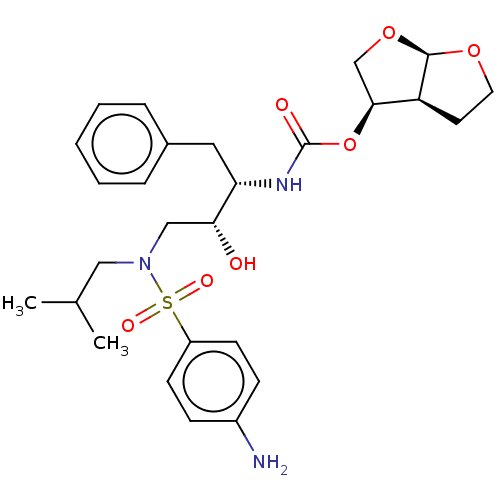
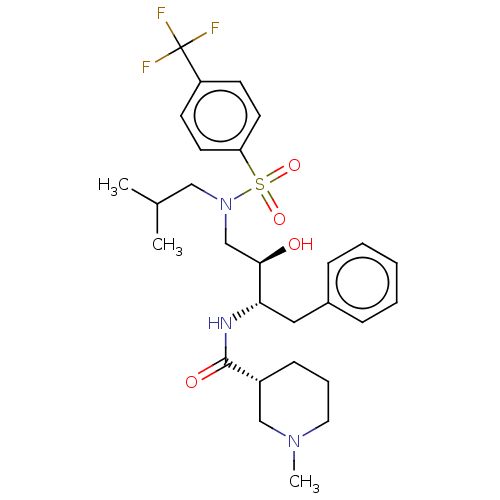
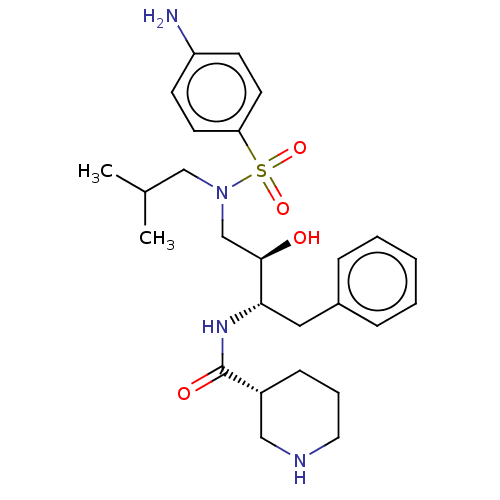
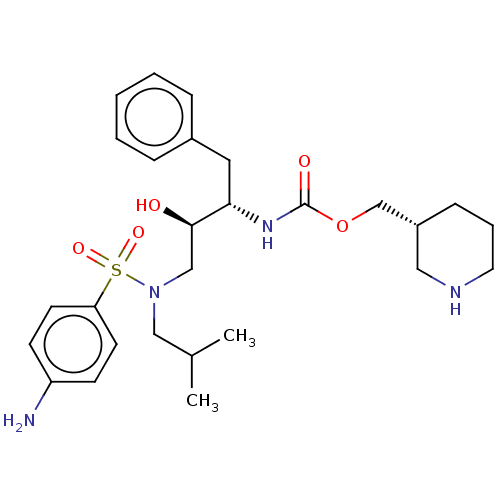
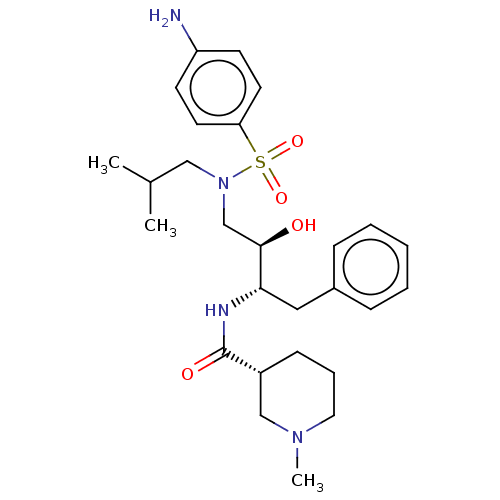
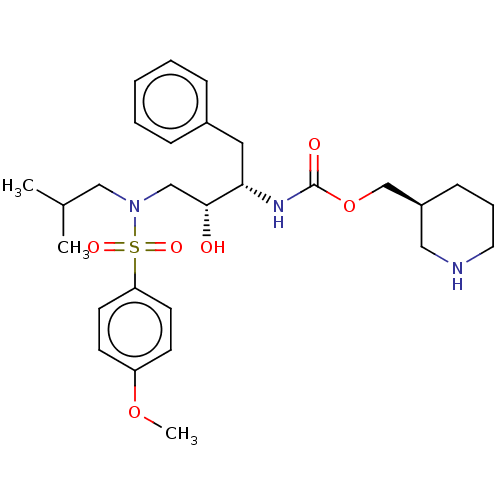
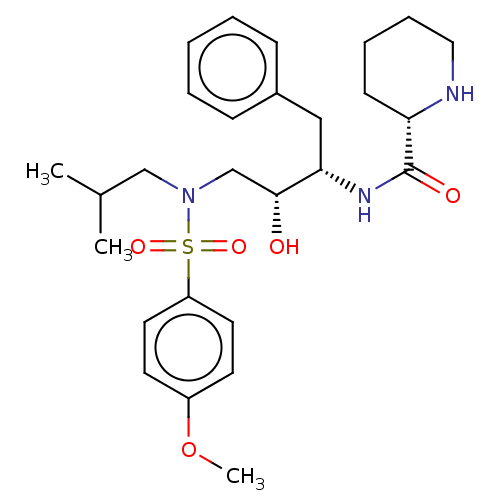
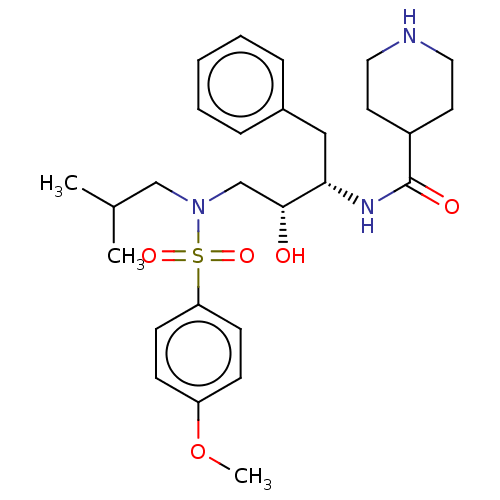
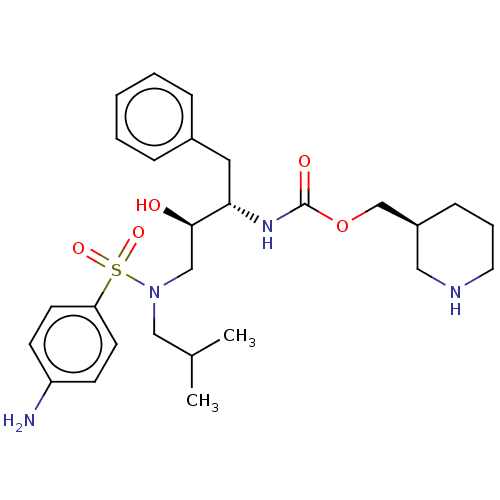
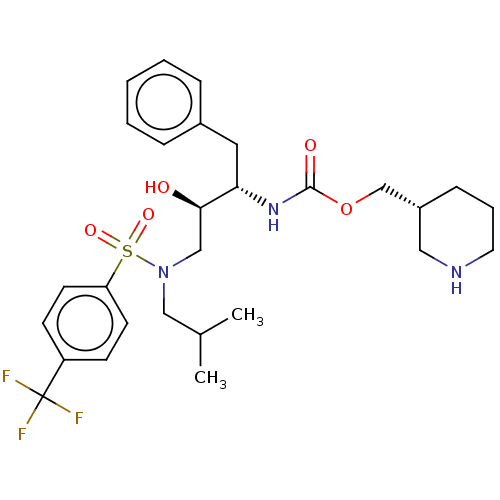
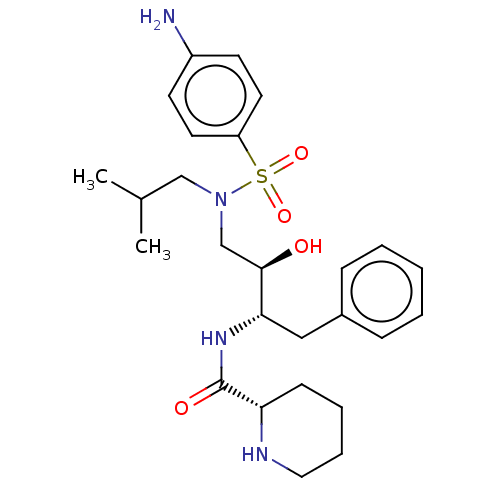
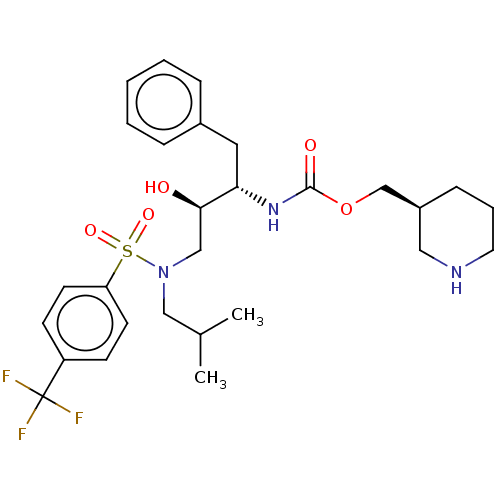
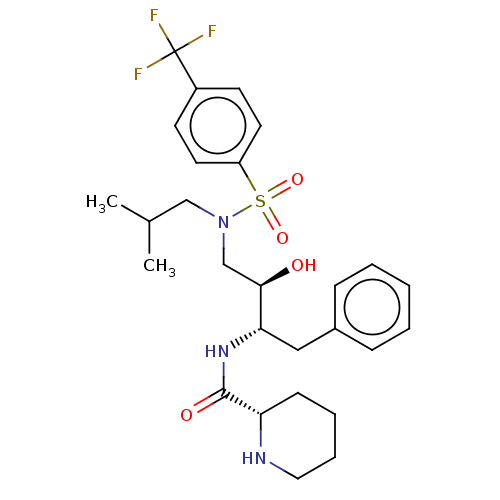
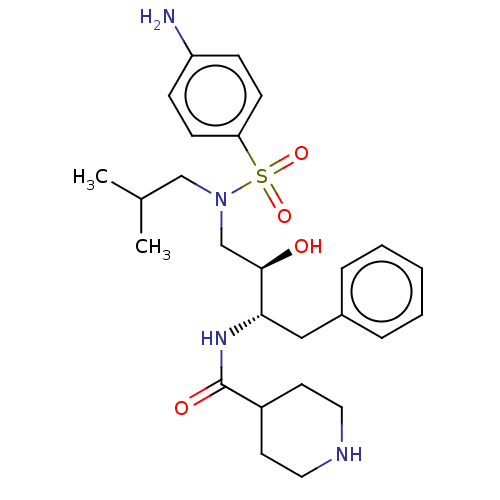
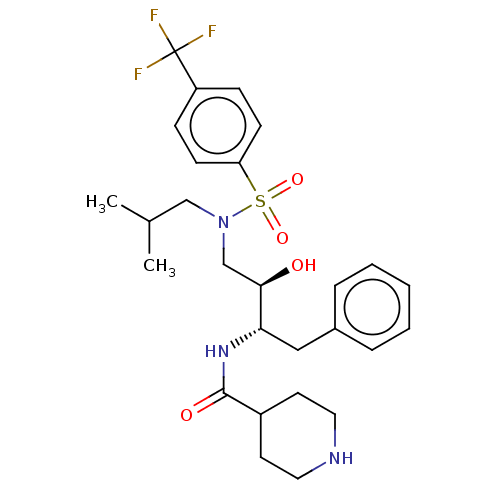
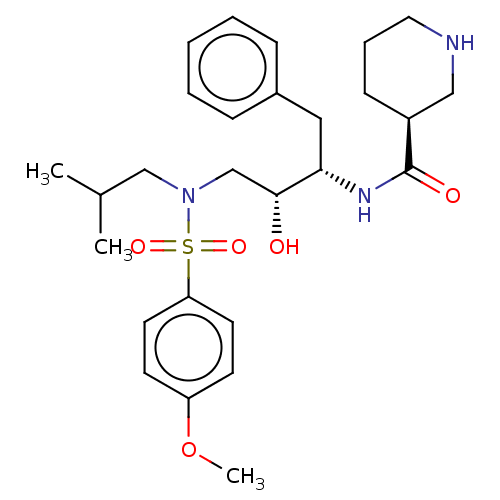
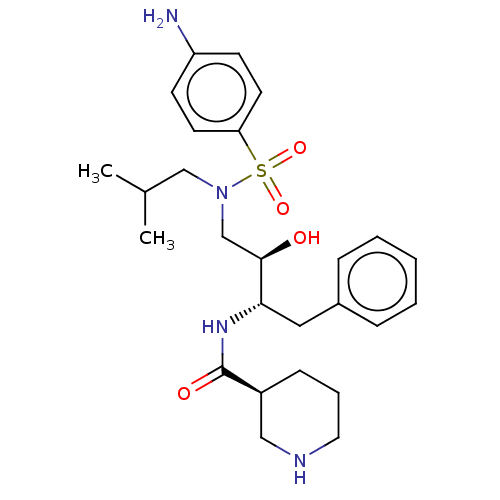
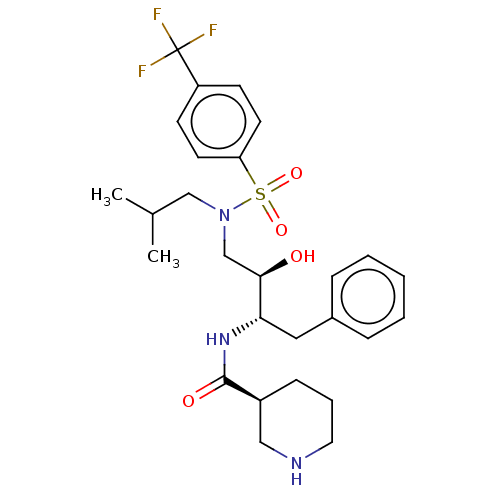
Report error Found 22 Enz. Inhib. hit(s) with all data for entry = 50014504
TargetGag-Pol polyprotein [489-587](Human immunodeficiency virus type 1)
Peking Union Medical College
Curated by ChEMBL
Peking Union Medical College
Curated by ChEMBL
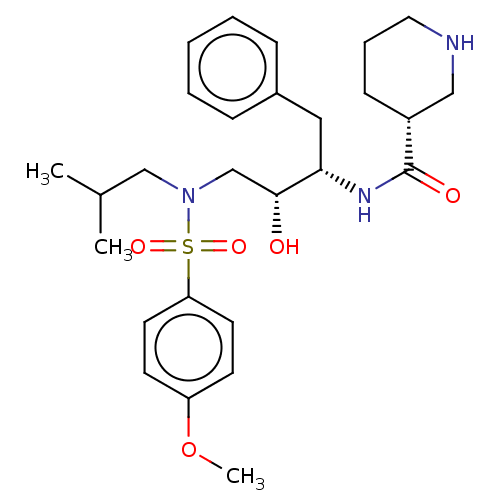
Affinity DataKi: 0.0290nMAssay Description:Inhibition of wild type HIV1 protease using Arg-Glu (EDANS)-Ser-Gln-Asn-Tyr-Pro-Ile-Val-Gln-Lys(DABCYL)-Arg as substrate preincubated for 20 to 30 mi...More data for this Ligand-Target Pair
TargetGag-Pol polyprotein [489-587](Human immunodeficiency virus type 1)
Peking Union Medical College
Curated by ChEMBL
Peking Union Medical College
