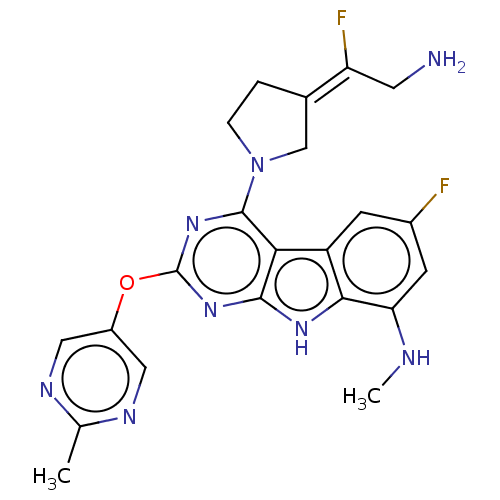
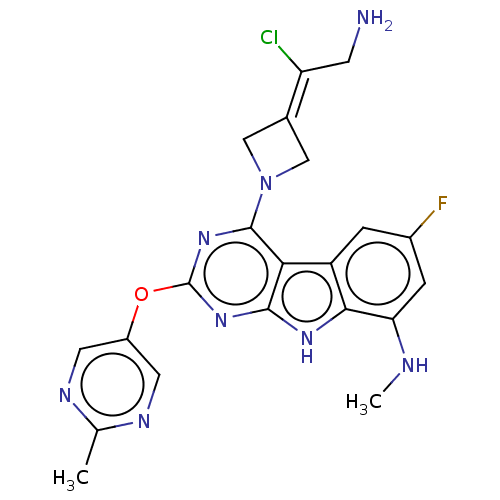
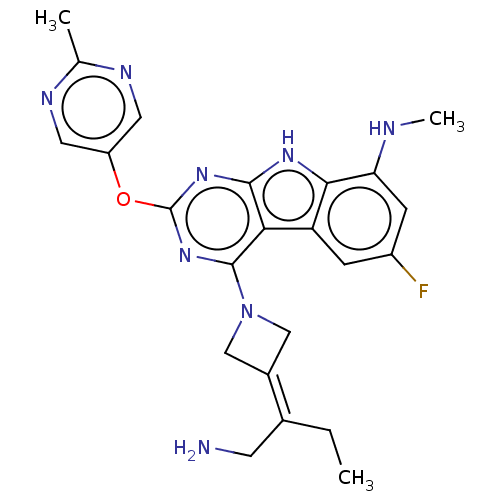
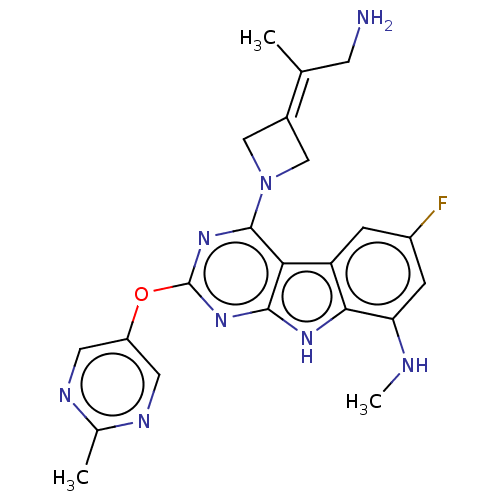
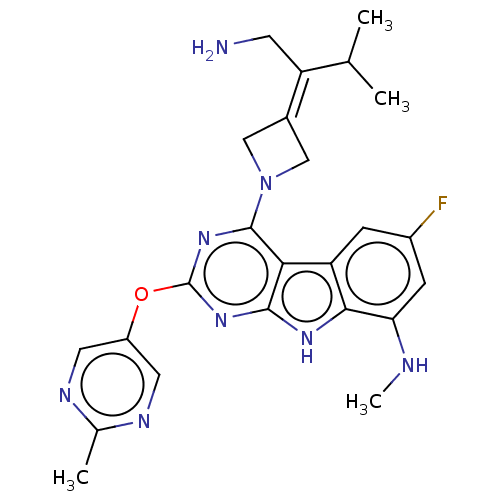
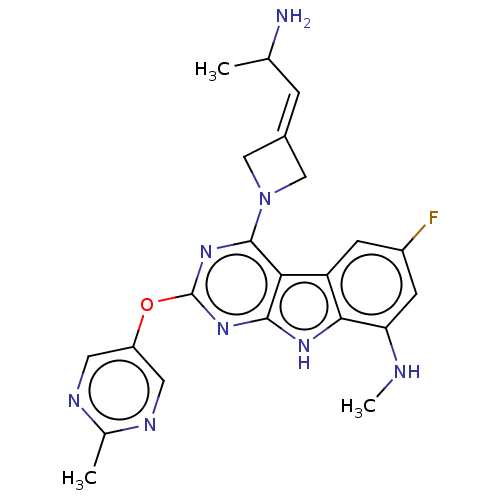
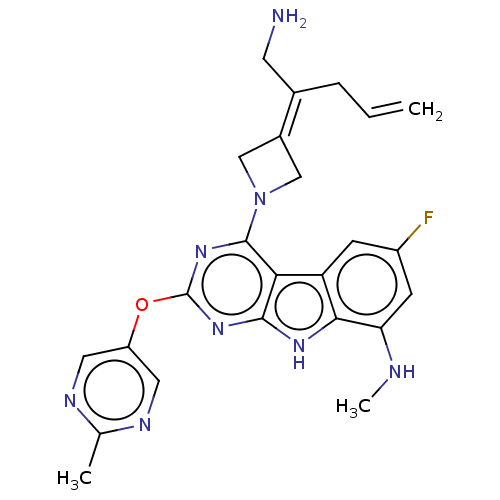
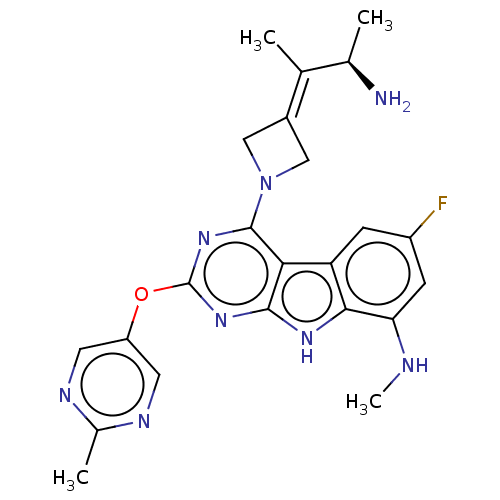
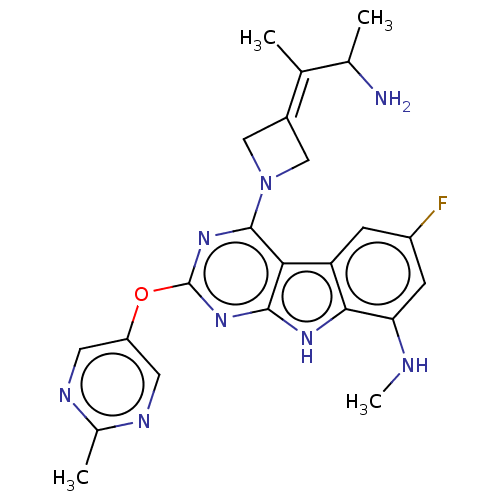
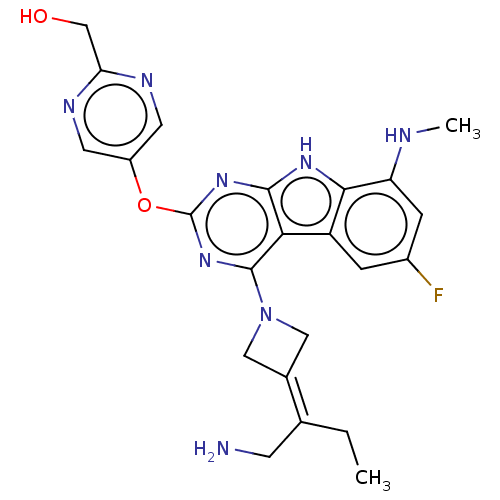
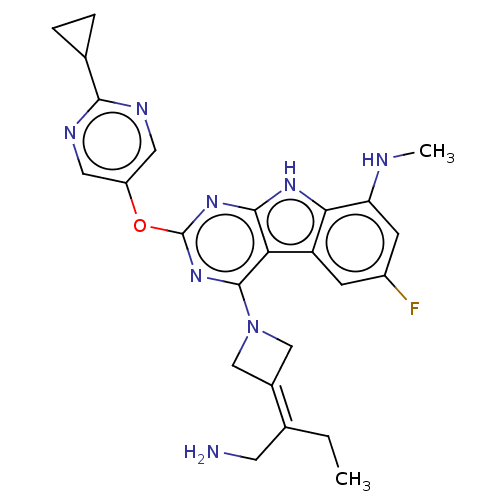
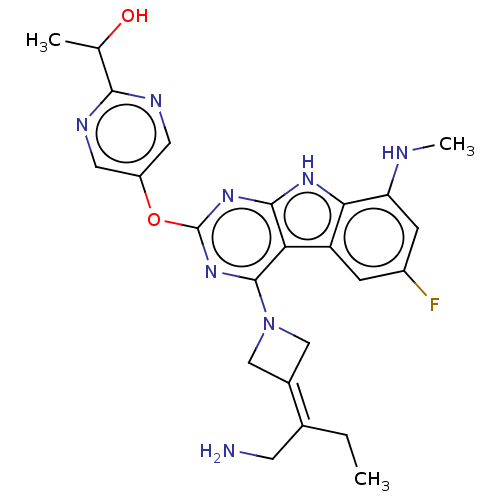
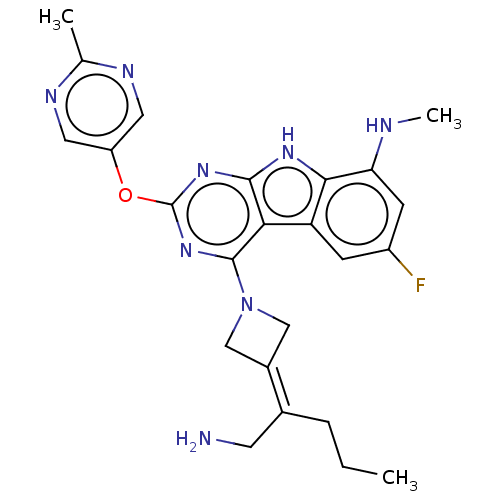
Report error Found 18 Enz. Inhib. hit(s) with all data for entry = 50015635
TargetPotassium voltage-gated channel subfamily H member 2(Human)
Shanghai Institute of Materia Medica
Curated by ChEMBL
Shanghai Institute of Materia Medica
Curated by ChEMBL
Affinity DataIC50: 40nMAssay Description:Inhibition of hERG potassium channel assessed as maximum inhibition rate at -80 mV holding potential incubated for 120 seconds by whole cell-automate...More data for this Ligand-Target Pair
TargetPotassium voltage-gated channel subfamily H member 2(Human)
Shanghai Institute of Materia Medica
Curated by ChEMBL
Shanghai Institute of Materia Medica
Curated by ChEMBL
Affinity DataIC50: 1.23E+4nMAssay Description:Inhibition of hERG potassium channel assessed as maximum inhibition rate at -80 mV holding potential incubated for 120 seconds by whole cell-automate...More data for this Ligand-Target Pair
TargetPotassium voltage-gated channel subfamily H member 2(Human)
Shanghai Institute of Materia Medica
Curated by ChEMBL
Shanghai Institute of Materia Medica
Curated by ChEMBL
Affinity DataIC50: 1.84E+4nMAssay Description:Inhibition of hERG potassium channel assessed as maximum inhibition rate at -80 mV holding potential incubated for 120 seconds by whole cell-automate...More data for this Ligand-Target Pair
TargetPotassium voltage-gated channel subfamily H member 2(Human)
Shanghai Institute of Materia Medica
Curated by ChEMBL
Shanghai Institute of Materia Medica
