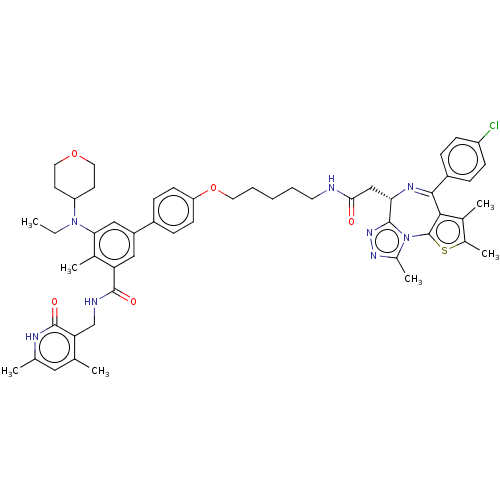
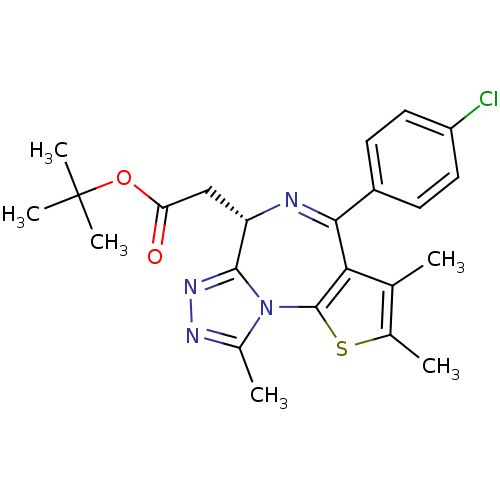
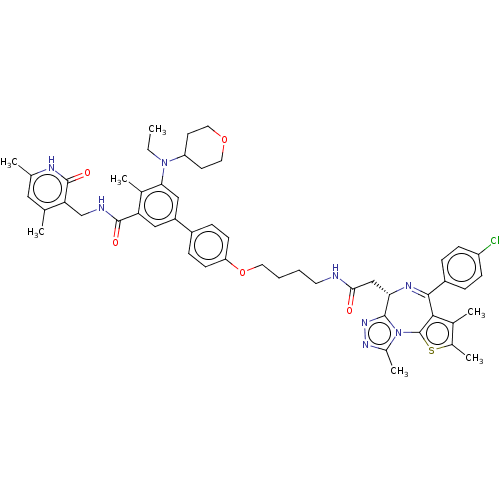
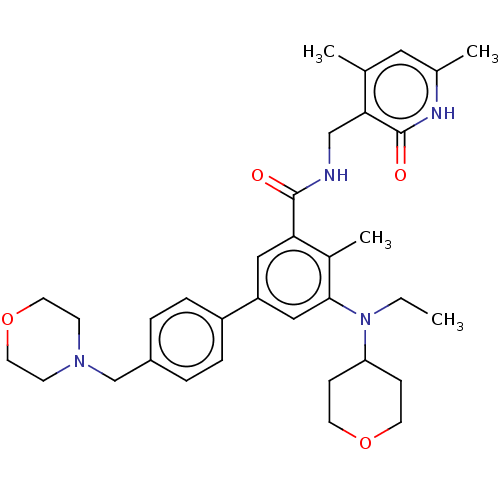
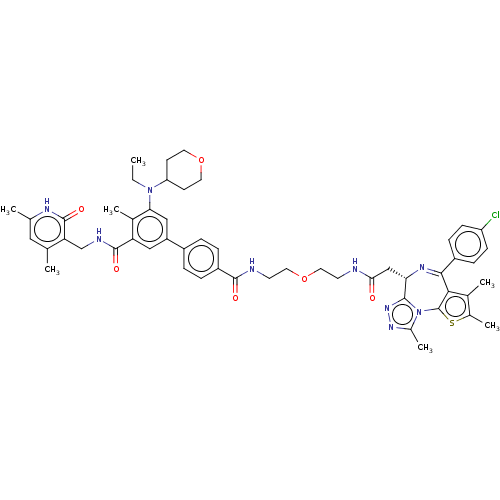
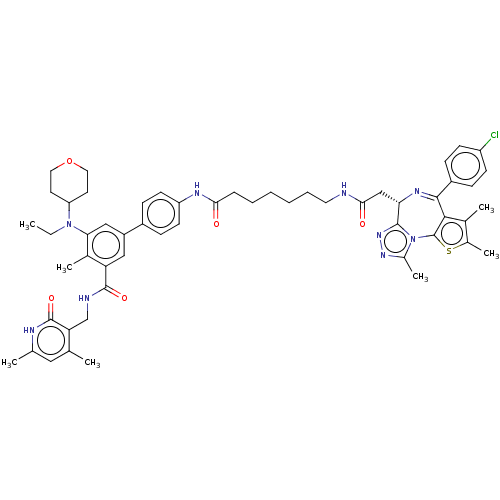
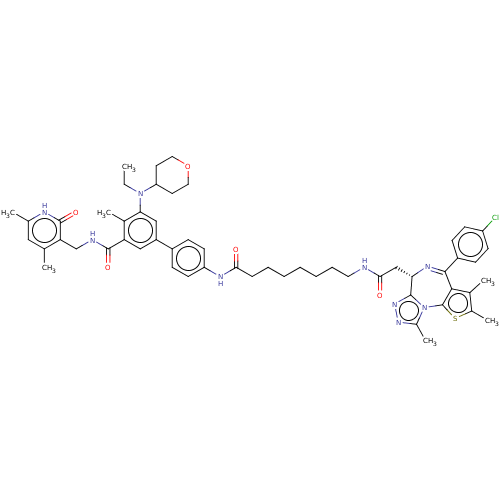
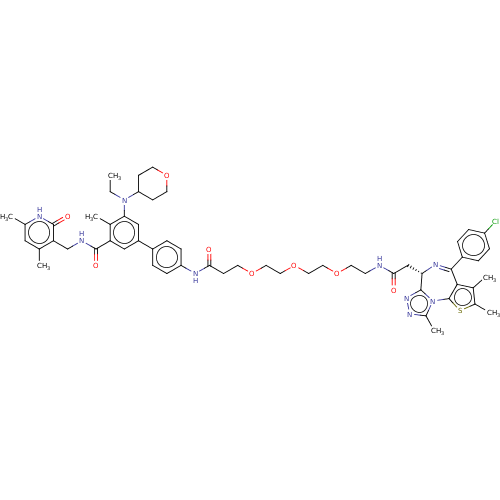
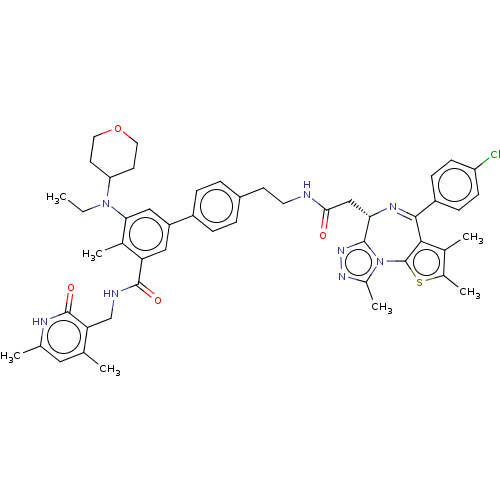
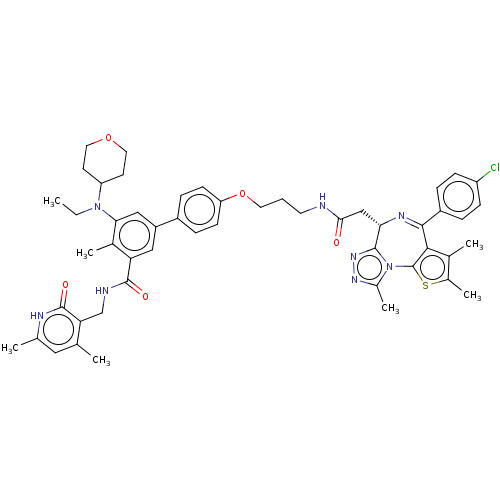
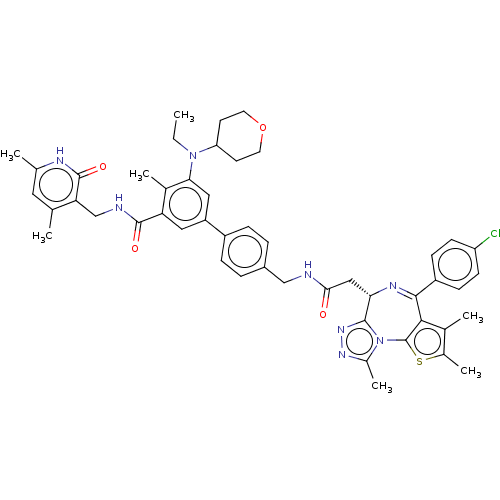
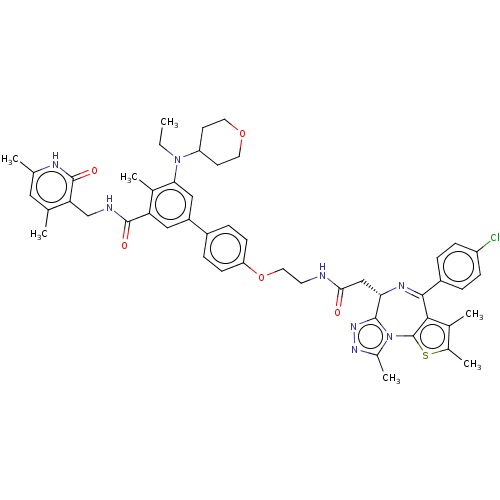
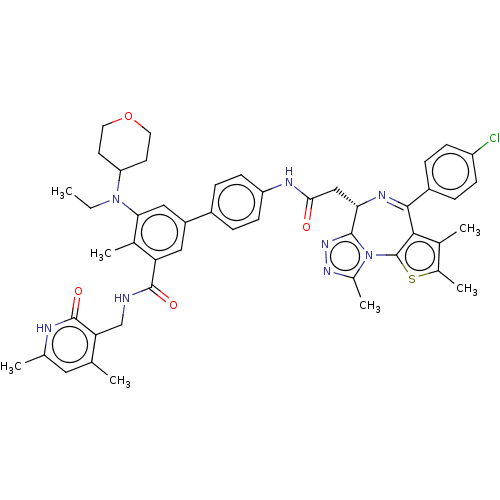
Report error Found 24 Enz. Inhib. hit(s) with all data for entry = 50017866
Affinity DataIC50: 34nMAssay Description:Displacement of fluorescein-labeled JQ1 from His6/TEV cleavage site fused human BRD4 (residues N44 to E168) expressed in Escherichia coli BL21 (DE3) ...More data for this Ligand-Target Pair
Affinity DataIC50: 68nMAssay Description:Displacement of fluorescein-labeled JQ1 from His6/TEV cleavage site fused human BRD4 (residues N44 to E168) expressed in Escherichia coli BL21 (DE3) ...More data for this Ligand-Target Pair
Affinity DataIC50: 82nMAssay Description:Displacement of fluorescein-labeled JQ1 from His6/TEV cleavage site fused human BRD4 (residues N44 to E168) expressed in Escherichia coli BL21 (DE3) ...More data for this Ligand-Target Pair
TargetHistone-lysine N-methyltransferase EZH2(Human)
Sun Yat-Sen University Cancer Center
Curated by ChEMBL
Sun Yat-Sen University Cancer Center
Curated by ChEMBL
Affinity DataIC50: 150nMAssay Description:Inhibition of EZH2 (unknown origin) using H3-derivede peptide (21 to 44) as substrate incubated for 60 min in presence of SAM by MTase-Glo assayMore data for this Ligand-Target Pair