Curated by ChEMBL
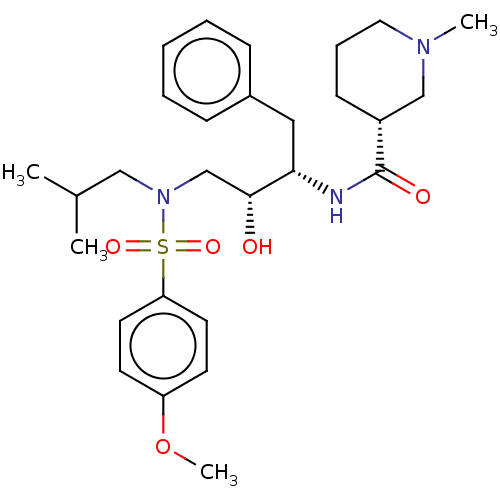
Affinity DataKi: 0.0400nMAssay Description:Inhibition of wild type HIV1 protease using Arg-Glu (EDANS)-Ser-Gln-Asn-Tyr-Pro-Ile-Val-Gln-Lys(DABCYL)-Arg as substrate preincubated for 20 to 30 mi...More data for this Ligand-Target Pair
TargetGag-Pol polyprotein [489-587](Human immunodeficiency virus type 1)
Peking Union Medical College
Curated by ChEMBL
Peking Union Medical College
Curated by ChEMBL
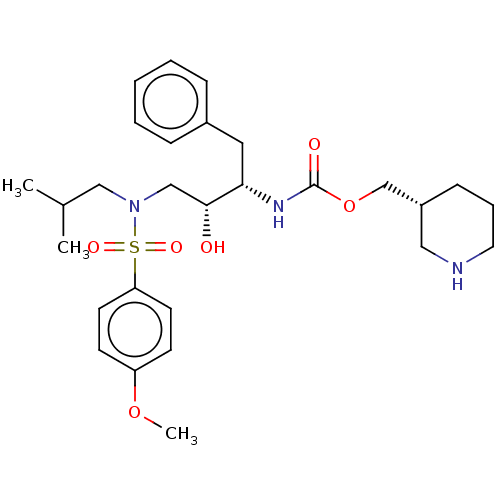
Affinity DataKi: 0.100nMAssay Description:Inhibition of wild type HIV1 protease using Arg-Glu (EDANS)-Ser-Gln-Asn-Tyr-Pro-Ile-Val-Gln-Lys(DABCYL)-Arg as substrate preincubated for 20 to 30 mi...More data for this Ligand-Target Pair
TargetGag-Pol polyprotein [489-587](Human immunodeficiency virus type 1)
Peking Union Medical College
Curated by ChEMBL
Peking Union Medical College
Curated by ChEMBL
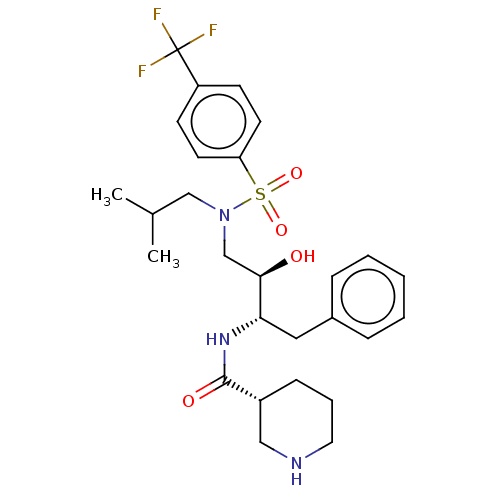
Affinity DataKi: 0.100nMAssay Description:Inhibition of wild type HIV1 protease using Arg-Glu (EDANS)-Ser-Gln-Asn-Tyr-Pro-Ile-Val-Gln-Lys(DABCYL)-Arg as substrate preincubated for 20 to 30 mi...More data for this Ligand-Target Pair
TargetGag-Pol polyprotein [489-587](Human immunodeficiency virus type 1)
Peking Union Medical College
Curated by ChEMBL
Peking Union Medical College
Curated by ChEMBL
