine-protein kinase 2(Human)
Stanford University
Curated by ChEMBL
Stanford University
Curated by ChEMBL
Affinity DataIC50: 8.10nMAssay Description:Inhibition of full-length LRRK2 G2019S mutant (unknown origin) using LRRKtide as substrate incubated for 1 hr in presence of ATP by TR-FRET based ADA...More data for this Ligand-Target Pair
TargetLeucine-rich repeat serine/threonine-protein kinase 2(Human)
Stanford University
Curated by ChEMBL
Stanford University
Curated by ChEMBL
Affinity DataIC50: 8.10nMAssay Description:Inhibition of full-length wild type LRRK2 (unknown origin) using LRRKtide as substrate incubated for 1 hr in presence of ATP by TR-FRET based ADAPTA ...More data for this Ligand-Target Pair
TargetLeucine-rich repeat serine/threonine-protein kinase 2(Human)
Stanford University
Curated by ChEMBL
Stanford University
Curated by ChEMBL
Affinity DataIC50: 8.70nMAssay Description:Inhibition of full-length LRRK2 G2019S mutant (unknown origin) using LRRKtide as substrate incubated for 1 hr in presence of ATP by TR-FRET based ADA...More data for this Ligand-Target Pair
TargetLeucine-rich repeat serine/threonine-protein kinase 2(Human)
Stanford University
Curated by ChEMBL
Stanford University
Curated by ChEMBL
Affinity DataIC50: 15nMAssay Description:Inhibition of full-length LRRK2 G2019S mutant (unknown origin) using LRRKtide as substrate incubated for 1 hr in presence of ATP by TR-FRET based ADA...More data for this Ligand-Target Pair
TargetLeucine-rich repeat serine/threonine-protein kinase 2(Human)
Stanford University
Curated by ChEMBL
Stanford University
Curated by ChEMBL
Affinity DataIC50: 15nMAssay Description:Inhibition of full-length LRRK2 G2019S mutant (unknown origin) using LRRKtide as substrate incubated for 1 hr in presence of ATP by TR-FRET based ADA...More data for this Ligand-Target Pair
TargetLeucine-rich repeat serine/threonine-protein kinase 2(Human)
Stanford University
Curated by ChEMBL
Stanford University
Curated by ChEMBL
Affinity DataIC50: 16nMAssay Description:Inhibition of full-length LRRK2 G2019S mutant (unknown origin) using LRRKtide as substrate incubated for 1 hr in presence of ATP by TR-FRET based ADA...More data for this Ligand-Target Pair
TargetLeucine-rich repeat serine/threonine-protein kinase 2(Human)
Stanford University
Curated by ChEMBL
Stanford University
Curated by ChEMBL
Affinity DataIC50: 17nMAssay Description:Inhibition of full-length LRRK2 G2019S mutant (unknown origin) using LRRKtide as substrate incubated for 1 hr in presence of ATP by TR-FRET based ADA...More data for this Ligand-Target Pair
TargetLeucine-rich repeat serine/threonine-protein kinase 2(Human)
Stanford University
Curated by ChEMBL
Stanford University
Curated by ChEMBL
Affinity DataIC50: 17nMAssay Description:Inhibition of full-length wild type LRRK2 (unknown origin) using LRRKtide as substrate incubated for 1 hr in presence of ATP by TR-FRET based ADAPTA ...More data for this Ligand-Target Pair
TargetLeucine-rich repeat serine/threonine-protein kinase 2(Human)
Stanford University
Curated by ChEMBL
Stanford University
Curated by ChEMBL
Affinity DataIC50: 19nMAssay Description:Inhibition of full-length LRRK2 G2019S mutant (unknown origin) using LRRKtide as substrate incubated for 1 hr in presence of ATP by TR-FRET based ADA...More data for this Ligand-Target Pair
TargetLeucine-rich repeat serine/threonine-protein kinase 2(Human)
Stanford University
Curated by ChEMBL
Stanford University
Curated by ChEMBL
Affinity DataIC50: 19nMAssay Description:Inhibition of full-length LRRK2 G2019S mutant (unknown origin) using LRRKtide as substrate incubated for 1 hr in presence of ATP by TR-FRET based ADA...More data for this Ligand-Target Pair
TargetLeucine-rich repeat serine/threonine-protein kinase 2(Human)
Stanford University
Curated by ChEMBL
Stanford University
Curated by ChEMBL
Affinity DataIC50: 20nMAssay Description:Inhibition of full-length wild type LRRK2 (unknown origin) using LRRKtide as substrate incubated for 1 hr in presence of ATP by TR-FRET based ADAPTA ...More data for this Ligand-Target Pair
TargetLeucine-rich repeat serine/threonine-protein kinase 2(Human)
Stanford University
Curated by ChEMBL
Stanford University
Curated by ChEMBL
Affinity DataIC50: 21nMAssay Description:Inhibition of full-length LRRK2 G2019S mutant (unknown origin) using LRRKtide as substrate incubated for 1 hr in presence of ATP by TR-FRET based ADA...More data for this Ligand-Target Pair
