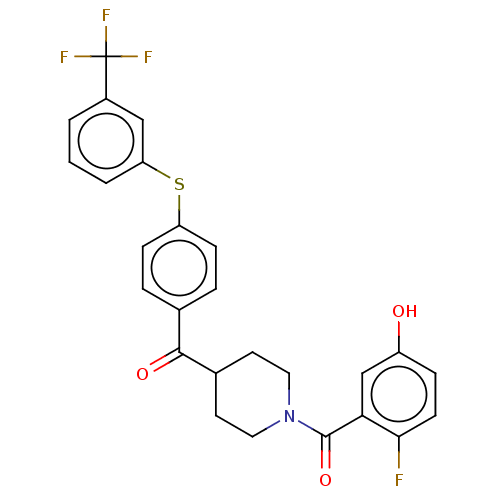
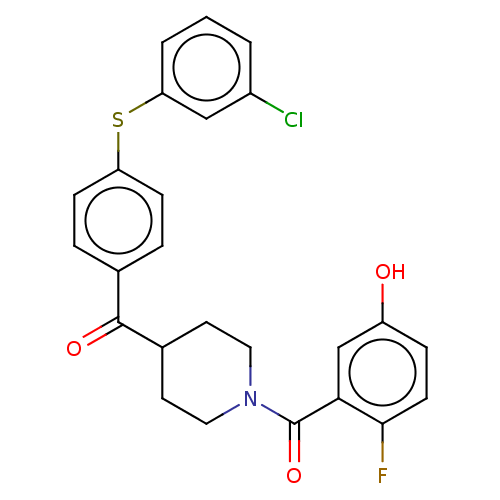
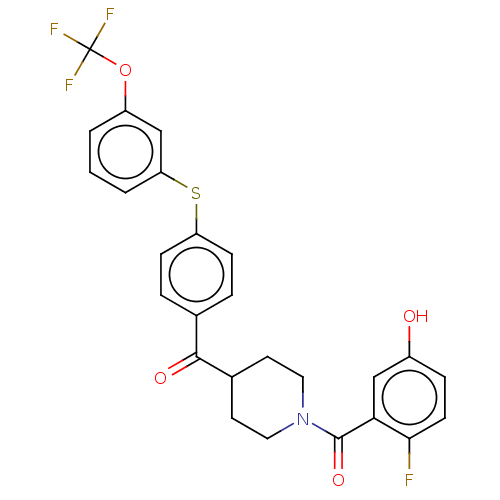
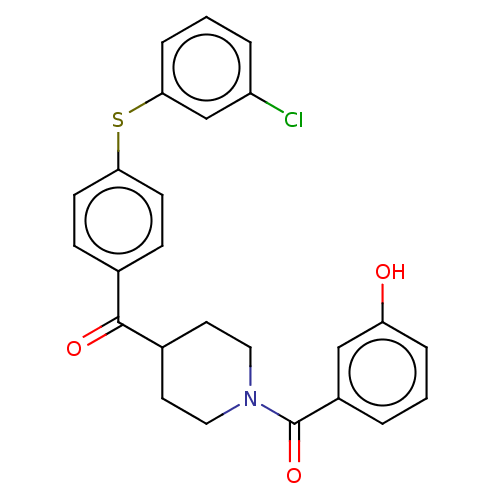
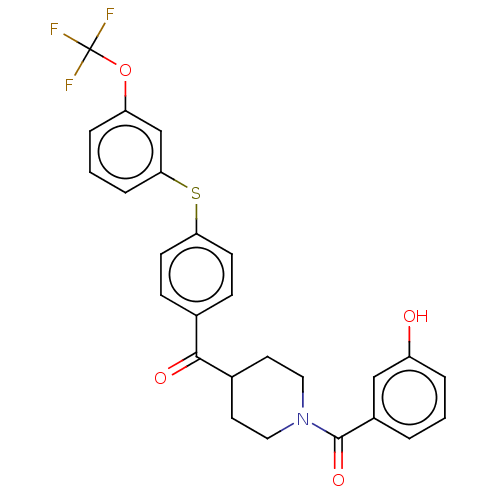
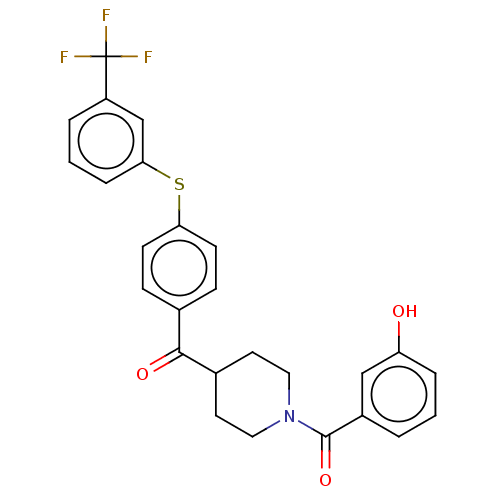
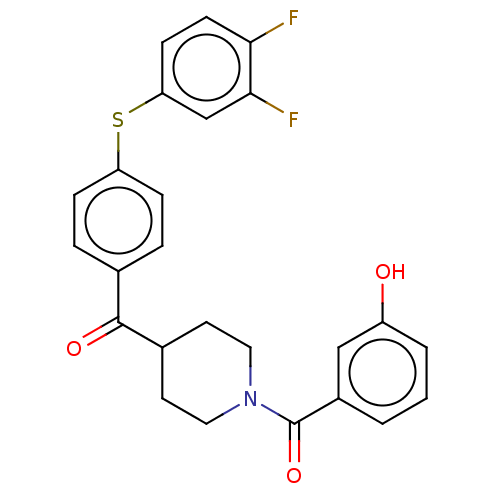
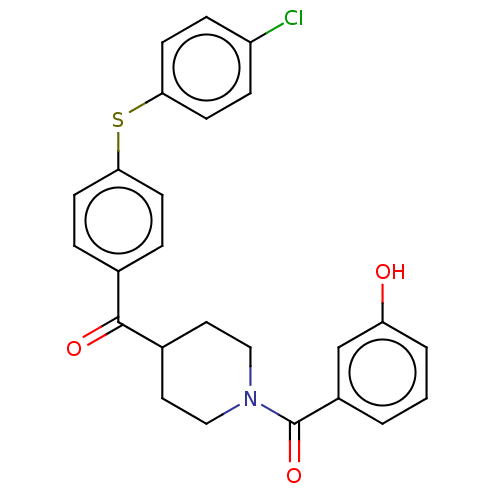
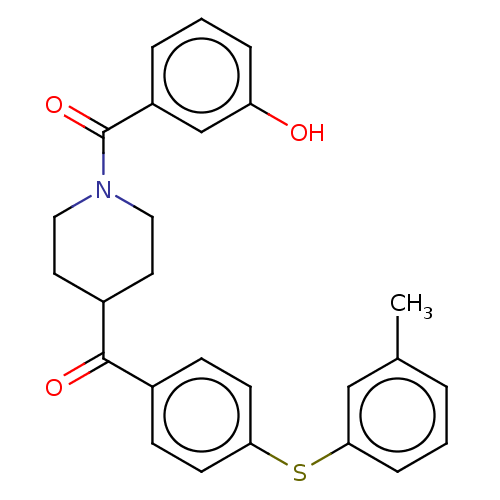
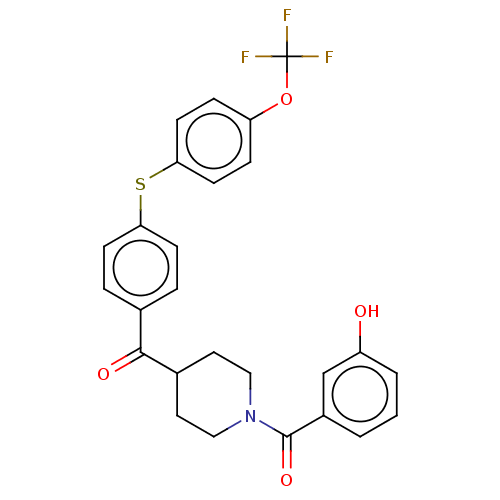
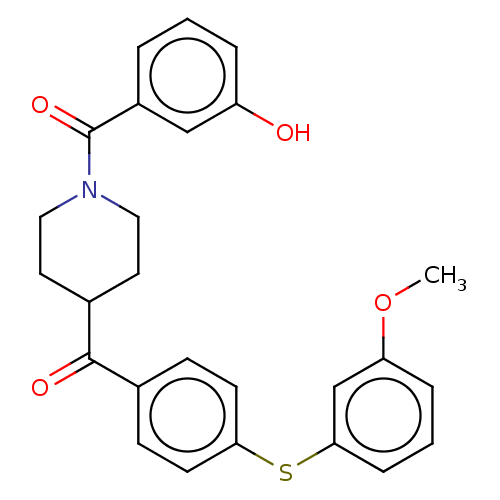
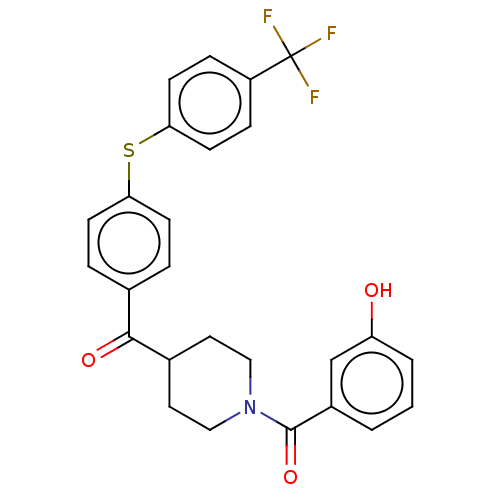
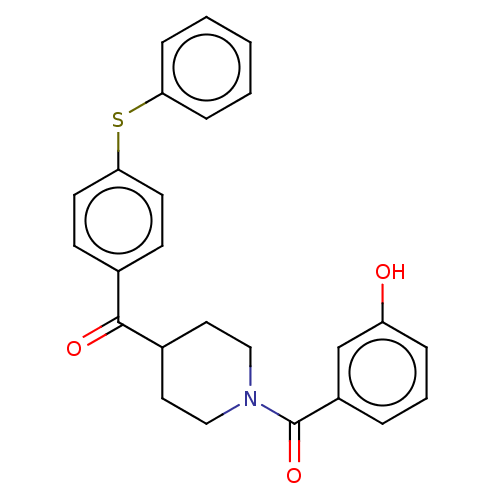
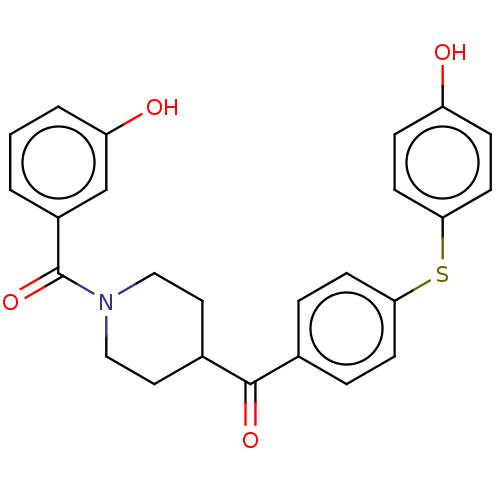
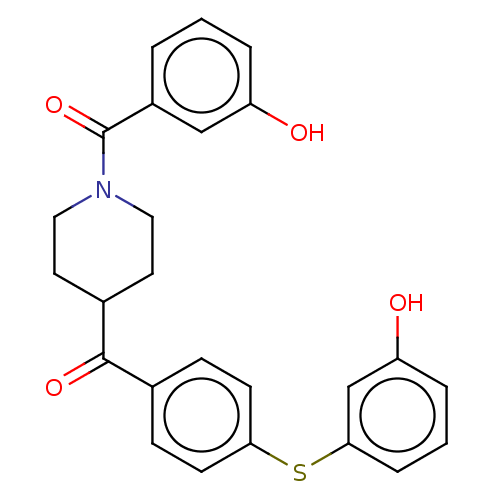
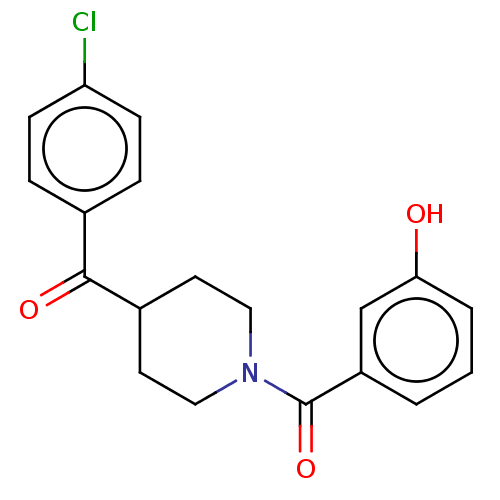
Report error Found 26 Enz. Inhib. hit(s) with all data for entry = 50014298
Affinity DataKi: 1.10nMAssay Description:Inhibition of human recombinant MAGL assessed as dissociation constant using 4-NPA as substrateMore data for this Ligand-Target Pair
Affinity DataKi: 1.10nMAssay Description:Inhibition of human recombinant MAGL assessed as dissociation constant using 4-NPA as substrateMore data for this Ligand-Target Pair
Affinity DataKi: 1.10nMAssay Description:Inhibition of human recombinant MAGL assessed as dissociation constant using 4-NPA as substrateMore data for this Ligand-Target Pair