Affinity DataIC50: 248nMAssay Description:Displacement of fluorescein-labeled JQ1 from His6/TEV cleavage site fused human BRD4 (residues N44 to E168) expressed in Escherichia coli BL21 (DE3) ...More data for this Ligand-Target Pair
TargetHistone-lysine N-methyltransferase EZH2(Human)
Sun Yat-Sen University Cancer Center
Curated by ChEMBL
Sun Yat-Sen University Cancer Center
Curated by ChEMBL
Affinity DataIC50: 300nMAssay Description:Inhibition of EZH2 (unknown origin) using H3-derivede peptide (21 to 44) as substrate incubated for 60 min in presence of SAM by MTase-Glo assayMore data for this Ligand-Target Pair
TargetHistone-lysine N-methyltransferase EZH2(Human)
Sun Yat-Sen University Cancer Center
Curated by ChEMBL
Sun Yat-Sen University Cancer Center
Curated by ChEMBL
Affinity DataIC50: 420nMAssay Description:Inhibition of EZH2 (unknown origin) using H3-derivede peptide (21 to 44) as substrate incubated for 60 min in presence of SAM by MTase-Glo assayMore data for this Ligand-Target Pair
TargetHistone-lysine N-methyltransferase EZH2(Human)
Sun Yat-Sen University Cancer Center
Curated by ChEMBL
Sun Yat-Sen University Cancer Center
Curated by ChEMBL
Affinity DataIC50: 490nMAssay Description:Inhibition of EZH2 (unknown origin) using H3-derivede peptide (21 to 44) as substrate incubated for 60 min in presence of SAM by MTase-Glo assayMore data for this Ligand-Target Pair
TargetHistone-lysine N-methyltransferase EZH2(Human)
Sun Yat-Sen University Cancer Center
Curated by ChEMBL
Sun Yat-Sen University Cancer Center