Curated by ChEMBL
Affinity DataIC50: 4.00E+4nMAssay Description:Inhibition of hERG potassium channel assessed as maximum inhibition rate at -80 mV holding potential incubated for 120 seconds by whole cell-automate...More data for this Ligand-Target Pair
TargetPotassium voltage-gated channel subfamily H member 2(Human)
Shanghai Institute of Materia Medica
Curated by ChEMBL
Shanghai Institute of Materia Medica
Curated by ChEMBL
Affinity DataIC50: 4.00E+4nMAssay Description:Inhibition of hERG potassium channel assessed as maximum inhibition rate at -80 mV holding potential incubated for 120 seconds by whole cell-automate...More data for this Ligand-Target Pair
TargetPotassium voltage-gated channel subfamily H member 2(Human)
Shanghai Institute of Materia Medica
Curated by ChEMBL
Shanghai Institute of Materia Medica
Curated by ChEMBL
Affinity DataIC50: 4.00E+4nMAssay Description:Inhibition of hERG potassium channel assessed as maximum inhibition rate at -80 mV holding potential incubated for 120 seconds by whole cell-automate...More data for this Ligand-Target Pair
TargetPotassium voltage-gated channel subfamily H member 2(Human)
Shanghai Institute of Materia Medica
Curated by ChEMBL
Shanghai Institute of Materia Medica
Curated by ChEMBL
Affinity DataIC50: 4.00E+4nMAssay Description:Inhibition of hERG potassium channel assessed as maximum inhibition rate at -80 mV holding potential incubated for 120 seconds by whole cell-automate...More data for this Ligand-Target Pair
TargetPotassium voltage-gated channel subfamily H member 2(Human)
Shanghai Institute of Materia Medica
Curated by ChEMBL
Shanghai Institute of Materia Medica
Curated by ChEMBL
Affinity DataIC50: 4.00E+4nMAssay Description:Inhibition of hERG potassium channel assessed as maximum inhibition rate at -80 mV holding potential incubated for 120 seconds by whole cell-automate...More data for this Ligand-Target Pair
TargetPotassium voltage-gated channel subfamily H member 2(Human)
Shanghai Institute of Materia Medica
