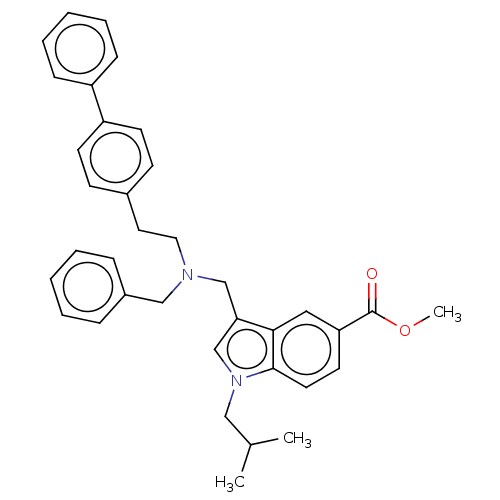
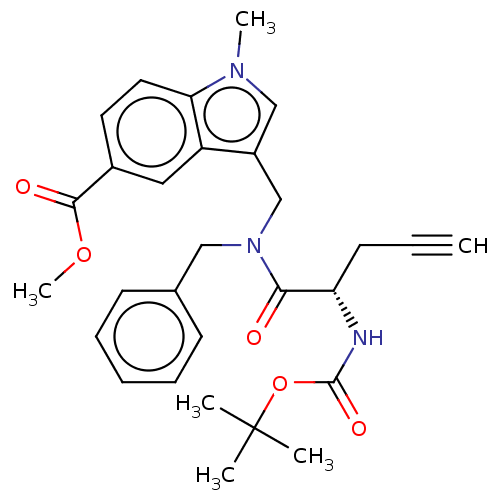
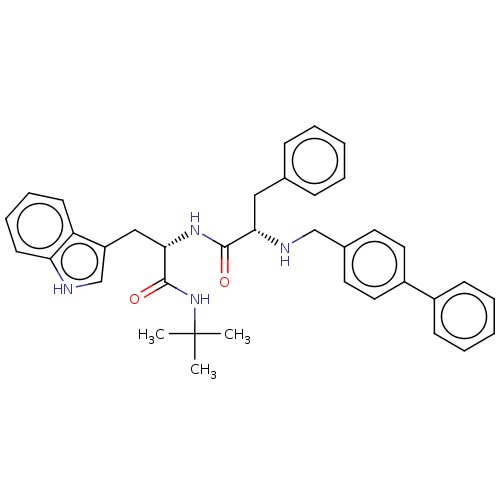
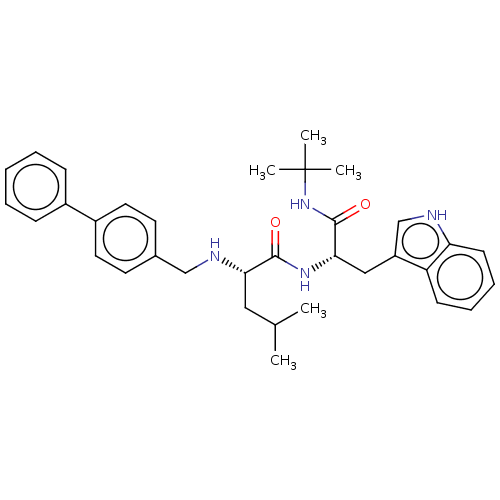
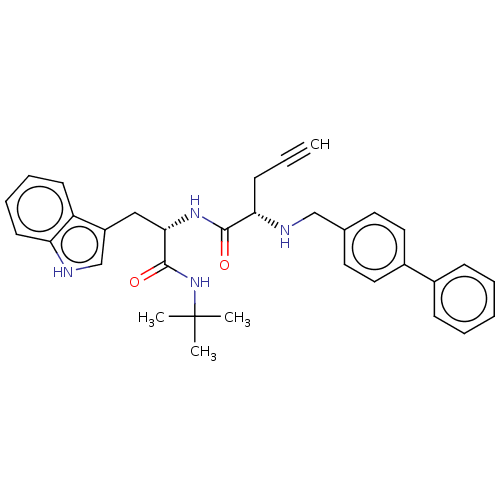
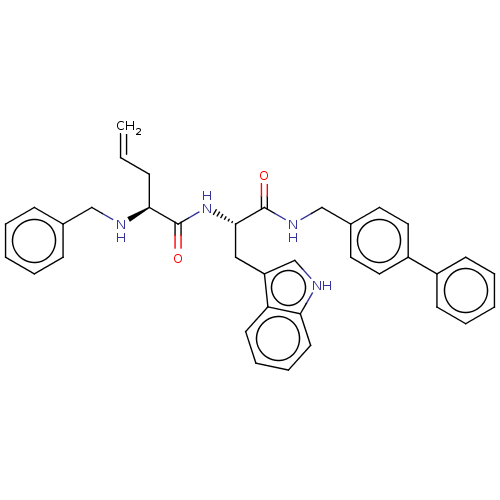
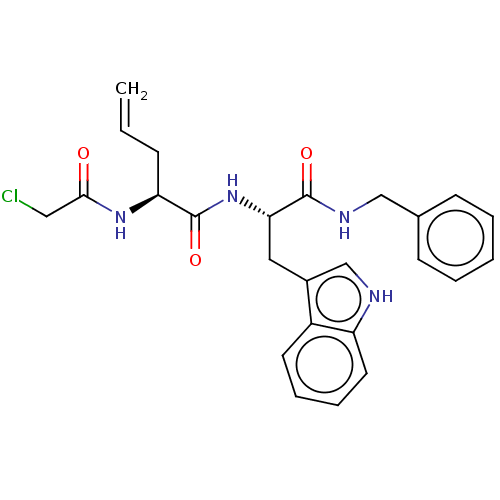
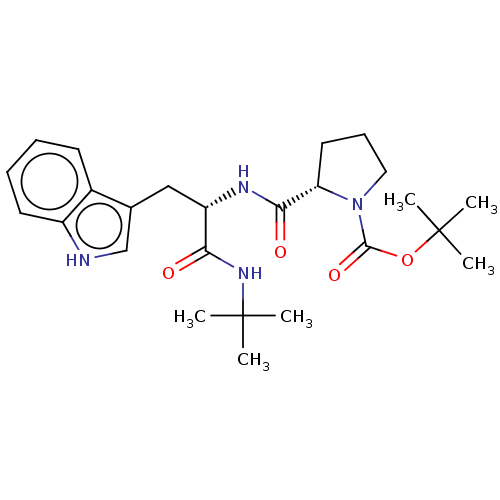
Report error Found 8 Enz. Inhib. hit(s) with all data for entry = 50015027
Affinity DataKd: 3.26E+3nMAssay Description:Binding affinity to CM5 sensor chip immobilized SARS-CoV-2 spike glycoprotein assessed as dissociation constant by SPR analysisMore data for this Ligand-Target Pair
Affinity DataKd: 1.14E+4nMAssay Description:Binding affinity to CM5 sensor chip immobilized SARS-CoV-2 spike glycoprotein assessed as dissociation constant by SPR analysisMore data for this Ligand-Target Pair
Affinity DataKd: 1.91E+4nMAssay Description:Binding affinity to CM5 sensor chip immobilized SARS-CoV-2 spike glycoprotein assessed as dissociation constant by SPR analysisMore data for this Ligand-Target Pair
Affinity DataKd: >2.50E+4nMAssay Description:Binding affinity to CM5 sensor chip immobilized SARS-CoV-2 spike glycoprotein assessed as dissociation constant by SPR analysisMore data for this Ligand-Target Pair
Affinity DataKd: >2.50E+4nMAssay Description:Binding affinity to CM5 sensor chip immobilized SARS-CoV-2 spike glycoprotein assessed as dissociation constant by SPR analysisMore data for this Ligand-Target Pair
Affinity DataKd: >2.50E+4nMAssay Description:Binding affinity to CM5 sensor chip immobilized SARS-CoV-2 spike glycoprotein assessed as dissociation constant by SPR analysisMore data for this Ligand-Target Pair
Affinity DataKd: >2.50E+4nMAssay Description:Binding affinity to CM5 sensor chip immobilized SARS-CoV-2 spike glycoprotein assessed as dissociation constant by SPR analysisMore data for this Ligand-Target Pair
Affinity DataKd: >2.50E+4nMAssay Description:Binding affinity to CM5 sensor chip immobilized SARS-CoV-2 spike glycoprotein assessed as dissociation constant by SPR analysisMore data for this Ligand-Target Pair