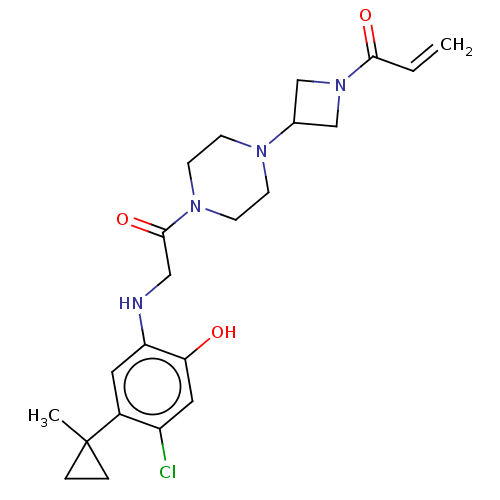
Report error Found 31 Enz. Inhib. hit(s) with all data for entry = 50015318
Affinity DataIC50: 2nMAssay Description:Inhibition of KRAS G12D mutant in human AGS cells assessed as reduction in ERK phosphorylation measured after 3 hrs by In-Cell Western assayMore data for this Ligand-Target Pair
Affinity DataIC50: 5nMAssay Description:Inhibition of human GDP-bound KRAS G12D mutant assessed as inhibition of SOS1-catalyzed nucleotide exchange by measuring KRAS G12D mutant/GTP/c-Raf1 ...More data for this Ligand-Target Pair
Affinity DataIC50: 14nMAssay Description:Inhibition of KRAS G12D mutant in human AGS cells assessed as reduction in ERK phosphorylation measured after 3 hrs by In-Cell Western assayMore data for this Ligand-Target Pair
Affinity DataIC50: 24nMAssay Description:Inhibition of KRAS G12D mutant in human AGS cells assessed as reduction in ERK phosphorylation measured after 3 hrs by In-Cell Western assayMore data for this Ligand-Target Pair
Affinity DataIC50: 24nMAssay Description:Inhibition of KRAS G12D mutant in human AGS cells assessed as reduction in ERK phosphorylation measured after 3 hrs by In-Cell Western assayMore data for this Ligand-Target Pair
Affinity DataIC50: 63nMAssay Description:Inhibition of KRAS G12D mutant in human AGS cells assessed as reduction in ERK phosphorylation measured after 3 hrs by In-Cell Western assayMore data for this Ligand-Target Pair
Affinity DataIC50: 64nMAssay Description:Inhibition of KRAS G12D mutant in human AGS cells assessed as reduction in ERK phosphorylation measured after 3 hrs by In-Cell Western assayMore data for this Ligand-Target Pair
Affinity DataIC50: 71nMAssay Description:Inhibition of KRAS G12D mutant in human AGS cells assessed as reduction in ERK phosphorylation measured after 3 hrs by In-Cell Western assayMore data for this Ligand-Target Pair
Affinity DataIC50: 97nMAssay Description:Inhibition of KRAS G12D mutant in human AGS cells assessed as reduction in ERK phosphorylation measured after 3 hrs by In-Cell Western assayMore data for this Ligand-Target Pair
Affinity DataKd: 182nMAssay Description:Binding affinity to human AviTEV6His-tagged KRAS isoform 4B wild type (1 to 169 residues) expressed in Escherichia coli BL21 (DE3) assessed as dissoc...More data for this Ligand-Target Pair
Affinity DataIC50: 190nMAssay Description:Inhibition of KRAS G12D mutant in human AGS cells assessed as reduction in ERK phosphorylation measured after 3 hrs by In-Cell Western assayMore data for this Ligand-Target Pair
Affinity DataIC50: 200nMAssay Description:Inhibition of KRAS G12D mutant in human AGS cells assessed as reduction in ERK phosphorylation measured after 3 hrs by In-Cell Western assayMore data for this Ligand-Target Pair
Affinity DataIC50: 270nMAssay Description:Inhibition of KRAS G12D mutant in human AGS cells assessed as reduction in ERK phosphorylation measured after 3 hrs by In-Cell Western assayMore data for this Ligand-Target Pair
Affinity DataIC50: 370nMAssay Description:Inhibition of KRAS G12D mutant in human AGS cells assessed as reduction in ERK phosphorylation measured after 3 hrs by In-Cell Western assayMore data for this Ligand-Target Pair
Affinity DataIC50: 380nMAssay Description:Inhibition of KRAS G12D mutant in human AGS cells assessed as reduction in ERK phosphorylation measured after 3 hrs by In-Cell Western assayMore data for this Ligand-Target Pair