Affinity DataIC50: 1.30nMAssay Description:Inhibition of human recombinant MAGL using 4-NPA as substrate by microplate reader methodMore data for this Ligand-Target Pair
Affinity DataIC50: 1.5nMAssay Description:Inhibition of human recombinant MAGL using 4-NPA as substrate by microplate reader methodMore data for this Ligand-Target Pair
Affinity DataIC50: 1.90nMAssay Description:Inhibition of human recombinant MAGL using 4-NPA as substrate by microplate reader methodMore data for this Ligand-Target Pair
Affinity DataIC50: 5.70nMAssay Description:Inhibition of human recombinant MAGL using 4-NPA as substrate by microplate reader methodMore data for this Ligand-Target Pair
Affinity DataIC50: 6.20nMAssay Description:Inhibition of human recombinant MAGL using 4-NPA as substrate by microplate reader methodMore data for this Ligand-Target Pair
Affinity DataIC50: 6.90nMAssay Description:Inhibition of human recombinant MAGL using 4-NPA as substrate by microplate reader methodMore data for this Ligand-Target Pair
Affinity DataIC50: 14nMAssay Description:Inhibition of human recombinant MAGL using 4-NPA as substrate by microplate reader methodMore data for this Ligand-Target Pair
Affinity DataIC50: 17nMAssay Description:Inhibition of human recombinant MAGL using 4-NPA as substrate by microplate reader methodMore data for this Ligand-Target Pair