Curated by ChEMBL
Shanghai Institute of Materia Medica
Curated by ChEMBL
Affinity DataIC50: 4.00E+4nMAssay Description:Inhibition of hERG potassium channel assessed as maximum inhibition rate at -80 mV holding potential incubated for 120 seconds by whole cell-automate...More data for this Ligand-Target Pair
TargetPotassium voltage-gated channel subfamily H member 2(Human)
Shanghai Institute of Materia Medica
Curated by ChEMBL
Shanghai Institute of Materia Medica
Curated by ChEMBL
Affinity DataIC50: 4.00E+4nMAssay Description:Inhibition of hERG potassium channel assessed as maximum inhibition rate at -80 mV holding potential incubated for 120 seconds by whole cell-automate...More data for this Ligand-Target Pair
TargetPotassium voltage-gated channel subfamily H member 2(Human)
Shanghai Institute of Materia Medica
Curated by ChEMBL
Shanghai Institute of Materia Medica
Curated by ChEMBL
Affinity DataIC50: 4.00E+4nMAssay Description:Inhibition of hERG potassium channel assessed as maximum inhibition rate at -80 mV holding potential incubated for 120 seconds by whole cell-automate...More data for this Ligand-Target Pair
TargetPotassium voltage-gated channel subfamily H member 2(Human)
Shanghai Institute of Materia Medica
Curated by ChEMBL
Shanghai Institute of Materia Medica
Curated by ChEMBL
Affinity DataIC50: 4.00E+4nMAssay Description:Inhibition of hERG potassium channel assessed as maximum inhibition rate at -80 mV holding potential incubated for 120 seconds by whole cell-automate...More data for this Ligand-Target Pair
TargetPotassium voltage-gated channel subfamily H member 2(Human)
Shanghai Institute of Materia Medica
Curated by ChEMBL
Shanghai Institute of Materia Medica
Curated by ChEMBL
Affinity DataIC50: 4.00E+4nMAssay Description:Inhibition of hERG potassium channel assessed as maximum inhibition rate at -80 mV holding potential incubated for 120 seconds by whole cell-automate...More data for this Ligand-Target Pair
TargetPotassium voltage-gated channel subfamily H member 2(Human)
