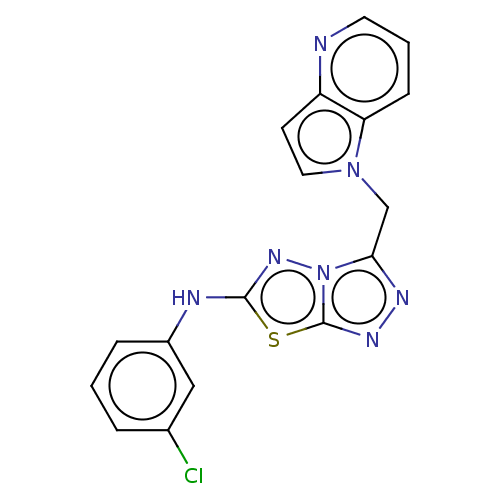
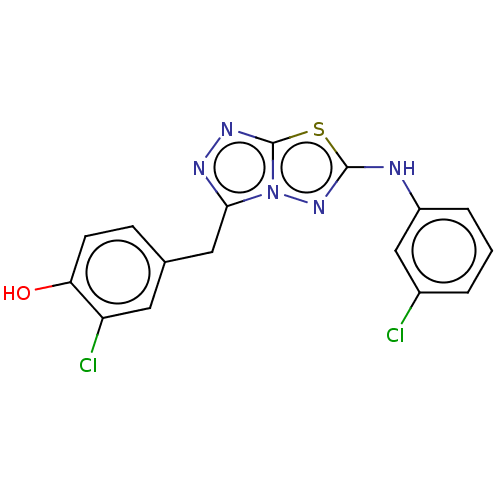
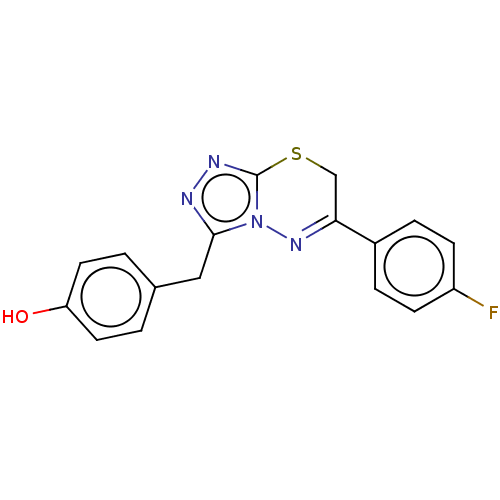
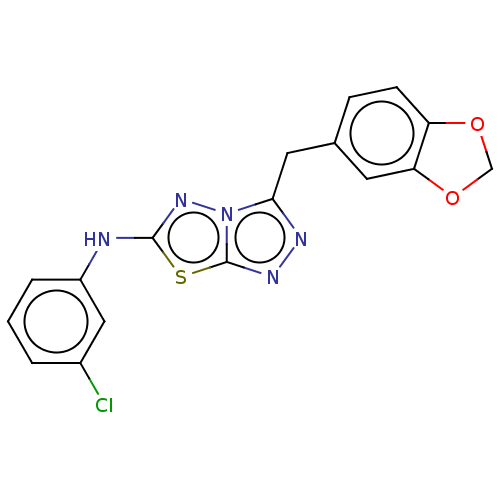
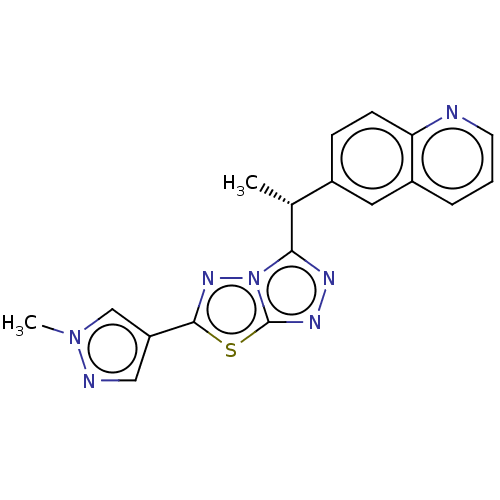
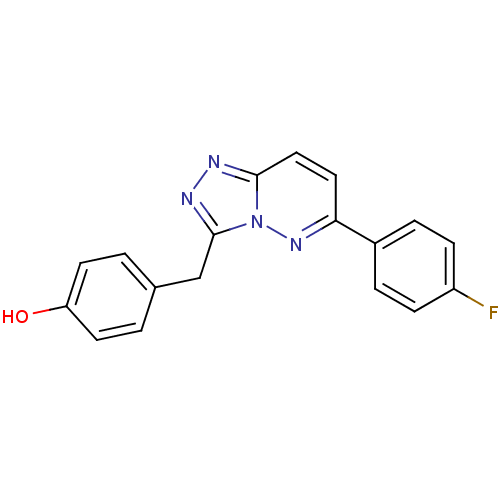
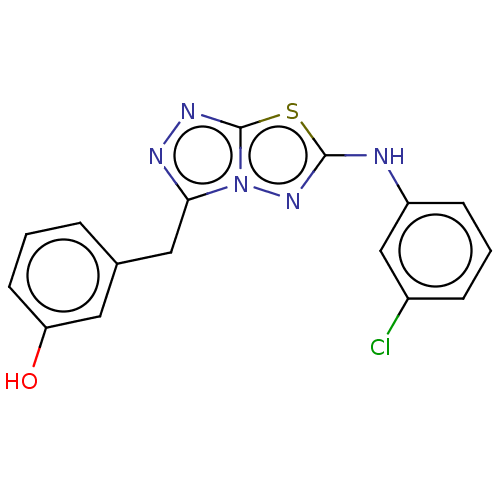
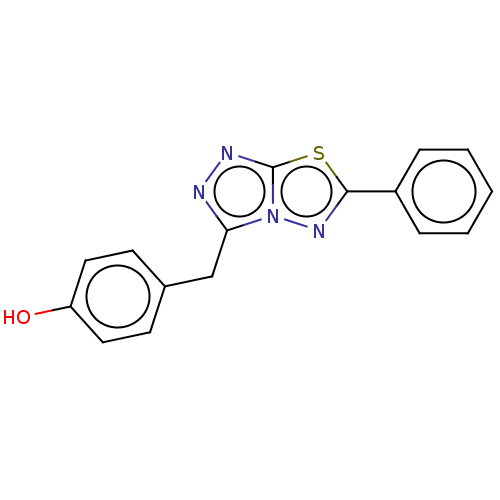
Report error Found 49 Enz. Inhib. hit(s) with all data for entry = 50012482
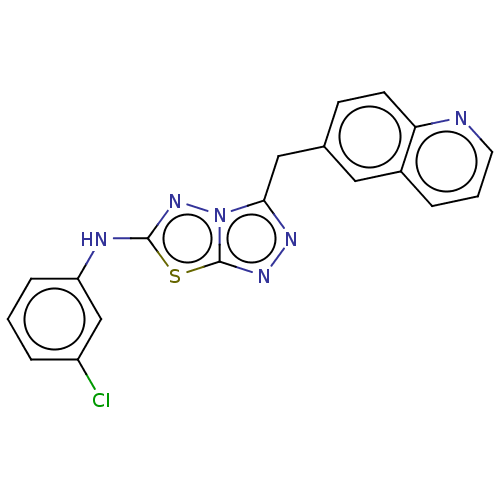
Affinity DataIC50: 3nMAssay Description:Inhibition of c-MET in human SNU-5 cells measured after 6 hrs by steady-glo luciferase reporter gene assayMore data for this Ligand-Target Pair
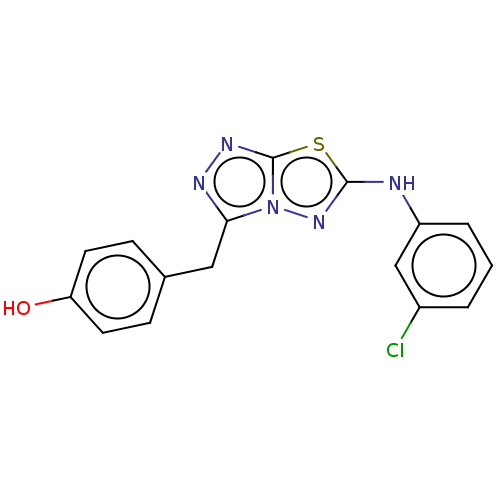
Affinity DataIC50: 3nMAssay Description:Inhibition of c-MET in human SNU-5 cells measured after 6 hrs by steady-glo luciferase reporter gene assayMore data for this Ligand-Target Pair
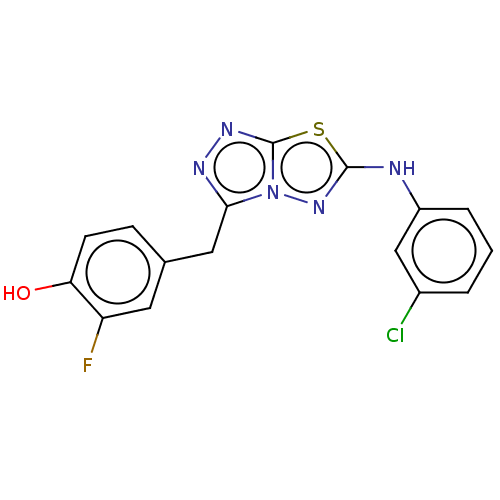
Affinity DataIC50: 4nMAssay Description:Inhibition of c-MET in human SNU-5 cells measured after 6 hrs by steady-glo luciferase reporter gene assayMore data for this Ligand-Target Pair
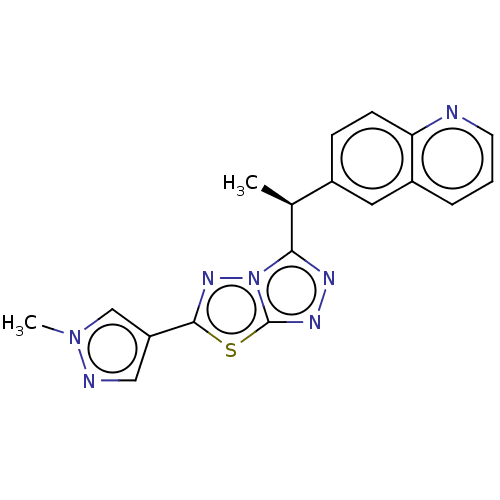
Affinity DataIC50: 5nMAssay Description:Inhibition of c-MET in human SNU-5 cells measured after 6 hrs by steady-glo luciferase reporter gene assayMore data for this Ligand-Target Pair
Affinity DataKi: 7nMAssay Description:Inhibition of c-Met (unknown origin) using poly Glu-Tyr as substrate preincubated for 10 mins followed by ATP addition by phosphoenolpyruvate/pyruvat...More data for this Ligand-Target Pair
Affinity DataKi: 8nMAssay Description:Inhibition of c-Met (unknown origin) using poly Glu-Tyr as substrate preincubated for 10 mins followed by ATP addition by phosphoenolpyruvate/pyruvat...More data for this Ligand-Target Pair
Affinity DataKi: 10nMAssay Description:Inhibition of c-Met (unknown origin) using poly Glu-Tyr as substrate preincubated for 10 mins followed by ATP addition by phosphoenolpyruvate/pyruvat...More data for this Ligand-Target Pair
Affinity DataKi: 15nMAssay Description:Inhibition of c-Met (unknown origin) using poly Glu-Tyr as substrate preincubated for 10 mins followed by ATP addition by phosphoenolpyruvate/pyruvat...More data for this Ligand-Target Pair