Affinity DataIC50: 530nMAssay Description:Inhibition of KRAS G12D mutant in human AGS cells assessed as reduction in ERK phosphorylation measured after 3 hrs by In-Cell Western assayMore data for this Ligand-Target Pair
Affinity DataKd: 560nMAssay Description:Binding affinity to human AviTEV6His-tagged KRAS isoform 4B wild type (1 to 169 residues) expressed in Escherichia coli BL21 (DE3) assessed as dissoc...More data for this Ligand-Target Pair
Affinity DataIC50: 650nMAssay Description:Inhibition of KRAS G12D mutant in human AGS cells assessed as reduction in ERK phosphorylation measured after 3 hrs by In-Cell Western assayMore data for this Ligand-Target Pair
Affinity DataIC50: 1.20E+3nMAssay Description:Inhibition of KRAS G12D mutant in human AGS cells assessed as reduction in ERK phosphorylation measured after 3 hrs by In-Cell Western assayMore data for this Ligand-Target Pair
Affinity DataIC50: 1.30E+3nMAssay Description:Inhibition of KRAS G12D mutant in human AGS cells assessed as reduction in ERK phosphorylation measured after 3 hrs by In-Cell Western assayMore data for this Ligand-Target Pair
Affinity DataIC50: 2.50E+3nMAssay Description:Inhibition of KRAS G12D mutant in human AGS cells assessed as reduction in ERK phosphorylation measured after 3 hrs by In-Cell Western assayMore data for this Ligand-Target Pair
Affinity DataIC50: 3.00E+3nMAssay Description:Inhibition of KRAS G12D mutant in human AGS cells assessed as reduction in ERK phosphorylation measured after 3 hrs by In-Cell Western assayMore data for this Ligand-Target Pair
Affinity DataIC50: 3.50E+3nMAssay Description:Inhibition of KRAS G12D mutant in human AGS cells assessed as reduction in ERK phosphorylation measured after 3 hrs by In-Cell Western assayMore data for this Ligand-Target Pair
Affinity DataKd: 6.20E+3nMAssay Description:Binding affinity to human AviTEV6His-tagged KRAS isoform 4B wild type (1 to 169 residues) expressed in Escherichia coli BL21 (DE3) assessed as dissoc...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMAssay Description:Inhibition of KRAS G12D mutant in human AGS cells assessed as reduction in ERK phosphorylation measured after 3 hrs by In-Cell Western assayMore data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMAssay Description:Inhibition of KRAS G12D mutant in human AGS cells assessed as reduction in ERK phosphorylation measured after 3 hrs by In-Cell Western assayMore data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMAssay Description:Inhibition of KRAS G12D mutant in human AGS cells assessed as reduction in ERK phosphorylation measured after 3 hrs by In-Cell Western assayMore data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMAssay Description:Inhibition of KRAS G12D mutant in human AGS cells assessed as reduction in ERK phosphorylation measured after 3 hrs by In-Cell Western assayMore data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMAssay Description:Inhibition of KRAS G12D mutant in human AGS cells assessed as reduction in ERK phosphorylation measured after 3 hrs by In-Cell Western assayMore data for this Ligand-Target Pair
Affinity DataKd: 3.60E+4nMAssay Description:Binding affinity to human AviTEV6His-tagged KRAS isoform 4B wild type (1 to 169 residues) expressed in Escherichia coli BL21 (DE3) assessed as dissoc...More data for this Ligand-Target Pair
Affinity DataKi: 2.00E+5nMAssay Description:Inhibition of KRAS G12C mutant (unknown origin)More data for this Ligand-Target Pair