TargetLeucine-rich repeat serine/threonine-protein kinase 2(Human)
Stanford University
Curated by ChEMBL
Stanford University
Curated by ChEMBL
Affinity DataIC50: 24nMAssay Description:Inhibition of full-length LRRK2 G2019S mutant (unknown origin) using LRRKtide as substrate incubated for 1 hr in presence of ATP by TR-FRET based ADA...More data for this Ligand-Target Pair
TargetLeucine-rich repeat serine/threonine-protein kinase 2(Human)
Stanford University
Curated by ChEMBL
Stanford University
Curated by ChEMBL
Affinity DataIC50: 25nMAssay Description:Inhibition of full-length wild type LRRK2 (unknown origin) using LRRKtide as substrate incubated for 1 hr in presence of ATP by TR-FRET based ADAPTA ...More data for this Ligand-Target Pair
TargetLeucine-rich repeat serine/threonine-protein kinase 2(Human)
Stanford University
Curated by ChEMBL
Stanford University
Curated by ChEMBL
Affinity DataIC50: 27nMAssay Description:Inhibition of full-length wild type LRRK2 (unknown origin) using LRRKtide as substrate incubated for 1 hr in presence of ATP by TR-FRET based ADAPTA ...More data for this Ligand-Target Pair
TargetLeucine-rich repeat serine/threonine-protein kinase 2(Mouse)
Stanford University
Curated by ChEMBL
Stanford University
Curated by ChEMBL
Affinity DataEC50: 30nMAssay Description:Inhibition of mouse full-length LRRK2 G2019S mutant transfected in HEK293 cells using LRRKtide as substrate in presence of ATP by incubated for 1 hr ...More data for this Ligand-Target Pair
TargetLeucine-rich repeat serine/threonine-protein kinase 2(Human)
Stanford University
Curated by ChEMBL
Stanford University
Curated by ChEMBL
Affinity DataIC50: 33nMAssay Description:Inhibition of full-length LRRK2 G2019S mutant (unknown origin) using LRRKtide as substrate incubated for 1 hr in presence of ATP by TR-FRET based ADA...More data for this Ligand-Target Pair
TargetLeucine-rich repeat serine/threonine-protein kinase 2(Human)
Stanford University
Curated by ChEMBL
Stanford University
Curated by ChEMBL
Affinity DataIC50: 35nMAssay Description:Inhibition of full-length LRRK2 G2019S mutant (unknown origin) using LRRKtide as substrate incubated for 1 hr in presence of ATP by TR-FRET based ADA...More data for this Ligand-Target Pair
TargetLeucine-rich repeat serine/threonine-protein kinase 2(Human)
Stanford University
Curated by ChEMBL
Stanford University
Curated by ChEMBL
Affinity DataIC50: 40nMAssay Description:Inhibition of full-length wild type LRRK2 (unknown origin) using LRRKtide as substrate incubated for 1 hr in presence of ATP by TR-FRET based ADAPTA ...More data for this Ligand-Target Pair
TargetLeucine-rich repeat serine/threonine-protein kinase 2(Human)
Stanford University
Curated by ChEMBL
Stanford University
Curated by ChEMBL
Affinity DataIC50: 44nMAssay Description:Inhibition of full-length wild type LRRK2 (unknown origin) using LRRKtide as substrate incubated for 1 hr in presence of ATP by TR-FRET based ADAPTA ...More data for this Ligand-Target Pair
TargetLeucine-rich repeat serine/threonine-protein kinase 2(Human)
Stanford University
Curated by ChEMBL
Stanford University
Curated by ChEMBL
Affinity DataIC50: 47nMAssay Description:Inhibition of full-length wild type LRRK2 (unknown origin) using LRRKtide as substrate incubated for 1 hr in presence of ATP by TR-FRET based ADAPTA ...More data for this Ligand-Target Pair
TargetLeucine-rich repeat serine/threonine-protein kinase 2(Human)
Stanford University
Curated by ChEMBL
Stanford University
Curated by ChEMBL
Affinity DataIC50: 50nMAssay Description:Inhibition of full-length LRRK2 G2019S mutant (unknown origin) using LRRKtide as substrate incubated for 1 hr in presence of ATP by TR-FRET based ADA...More data for this Ligand-Target Pair
TargetLeucine-rich repeat serine/threonine-protein kinase 2(Human)
Stanford University
Curated by ChEMBL
Stanford University
Curated by ChEMBL
Affinity DataIC50: 51nMAssay Description:Inhibition of full-length wild type LRRK2 (unknown origin) using LRRKtide as substrate incubated for 1 hr in presence of ATP by TR-FRET based ADAPTA ...More data for this Ligand-Target Pair
TargetLeucine-rich repeat serine/threonine-protein kinase 2(Human)
Stanford University
Curated by ChEMBL
Stanford University
Curated by ChEMBL
Affinity DataIC50: 57nMAssay Description:Inhibition of full-length LRRK2 G2019S mutant (unknown origin) using LRRKtide as substrate incubated for 1 hr in presence of ATP by TR-FRET based ADA...More data for this Ligand-Target Pair