Affinity DataKi: 20nMAssay Description:Inhibition of c-Met (unknown origin) using poly Glu-Tyr as substrate preincubated for 10 mins followed by ATP addition by phosphoenolpyruvate/pyruvat...More data for this Ligand-Target Pair
Affinity DataIC50: 20nMAssay Description:Inhibition of c-MET in human SNU-5 cells measured after 6 hrs by steady-glo luciferase reporter gene assayMore data for this Ligand-Target Pair
Affinity DataKi: 20nMAssay Description:Inhibition of c-Met (unknown origin) using poly Glu-Tyr as substrate preincubated for 10 mins followed by ATP addition by phosphoenolpyruvate/pyruvat...More data for this Ligand-Target Pair
Affinity DataKi: 21nMAssay Description:Inhibition of c-Met (unknown origin) using poly Glu-Tyr as substrate preincubated for 10 mins followed by ATP addition by phosphoenolpyruvate/pyruvat...More data for this Ligand-Target Pair
Affinity DataKi: 24nMAssay Description:Inhibition of c-Met (unknown origin) using poly Glu-Tyr as substrate preincubated for 10 mins followed by ATP addition by phosphoenolpyruvate/pyruvat...More data for this Ligand-Target Pair
Affinity DataKi: 30nMAssay Description:Inhibition of c-Met (unknown origin) using poly Glu-Tyr as substrate preincubated for 10 mins followed by ATP addition by phosphoenolpyruvate/pyruvat...More data for this Ligand-Target Pair
Affinity DataIC50: 30nMAssay Description:Inhibition of c-MET in human SNU-5 cells measured after 6 hrs by steady-glo luciferase reporter gene assayMore data for this Ligand-Target Pair
Affinity DataIC50: 30nMAssay Description:Inhibition of c-MET in human SNU-5 cells measured after 6 hrs by steady-glo luciferase reporter gene assayMore data for this Ligand-Target Pair
Affinity DataIC50: 32nMAssay Description:Inhibition of c-MET in human SNU-5 cells measured after 6 hrs by steady-glo luciferase reporter gene assayMore data for this Ligand-Target Pair
Affinity DataKi: 40nMAssay Description:Inhibition of c-Met (unknown origin) using poly Glu-Tyr as substrate preincubated for 10 mins followed by ATP addition by phosphoenolpyruvate/pyruvat...More data for this Ligand-Target Pair
Affinity DataKi: 40nMAssay Description:Inhibition of c-Met (unknown origin) using poly Glu-Tyr as substrate preincubated for 10 mins followed by ATP addition by phosphoenolpyruvate/pyruvat...More data for this Ligand-Target Pair