Affinity DataKi: 0.180nMAssay Description:Inhibition of wild type HIV1 protease using Arg-Glu (EDANS)-Ser-Gln-Asn-Tyr-Pro-Ile-Val-Gln-Lys(DABCYL)-Arg as substrate preincubated for 20 to 30 mi...More data for this Ligand-Target Pair
TargetGag-Pol polyprotein [489-587](Human immunodeficiency virus type 1)
Peking Union Medical College
Curated by ChEMBL
Peking Union Medical College
Curated by ChEMBL
Affinity DataKi: 0.190nMAssay Description:Inhibition of wild type HIV1 protease using Arg-Glu (EDANS)-Ser-Gln-Asn-Tyr-Pro-Ile-Val-Gln-Lys(DABCYL)-Arg as substrate preincubated for 20 to 30 mi...More data for this Ligand-Target Pair
TargetGag-Pol polyprotein [489-587](Human immunodeficiency virus type 1)
Peking Union Medical College
Curated by ChEMBL
Peking Union Medical College
Curated by ChEMBL
Affinity DataKi: 0.270nMAssay Description:Inhibition of wild type HIV1 protease using Arg-Glu (EDANS)-Ser-Gln-Asn-Tyr-Pro-Ile-Val-Gln-Lys(DABCYL)-Arg as substrate preincubated for 20 to 30 mi...More data for this Ligand-Target Pair
TargetGag-Pol polyprotein [489-587](Human immunodeficiency virus type 1)
Peking Union Medical College
Curated by ChEMBL
Peking Union Medical College
Curated by ChEMBL
Affinity DataKi: 0.280nMAssay Description:Inhibition of wild type HIV1 protease using Arg-Glu (EDANS)-Ser-Gln-Asn-Tyr-Pro-Ile-Val-Gln-Lys(DABCYL)-Arg as substrate preincubated for 20 to 30 mi...More data for this Ligand-Target Pair
