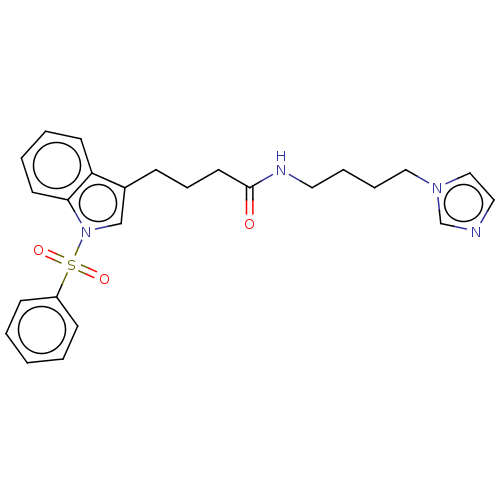
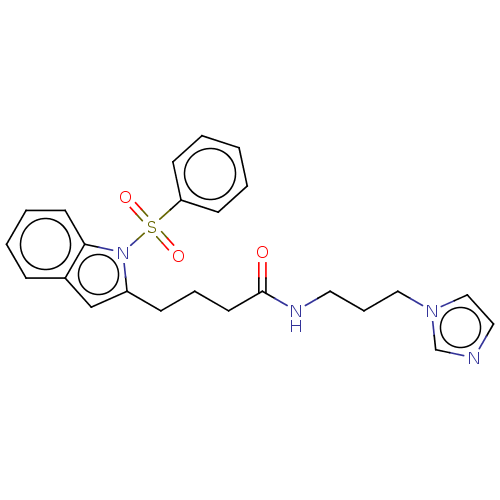
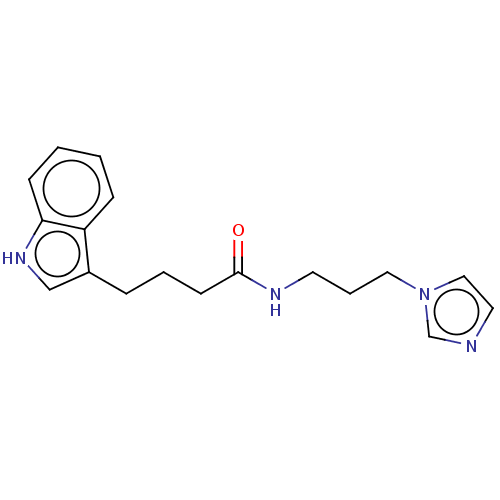
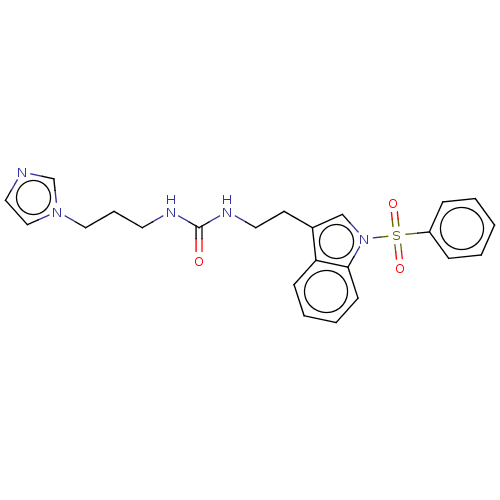
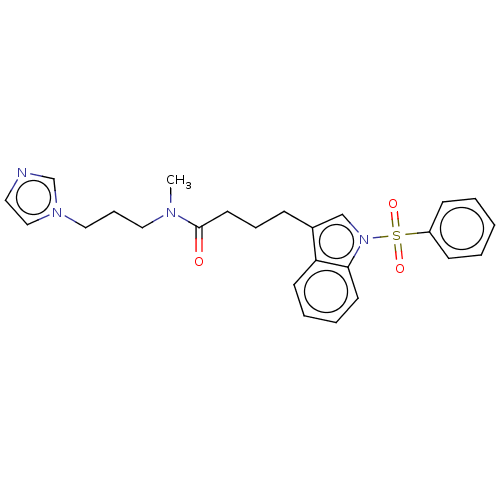
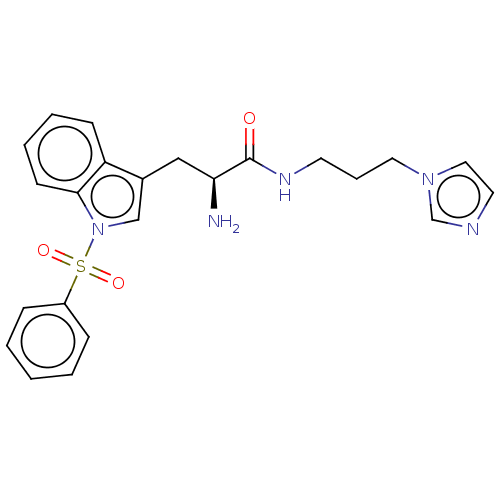
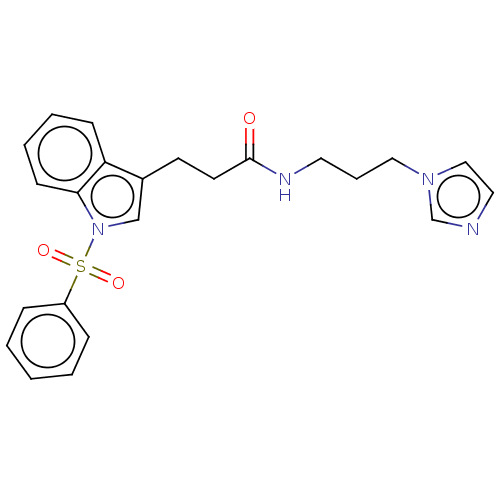
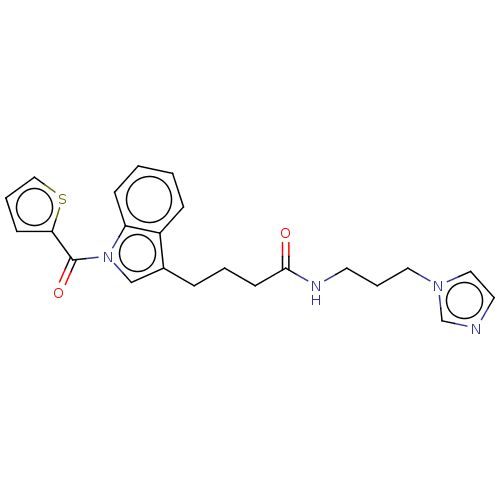
Report error Found 22 Enz. Inhib. hit(s) with all data for entry = 50017541
Affinity DataEC50: 5.01E+3nMAssay Description:Activation of human recombinant IDE using ATTO 655-Cys-Lys-Leu-Val-Phe-Phe-Ala-Glu-Asp-trp as substrate incubated for 10 mins followed by substrate a...More data for this Ligand-Target Pair
Affinity DataEC50: 5.50E+3nMAssay Description:Activation of human recombinant IDE using ATTO 655-Cys-Lys-Leu-Val-Phe-Phe-Ala-Glu-Asp-trp as substrate incubated for 10 mins followed by substrate a...More data for this Ligand-Target Pair
Affinity DataEC50: 6.76E+3nMAssay Description:Activation of human recombinant IDE using ATTO 655-Cys-Lys-Leu-Val-Phe-Phe-Ala-Glu-Asp-trp as substrate incubated for 10 mins followed by substrate a...More data for this Ligand-Target Pair
Affinity DataEC50: 7.24E+3nMAssay Description:Activation of human recombinant IDE using ATTO 655-Cys-Lys-Leu-Val-Phe-Phe-Ala-Glu-Asp-trp as substrate incubated for 10 mins followed by substrate a...More data for this Ligand-Target Pair
Affinity DataEC50: 7.94E+3nMAssay Description:Activation of human recombinant IDE using ATTO 655-Cys-Lys-Leu-Val-Phe-Phe-Ala-Glu-Asp-trp as substrate incubated for 10 mins followed by substrate a...More data for this Ligand-Target Pair
Affinity DataEC50: 8.13E+3nMAssay Description:Activation of human recombinant IDE using ATTO 655-Cys-Lys-Leu-Val-Phe-Phe-Ala-Glu-Asp-trp as substrate incubated for 10 mins followed by substrate a...More data for this Ligand-Target Pair