Affinity DataIC50: 18nMAssay Description:Inhibition of human recombinant MAGL using 4-NPA as substrate by microplate reader methodMore data for this Ligand-Target Pair
Affinity DataIC50: 21nMAssay Description:Inhibition of human recombinant MAGL using 4-NPA as substrate by microplate reader methodMore data for this Ligand-Target Pair
Affinity DataIC50: 23nMAssay Description:Inhibition of human recombinant MAGL using 4-NPA as substrate by microplate reader methodMore data for this Ligand-Target Pair
Affinity DataIC50: 25nMAssay Description:Inhibition of human recombinant MAGL using 4-NPA as substrate by microplate reader methodMore data for this Ligand-Target Pair
Affinity DataIC50: 26nMAssay Description:Inhibition of human recombinant MAGL using 4-NPA as substrate by microplate reader methodMore data for this Ligand-Target Pair
Affinity DataIC50: 32nMAssay Description:Inhibition of human recombinant MAGL using 4-NPA as substrate by microplate reader methodMore data for this Ligand-Target Pair
Affinity DataIC50: 50nMAssay Description:Inhibition of human recombinant MAGL using 4-NPA as substrate by microplate reader methodMore data for this Ligand-Target Pair
Affinity DataIC50: 51nMAssay Description:Inhibition of human recombinant MAGL using 4-NPA as substrate by microplate reader methodMore data for this Ligand-Target Pair