TargetGag-Pol polyprotein [489-587](Human immunodeficiency virus type 1)
Peking Union Medical College
Curated by ChEMBL
Peking Union Medical College
Curated by ChEMBL
Affinity DataKi: 0.300nMAssay Description:Inhibition of wild type HIV1 protease using Arg-Glu (EDANS)-Ser-Gln-Asn-Tyr-Pro-Ile-Val-Gln-Lys(DABCYL)-Arg as substrate preincubated for 20 to 30 mi...More data for this Ligand-Target Pair
TargetGag-Pol polyprotein [489-587](Human immunodeficiency virus type 1)
Peking Union Medical College
Curated by ChEMBL
Peking Union Medical College
Curated by ChEMBL
Affinity DataKi: 0.300nMAssay Description:Inhibition of wild type HIV1 protease using Arg-Glu (EDANS)-Ser-Gln-Asn-Tyr-Pro-Ile-Val-Gln-Lys(DABCYL)-Arg as substrate preincubated for 20 to 30 mi...More data for this Ligand-Target Pair
TargetGag-Pol polyprotein [489-587](Human immunodeficiency virus type 1)
Peking Union Medical College
Curated by ChEMBL
Peking Union Medical College
Curated by ChEMBL
Affinity DataKi: 0.330nMAssay Description:Inhibition of wild type HIV1 protease using Arg-Glu (EDANS)-Ser-Gln-Asn-Tyr-Pro-Ile-Val-Gln-Lys(DABCYL)-Arg as substrate preincubated for 20 to 30 mi...More data for this Ligand-Target Pair
TargetGag-Pol polyprotein [489-587](Human immunodeficiency virus type 1)
Peking Union Medical College
Curated by ChEMBL
Peking Union Medical College
Curated by ChEMBL