Affinity DataIC50: 40nMAssay Description:Inhibition of c-MET in human SNU-5 cells measured after 6 hrs by steady-glo luciferase reporter gene assayMore data for this Ligand-Target Pair
Affinity DataKi: 50nMAssay Description:Inhibition of c-Met (unknown origin) using poly Glu-Tyr as substrate preincubated for 10 mins followed by ATP addition by phosphoenolpyruvate/pyruvat...More data for this Ligand-Target Pair
Affinity DataKi: 50nMAssay Description:Inhibition of c-Met (unknown origin) using poly Glu-Tyr as substrate preincubated for 10 mins followed by ATP addition by phosphoenolpyruvate/pyruvat...More data for this Ligand-Target Pair
Affinity DataKi: 60nMAssay Description:Inhibition of c-Met (unknown origin) using poly Glu-Tyr as substrate preincubated for 10 mins followed by ATP addition by phosphoenolpyruvate/pyruvat...More data for this Ligand-Target Pair
Affinity DataKi: 80nMAssay Description:Inhibition of c-Met (unknown origin) using poly Glu-Tyr as substrate preincubated for 10 mins followed by ATP addition by phosphoenolpyruvate/pyruvat...More data for this Ligand-Target Pair
Affinity DataIC50: 80nMAssay Description:Inhibition of c-MET in human SNU-5 cells measured after 6 hrs by steady-glo luciferase reporter gene assayMore data for this Ligand-Target Pair
Affinity DataKi: 80nMAssay Description:Inhibition of c-Met (unknown origin) using poly Glu-Tyr as substrate preincubated for 10 mins followed by ATP addition by phosphoenolpyruvate/pyruvat...More data for this Ligand-Target Pair
Affinity DataIC50: 84nMAssay Description:Inhibition of c-MET in human SNU-5 cells measured after 6 hrs by steady-glo luciferase reporter gene assayMore data for this Ligand-Target Pair
TargetcGMP-inhibited 3',5'-cyclic phosphodiesterase 3A(Human)
Vertex Pharmaceuticals
Curated by ChEMBL
Vertex Pharmaceuticals
Curated by ChEMBL
Affinity DataIC50: 100nMAssay Description:Inhibition of recombinant human N-terminal GST/His-tagged PDE3A (669 to 1141 residues) expressed in baculovirus infected Sf9 insect cells using iFL-c...More data for this Ligand-Target Pair
Affinity DataKi: 200nMAssay Description:Inhibition of c-Met (unknown origin) using poly Glu-Tyr as substrate preincubated for 10 mins followed by ATP addition by phosphoenolpyruvate/pyruvat...More data for this Ligand-Target Pair