Curated by ChEMBL
Affinity DataIC50: 530nMAssay Description:Inhibition of EZH2 (unknown origin) using H3-derivede peptide (21 to 44) as substrate incubated for 60 min in presence of SAM by MTase-Glo assayMore data for this Ligand-Target Pair
TargetHistone-lysine N-methyltransferase EZH2(Human)
Sun Yat-Sen University Cancer Center
Curated by ChEMBL
Sun Yat-Sen University Cancer Center
Curated by ChEMBL
Affinity DataIC50: 550nMAssay Description:Inhibition of EZH2 (unknown origin) using H3-derivede peptide (21 to 44) as substrate incubated for 60 min in presence of SAM by MTase-Glo assayMore data for this Ligand-Target Pair
Affinity DataIC50: 580nMAssay Description:Displacement of fluorescein-labeled JQ1 from His6/TEV cleavage site fused human BRD4 (residues N44 to E168) expressed in Escherichia coli BL21 (DE3) ...More data for this Ligand-Target Pair
Affinity DataIC50: 604nMAssay Description:Displacement of fluorescein-labeled JQ1 from His6/TEV cleavage site fused human BRD4 (residues N44 to E168) expressed in Escherichia coli BL21 (DE3) ...More data for this Ligand-Target Pair
Affinity DataIC50: 945nMAssay Description:Displacement of fluorescein-labeled JQ1 from His6/TEV cleavage site fused human BRD4 (residues N44 to E168) expressed in Escherichia coli BL21 (DE3) ...More data for this Ligand-Target Pair
TargetHistone-lysine N-methyltransferase EZH2(Human)
Sun Yat-Sen University Cancer Center
Curated by ChEMBL
Sun Yat-Sen University Cancer Center
Curated by ChEMBL
Affinity DataIC50: 980nMAssay Description:Inhibition of EZH2 (unknown origin) using H3-derivede peptide (21 to 44) as substrate incubated for 60 min in presence of SAM by MTase-Glo assayMore data for this Ligand-Target Pair