Affinity DataKi: 0.550nMAssay Description:Inhibition of wild type HIV1 protease using Arg-Glu (EDANS)-Ser-Gln-Asn-Tyr-Pro-Ile-Val-Gln-Lys(DABCYL)-Arg as substrate preincubated for 20 to 30 mi...More data for this Ligand-Target Pair
TargetGag-Pol polyprotein [489-587](Human immunodeficiency virus type 1)
Peking Union Medical College
Curated by ChEMBL
Peking Union Medical College
Curated by ChEMBL
Affinity DataKi: 1nMAssay Description:Inhibition of wild type HIV1 protease using Arg-Glu (EDANS)-Ser-Gln-Asn-Tyr-Pro-Ile-Val-Gln-Lys(DABCYL)-Arg as substrate preincubated for 20 to 30 mi...More data for this Ligand-Target Pair
TargetGag-Pol polyprotein [489-587](Human immunodeficiency virus type 1)
Peking Union Medical College
Curated by ChEMBL
Peking Union Medical College
Curated by ChEMBL
Affinity DataKi: 1.70nMAssay Description:Inhibition of wild type HIV1 protease using Arg-Glu (EDANS)-Ser-Gln-Asn-Tyr-Pro-Ile-Val-Gln-Lys(DABCYL)-Arg as substrate preincubated for 20 to 30 mi...More data for this Ligand-Target Pair
TargetGag-Pol polyprotein [489-587](Human immunodeficiency virus type 1)
Peking Union Medical College
Curated by ChEMBL
Peking Union Medical College
Curated by ChEMBL
Affinity DataKi: 2nMAssay Description:Inhibition of wild type HIV1 protease using Arg-Glu (EDANS)-Ser-Gln-Asn-Tyr-Pro-Ile-Val-Gln-Lys(DABCYL)-Arg as substrate preincubated for 20 to 30 mi...More data for this Ligand-Target Pair