Shanghai Institute of Materia Medica
Curated by ChEMBL
Shanghai Institute of Materia Medica
Curated by ChEMBL
Affinity DataIC50: 4.00E+4nMAssay Description:Inhibition of hERG potassium channel assessed as maximum inhibition rate at -80 mV holding potential incubated for 120 seconds by whole cell-automate...More data for this Ligand-Target Pair
TargetPotassium voltage-gated channel subfamily H member 2(Human)
Shanghai Institute of Materia Medica
Curated by ChEMBL
Shanghai Institute of Materia Medica
Curated by ChEMBL
Affinity DataIC50: 4.00E+4nMAssay Description:Inhibition of hERG potassium channel assessed as maximum inhibition rate at -80 mV holding potential incubated for 120 seconds by whole cell-automate...More data for this Ligand-Target Pair
TargetPotassium voltage-gated channel subfamily H member 2(Human)
Shanghai Institute of Materia Medica
Curated by ChEMBL
Shanghai Institute of Materia Medica
Curated by ChEMBL
Affinity DataIC50: 4.00E+4nMAssay Description:Inhibition of hERG potassium channel assessed as maximum inhibition rate at -80 mV holding potential incubated for 120 seconds by whole cell-automate...More data for this Ligand-Target Pair
TargetPotassium voltage-gated channel subfamily H member 2(Human)
Shanghai Institute of Materia Medica
Curated by ChEMBL
Shanghai Institute of Materia Medica
Curated by ChEMBL
Affinity DataIC50: 4.00E+4nMAssay Description:Inhibition of hERG potassium channel assessed as maximum inhibition rate at -80 mV holding potential incubated for 120 seconds by whole cell-automate...More data for this Ligand-Target Pair
TargetPotassium voltage-gated channel subfamily H member 2(Human)
Shanghai Institute of Materia Medica
Curated by ChEMBL
Shanghai Institute of Materia Medica
Curated by ChEMBL
Affinity DataIC50: 5.15E+4nMAssay Description:Inhibition of hERG potassium channel assessed as maximum inhibition rate at -80 mV holding potential incubated for 120 seconds by whole cell-automate...More data for this Ligand-Target Pair