TargetcGMP-inhibited 3',5'-cyclic phosphodiesterase 3A(Human)
Vertex Pharmaceuticals
Curated by ChEMBL
Vertex Pharmaceuticals
Curated by ChEMBL
Affinity DataIC50: 200nMAssay Description:Inhibition of recombinant human N-terminal GST/His-tagged PDE3A (669 to 1141 residues) expressed in baculovirus infected Sf9 insect cells using iFL-c...More data for this Ligand-Target Pair
Affinity DataIC50: 200nMAssay Description:Inhibition of c-MET in human SNU-5 cells measured after 6 hrs by steady-glo luciferase reporter gene assayMore data for this Ligand-Target Pair
Affinity DataKi: 200nMAssay Description:Inhibition of c-Met (unknown origin) using poly Glu-Tyr as substrate preincubated for 10 mins followed by ATP addition by phosphoenolpyruvate/pyruvat...More data for this Ligand-Target Pair
Affinity DataKi: 240nMAssay Description:Inhibition of c-Met (unknown origin) using poly Glu-Tyr as substrate preincubated for 10 mins followed by ATP addition by phosphoenolpyruvate/pyruvat...More data for this Ligand-Target Pair
Affinity DataIC50: 300nMAssay Description:Inhibition of c-MET in human SNU-5 cells measured after 6 hrs by steady-glo luciferase reporter gene assayMore data for this Ligand-Target Pair
Affinity DataIC50: 300nMAssay Description:Inhibition of c-MET in human SNU-5 cells measured after 6 hrs by steady-glo luciferase reporter gene assayMore data for this Ligand-Target Pair
Affinity DataIC50: 380nMAssay Description:Inhibition of c-MET in human SNU-5 cells measured after 6 hrs by steady-glo luciferase reporter gene assayMore data for this Ligand-Target Pair
Affinity DataKi: 400nMAssay Description:Inhibition of c-Met (unknown origin) using poly Glu-Tyr as substrate preincubated for 10 mins followed by ATP addition by phosphoenolpyruvate/pyruvat...More data for this Ligand-Target Pair
Affinity DataKi: 400nMAssay Description:Inhibition of c-Met (unknown origin) using poly Glu-Tyr as substrate preincubated for 10 mins followed by ATP addition by phosphoenolpyruvate/pyruvat...More data for this Ligand-Target Pair
TargetcGMP-inhibited 3',5'-cyclic phosphodiesterase 3A(Human)
Vertex Pharmaceuticals
Curated by ChEMBL
Vertex Pharmaceuticals
Curated by ChEMBL