Affinity DataIC50: 74nMAssay Description:Inhibition of human recombinant MAGL using 4-NPA as substrate by microplate reader methodMore data for this Ligand-Target Pair
Affinity DataIC50: 96nMAssay Description:Inhibition of human recombinant MAGL using 4-NPA as substrate by microplate reader methodMore data for this Ligand-Target Pair
Affinity DataIC50: 100nMAssay Description:Inhibition of human recombinant MAGL using 4-NPA as substrate by microplate reader methodMore data for this Ligand-Target Pair
Affinity DataIC50: 845nMAssay Description:Inhibition of human recombinant MAGL using 4-NPA as substrate by microplate reader methodMore data for this Ligand-Target Pair
Affinity DataIC50: 2.00E+4nMAssay Description:Inhibition of human recombinant FAAH using AMC arachidonoyl amyl as substrate by fluorescence based assayMore data for this Ligand-Target Pair
Affinity DataIC50: 2.00E+4nMAssay Description:Inhibition of human recombinant FAAH using AMC arachidonoyl amyl as substrate by fluorescence based assayMore data for this Ligand-Target Pair
Affinity DataIC50: 2.00E+4nMAssay Description:Inhibition of human recombinant FAAH using AMC arachidonoyl amyl as substrate by fluorescence based assayMore data for this Ligand-Target Pair