TargetHistone-lysine N-methyltransferase EZH2(Human)
Sun Yat-Sen University Cancer Center
Curated by ChEMBL
Sun Yat-Sen University Cancer Center
Curated by ChEMBL
Affinity DataIC50: 1.04E+3nMAssay Description:Inhibition of EZH2 (unknown origin) using H3-derivede peptide (21 to 44) as substrate incubated for 60 min in presence of SAM by MTase-Glo assayMore data for this Ligand-Target Pair
Affinity DataIC50: 1.48E+3nMAssay Description:Displacement of fluorescein-labeled JQ1 from His6/TEV cleavage site fused human BRD4 (residues N44 to E168) expressed in Escherichia coli BL21 (DE3) ...More data for this Ligand-Target Pair
Affinity DataIC50: 1.54E+3nMAssay Description:Displacement of fluorescein-labeled JQ1 from His6/TEV cleavage site fused human BRD4 (residues N44 to E168) expressed in Escherichia coli BL21 (DE3) ...More data for this Ligand-Target Pair
Affinity DataIC50: 1.72E+3nMAssay Description:Displacement of fluorescein-labeled JQ1 from His6/TEV cleavage site fused human BRD4 (residues N44 to E168) expressed in Escherichia coli BL21 (DE3) ...More data for this Ligand-Target Pair
Affinity DataIC50: 2.34E+3nMAssay Description:Displacement of fluorescein-labeled JQ1 from His6/TEV cleavage site fused human BRD4 (residues N44 to E168) expressed in Escherichia coli BL21 (DE3) ...More data for this Ligand-Target Pair
Affinity DataIC50: 2.84E+3nMAssay Description:Displacement of fluorescein-labeled JQ1 from His6/TEV cleavage site fused human BRD4 (residues N44 to E168) expressed in Escherichia coli BL21 (DE3) ...More data for this Ligand-Target Pair