Affinity DataIC50: 500nMAssay Description:Inhibition of recombinant human N-terminal GST/His-tagged PDE3A (669 to 1141 residues) expressed in baculovirus infected Sf9 insect cells using iFL-c...More data for this Ligand-Target Pair
Affinity DataIC50: 800nMAssay Description:Inhibition of c-MET in human SNU-5 cells measured after 6 hrs by steady-glo luciferase reporter gene assayMore data for this Ligand-Target Pair
Affinity DataIC50: 1.40E+3nMAssay Description:Inhibition of c-MET in human SNU-5 cells measured after 6 hrs by steady-glo luciferase reporter gene assayMore data for this Ligand-Target Pair
TargetcGMP-inhibited 3',5'-cyclic phosphodiesterase 3A(Human)
Vertex Pharmaceuticals
Curated by ChEMBL
Vertex Pharmaceuticals
Curated by ChEMBL
Affinity DataIC50: 3.00E+3nMAssay Description:Inhibition of recombinant human N-terminal GST/His-tagged PDE3A (669 to 1141 residues) expressed in baculovirus infected Sf9 insect cells using iFL-c...More data for this Ligand-Target Pair
Affinity DataKi: >4.00E+3nMAssay Description:Inhibition of SKY (unknown origin)More data for this Ligand-Target Pair
Affinity DataKi: >4.00E+3nMAssay Description:Inhibition of SRC (unknown origin)More data for this Ligand-Target Pair
Affinity DataKi: >4.00E+3nMAssay Description:Inhibition of KDR (unknown origin)More data for this Ligand-Target Pair
Affinity DataKi: >4.00E+3nMAssay Description:Inhibition of JAK2 (unknown origin)More data for this Ligand-Target Pair
Affinity DataKi: >4.00E+3nMAssay Description:Inhibition of FLT3 (unknown origin)More data for this Ligand-Target Pair
TargetcGMP-inhibited 3',5'-cyclic phosphodiesterase 3A(Human)
Vertex Pharmaceuticals
Curated by ChEMBL
Vertex Pharmaceuticals
Curated by ChEMBL
Affinity DataIC50: 7.90E+3nMAssay Description:Inhibition of recombinant human N-terminal GST/His-tagged PDE3A (669 to 1141 residues) expressed in baculovirus infected Sf9 insect cells using iFL-c...More data for this Ligand-Target Pair
TargetcGMP-inhibited 3',5'-cyclic phosphodiesterase 3A(Human)
Vertex Pharmaceuticals
Curated by ChEMBL
Vertex Pharmaceuticals
Curated by ChEMBL
Affinity DataIC50: 8.07E+3nMAssay Description:Inhibition of recombinant human N-terminal GST/His-tagged PDE3A (669 to 1141 residues) expressed in baculovirus infected Sf9 insect cells using iFL-c...More data for this Ligand-Target Pair