TargetGag-Pol polyprotein [489-587](Human immunodeficiency virus type 1)
Peking Union Medical College
Curated by ChEMBL
Peking Union Medical College
Curated by ChEMBL
Affinity DataKi: 2.60nMAssay Description:Inhibition of wild type HIV1 protease using Arg-Glu (EDANS)-Ser-Gln-Asn-Tyr-Pro-Ile-Val-Gln-Lys(DABCYL)-Arg as substrate preincubated for 20 to 30 mi...More data for this Ligand-Target Pair
TargetGag-Pol polyprotein [489-587](Human immunodeficiency virus type 1)
Peking Union Medical College
Curated by ChEMBL
Peking Union Medical College
Curated by ChEMBL
Affinity DataKi: 4.60nMAssay Description:Inhibition of wild type HIV1 protease using Arg-Glu (EDANS)-Ser-Gln-Asn-Tyr-Pro-Ile-Val-Gln-Lys(DABCYL)-Arg as substrate preincubated for 20 to 30 mi...More data for this Ligand-Target Pair
TargetGag-Pol polyprotein [489-587](Human immunodeficiency virus type 1)
Peking Union Medical College
Curated by ChEMBL
Peking Union Medical College
Curated by ChEMBL
Affinity DataKi: 5.5nMAssay Description:Inhibition of wild type HIV1 protease using Arg-Glu (EDANS)-Ser-Gln-Asn-Tyr-Pro-Ile-Val-Gln-Lys(DABCYL)-Arg as substrate preincubated for 20 to 30 mi...More data for this Ligand-Target Pair
TargetGag-Pol polyprotein [489-587](Human immunodeficiency virus type 1)
Peking Union Medical College
Curated by ChEMBL
Peking Union Medical College
Curated by ChEMBL