Affinity DataEC50: 8.13E+3nMAssay Description:Activation of human recombinant IDE using ATTO 655-Cys-Lys-Leu-Val-Phe-Phe-Ala-Glu-Asp-trp as substrate incubated for 10 mins followed by substrate a...More data for this Ligand-Target Pair
Affinity DataEC50: 1.00E+4nMAssay Description:Activation of human recombinant IDE using ATTO 655-Cys-Lys-Leu-Val-Phe-Phe-Ala-Glu-Asp-trp as substrate incubated for 10 mins followed by substrate a...More data for this Ligand-Target Pair
Affinity DataEC50: 1.00E+4nMAssay Description:Activation of human recombinant IDE using ATTO 655-Cys-Lys-Leu-Val-Phe-Phe-Ala-Glu-Asp-trp as substrate incubated for 10 mins followed by substrate a...More data for this Ligand-Target Pair
Affinity DataEC50: 1.02E+4nMAssay Description:Activation of human recombinant IDE using ATTO 655-Cys-Lys-Leu-Val-Phe-Phe-Ala-Glu-Asp-trp as substrate incubated for 10 mins followed by substrate a...More data for this Ligand-Target Pair
Affinity DataEC50: 1.02E+4nMAssay Description:Activation of human recombinant IDE using ATTO 655-Cys-Lys-Leu-Val-Phe-Phe-Ala-Glu-Asp-trp as substrate incubated for 10 mins followed by substrate a...More data for this Ligand-Target Pair
Affinity DataEC50: 1.23E+4nMAssay Description:Activation of human recombinant IDE using ATTO 655-Cys-Lys-Leu-Val-Phe-Phe-Ala-Glu-Asp-trp as substrate incubated for 10 mins followed by substrate a...More data for this Ligand-Target Pair
Affinity DataEC50: 1.32E+4nMAssay Description:Activation of human recombinant IDE using ATTO 655-Cys-Lys-Leu-Val-Phe-Phe-Ala-Glu-Asp-trp as substrate incubated for 10 mins followed by substrate a...More data for this Ligand-Target Pair
Affinity DataEC50: 1.38E+4nMAssay Description:Activation of human recombinant IDE using ATTO 655-Cys-Lys-Leu-Val-Phe-Phe-Ala-Glu-Asp-trp as substrate incubated for 10 mins followed by substrate a...More data for this Ligand-Target Pair