TargetHistone-lysine N-methyltransferase EZH2(Human)
Sun Yat-Sen University Cancer Center
Curated by ChEMBL
Sun Yat-Sen University Cancer Center
Curated by ChEMBL
Affinity DataIC50: 7.55E+3nMAssay Description:Inhibition of EZH2 (unknown origin) using H3-derivede peptide (21 to 44) as substrate incubated for 60 min in presence of SAM by MTase-Glo assayMore data for this Ligand-Target Pair
TargetHistone-lysine N-methyltransferase EZH2(Human)
Sun Yat-Sen University Cancer Center
Curated by ChEMBL
Sun Yat-Sen University Cancer Center
Curated by ChEMBL
Affinity DataIC50: 1.61E+4nMAssay Description:Inhibition of EZH2 (unknown origin) using H3-derivede peptide (21 to 44) as substrate incubated for 60 min in presence of SAM by MTase-Glo assayMore data for this Ligand-Target Pair
TargetHistone-lysine N-methyltransferase EZH2(Human)
Sun Yat-Sen University Cancer Center
Curated by ChEMBL
Sun Yat-Sen University Cancer Center
Curated by ChEMBL
Affinity DataIC50: 2.98E+4nMAssay Description:Inhibition of EZH2 (unknown origin) using H3-derivede peptide (21 to 44) as substrate incubated for 60 min in presence of SAM by MTase-Glo assayMore data for this Ligand-Target Pair
TargetHistone-lysine N-methyltransferase EZH2(Human)
Sun Yat-Sen University Cancer Center
Curated by ChEMBL
Sun Yat-Sen University Cancer Center
Curated by ChEMBL
Affinity DataIC50: 7.78E+4nMAssay Description:Inhibition of EZH2 (unknown origin) using H3-derivede peptide (21 to 44) as substrate incubated for 60 min in presence of SAM by MTase-Glo assayMore data for this Ligand-Target Pair