Affinity DataKi: 5.90nMAssay Description:Inhibition of wild type HIV1 protease using Arg-Glu (EDANS)-Ser-Gln-Asn-Tyr-Pro-Ile-Val-Gln-Lys(DABCYL)-Arg as substrate preincubated for 20 to 30 mi...More data for this Ligand-Target Pair
TargetGag-Pol polyprotein [489-587](Human immunodeficiency virus type 1)
Peking Union Medical College
Curated by ChEMBL
Peking Union Medical College
Curated by ChEMBL
Affinity DataKi: 10nMAssay Description:Inhibition of wild type HIV1 protease using Arg-Glu (EDANS)-Ser-Gln-Asn-Tyr-Pro-Ile-Val-Gln-Lys(DABCYL)-Arg as substrate preincubated for 20 to 30 mi...More data for this Ligand-Target Pair
TargetGag-Pol polyprotein [489-587](Human immunodeficiency virus type 1)
Peking Union Medical College
Curated by ChEMBL
Peking Union Medical College
Curated by ChEMBL
Affinity DataKi: 41nMAssay Description:Inhibition of wild type HIV1 protease using Arg-Glu (EDANS)-Ser-Gln-Asn-Tyr-Pro-Ile-Val-Gln-Lys(DABCYL)-Arg as substrate preincubated for 20 to 30 mi...More data for this Ligand-Target Pair
TargetGag-Pol polyprotein [489-587](Human immunodeficiency virus type 1)
Peking Union Medical College
Curated by ChEMBL
Peking Union Medical College
Curated by ChEMBL
Affinity DataKi: 44nMAssay Description:Inhibition of wild type HIV1 protease using Arg-Glu (EDANS)-Ser-Gln-Asn-Tyr-Pro-Ile-Val-Gln-Lys(DABCYL)-Arg as substrate preincubated for 20 to 30 mi...More data for this Ligand-Target Pair