Affinity DataEC50: 1.41E+4nMAssay Description:Activation of human recombinant IDE using ATTO 655-Cys-Lys-Leu-Val-Phe-Phe-Ala-Glu-Asp-trp as substrate incubated for 10 mins followed by substrate a...More data for this Ligand-Target Pair
Affinity DataEC50: 1.45E+4nMAssay Description:Activation of human recombinant IDE using ATTO 655-Cys-Lys-Leu-Val-Phe-Phe-Ala-Glu-Asp-trp as substrate incubated for 10 mins followed by substrate a...More data for this Ligand-Target Pair
Affinity DataEC50: 1.55E+4nMAssay Description:Activation of human recombinant IDE using ATTO 655-Cys-Lys-Leu-Val-Phe-Phe-Ala-Glu-Asp-trp as substrate incubated for 10 mins followed by substrate a...More data for this Ligand-Target Pair
Affinity DataEC50: 2.14E+4nMAssay Description:Activation of human recombinant IDE using ATTO 655-Cys-Lys-Leu-Val-Phe-Phe-Ala-Glu-Asp-trp as substrate incubated for 10 mins followed by substrate a...More data for this Ligand-Target Pair
Affinity DataEC50: 2.69E+4nMAssay Description:Activation of human recombinant IDE using ATTO 655-Cys-Lys-Leu-Val-Phe-Phe-Ala-Glu-Asp-trp as substrate incubated for 10 mins followed by substrate a...More data for this Ligand-Target Pair
Affinity DataEC50: 2.69E+4nMAssay Description:Activation of human recombinant IDE using ATTO 655-Cys-Lys-Leu-Val-Phe-Phe-Ala-Glu-Asp-trp as substrate incubated for 10 mins followed by substrate a...More data for this Ligand-Target Pair
Affinity DataEC50: 2.88E+4nMAssay Description:Activation of human recombinant IDE using ATTO 655-Cys-Lys-Leu-Val-Phe-Phe-Ala-Glu-Asp-trp as substrate incubated for 10 mins followed by substrate a...More data for this Ligand-Target Pair
Affinity DataEC50: 3.89E+4nMAssay Description:Activation of human recombinant IDE using ATTO 655-Cys-Lys-Leu-Val-Phe-Phe-Ala-Glu-Asp-trp as substrate incubated for 10 mins followed by substrate a...More data for this Ligand-Target Pair