TargetLeucine-rich repeat serine/threonine-protein kinase 2(Human)
Stanford University
Curated by ChEMBL
Stanford University
Curated by ChEMBL
Affinity DataIC50: 61nMAssay Description:Inhibition of full-length LRRK2 G2019S mutant (unknown origin) using LRRKtide as substrate incubated for 1 hr in presence of ATP by TR-FRET based ADA...More data for this Ligand-Target Pair
TargetLeucine-rich repeat serine/threonine-protein kinase 2(Human)
Stanford University
Curated by ChEMBL
Stanford University
Curated by ChEMBL
Affinity DataIC50: 63nMAssay Description:Inhibition of full-length wild type LRRK2 (unknown origin) using LRRKtide as substrate incubated for 1 hr in presence of ATP by TR-FRET based ADAPTA ...More data for this Ligand-Target Pair
TargetLeucine-rich repeat serine/threonine-protein kinase 2(Human)
Stanford University
Curated by ChEMBL
Stanford University
Curated by ChEMBL
Affinity DataIC50: 66nMAssay Description:Inhibition of full-length wild type LRRK2 (unknown origin) using LRRKtide as substrate incubated for 1 hr in presence of ATP by TR-FRET based ADAPTA ...More data for this Ligand-Target Pair
TargetLeucine-rich repeat serine/threonine-protein kinase 2(Human)
Stanford University
Curated by ChEMBL
Stanford University
Curated by ChEMBL
Affinity DataIC50: 67nMAssay Description:Inhibition of full-length wild type LRRK2 (unknown origin) using LRRKtide as substrate incubated for 1 hr in presence of ATP by TR-FRET based ADAPTA ...More data for this Ligand-Target Pair
TargetLeucine-rich repeat serine/threonine-protein kinase 2(Human)
Stanford University
Curated by ChEMBL
Stanford University
Curated by ChEMBL
Affinity DataIC50: 70nMAssay Description:Inhibition of full-length wild type LRRK2 (unknown origin) using LRRKtide as substrate incubated for 1 hr in presence of ATP by TR-FRET based ADAPTA ...More data for this Ligand-Target Pair
TargetLeucine-rich repeat serine/threonine-protein kinase 2(Human)
Stanford University
Curated by ChEMBL
Stanford University
Curated by ChEMBL
Affinity DataIC50: 73nMAssay Description:Inhibition of full-length wild type LRRK2 (unknown origin) using LRRKtide as substrate incubated for 1 hr in presence of ATP by TR-FRET based ADAPTA ...More data for this Ligand-Target Pair
TargetLeucine-rich repeat serine/threonine-protein kinase 2(Human)
Stanford University
Curated by ChEMBL
Stanford University
Curated by ChEMBL
Affinity DataIC50: 76nMAssay Description:Inhibition of full-length LRRK2 G2019S mutant (unknown origin) using LRRKtide as substrate incubated for 1 hr in presence of ATP by TR-FRET based ADA...More data for this Ligand-Target Pair
TargetLeucine-rich repeat serine/threonine-protein kinase 2(Human)
Stanford University
Curated by ChEMBL
Stanford University
Curated by ChEMBL
Affinity DataIC50: 78nMAssay Description:Inhibition of full-length wild type LRRK2 (unknown origin) using LRRKtide as substrate incubated for 1 hr in presence of ATP by TR-FRET based ADAPTA ...More data for this Ligand-Target Pair
TargetLeucine-rich repeat serine/threonine-protein kinase 2(Human)
Stanford University
Curated by ChEMBL
Stanford University
Curated by ChEMBL
Affinity DataIC50: 88nMAssay Description:Inhibition of full-length LRRK2 G2019S mutant (unknown origin) using LRRKtide as substrate incubated for 1 hr in presence of ATP by TR-FRET based ADA...More data for this Ligand-Target Pair
TargetLeucine-rich repeat serine/threonine-protein kinase 2(Human)
Stanford University
Curated by ChEMBL
Stanford University
Curated by ChEMBL
Affinity DataIC50: 89nMAssay Description:Inhibition of full-length wild type LRRK2 (unknown origin) using LRRKtide as substrate incubated for 1 hr in presence of ATP by TR-FRET based ADAPTA ...More data for this Ligand-Target Pair
TargetLeucine-rich repeat serine/threonine-protein kinase 2(Human)
Stanford University
Curated by ChEMBL
Stanford University
Curated by ChEMBL
Affinity DataIC50: 96nMAssay Description:Inhibition of full-length LRRK2 G2019S mutant (unknown origin) using LRRKtide as substrate incubated for 1 hr in presence of ATP by TR-FRET based ADA...More data for this Ligand-Target Pair
TargetLeucine-rich repeat serine/threonine-protein kinase 2(Human)
Stanford University
Curated by ChEMBL
Stanford University
Curated by ChEMBL
Affinity DataIC50: 101nMAssay Description:Inhibition of full-length LRRK2 G2019S mutant (unknown origin) using LRRKtide as substrate incubated for 1 hr in presence of ATP by TR-FRET based ADA...More data for this Ligand-Target Pair