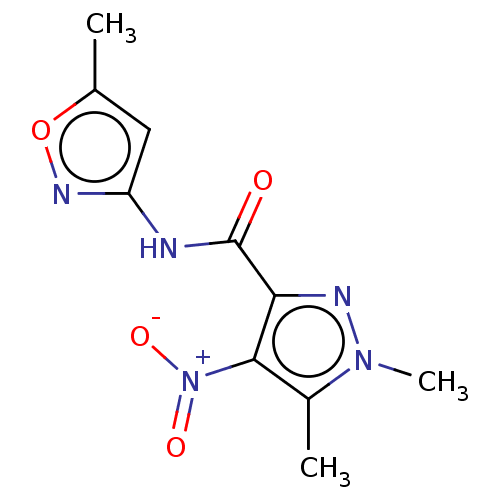
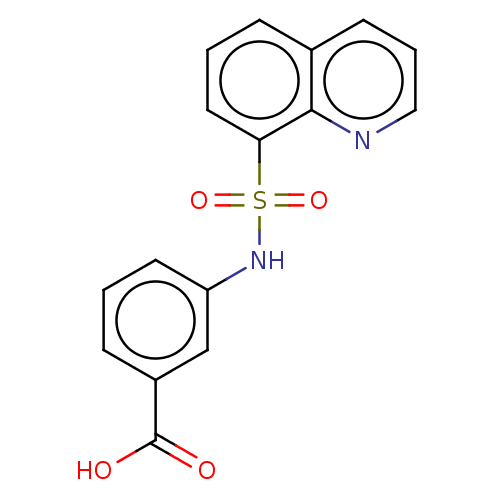
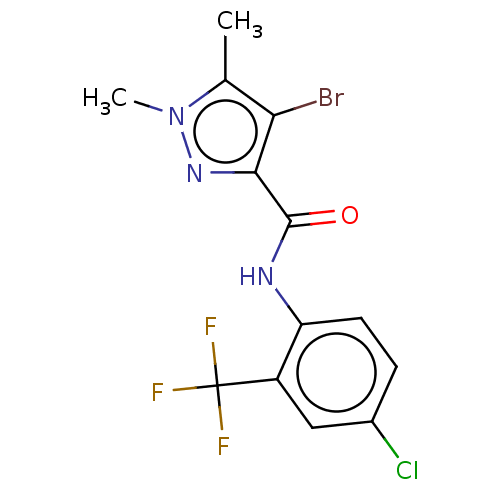
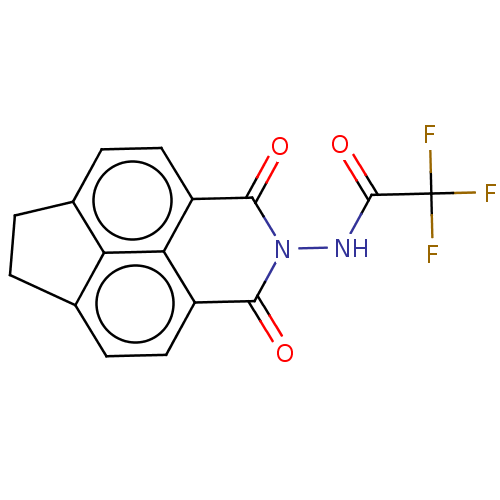
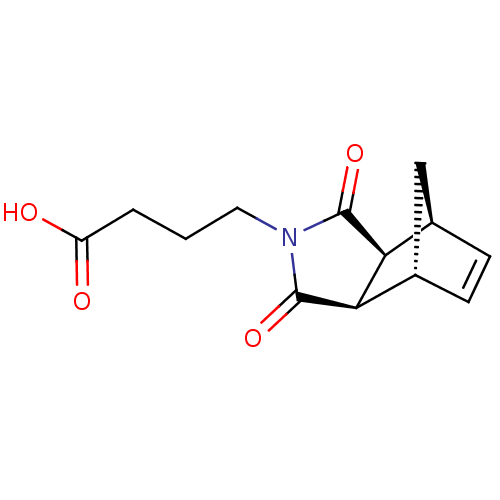
Report error Found 17 Enz. Inhib. hit(s) with all data for entry = 50013907
Affinity DataIC50: 4.09E+3nMAssay Description:Inhibition of recombinant human PHD2 (181 to 426 residues) using DLDLEMLAPYIPMDDDFQL peptide as substrate preincubated for 15 mins followed by substr...More data for this Ligand-Target Pair
Affinity DataIC50: 2.60E+4nMAssay Description:Inhibition of recombinant human PHD2 (181 to 426 residues) using DLDLEMLAPYIPMDDDFQL peptide as substrate preincubated for 15 mins followed by substr...More data for this Ligand-Target Pair
Affinity DataIC50: 7.77E+4nMAssay Description:Inhibition of recombinant human PHD2 (181 to 426 residues) using DLDLEMLAPYIPMDDDFQL peptide as substrate preincubated for 15 mins followed by substr...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+5nMAssay Description:Inhibition of recombinant human PHD2 (181 to 426 residues) using DLDLEMLAPYIPMDDDFQL peptide as substrate preincubated for 15 mins followed by substr...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+5nMAssay Description:Inhibition of recombinant human PHD2 (181 to 426 residues) using DLDLEMLAPYIPMDDDFQL peptide as substrate preincubated for 15 mins followed by substr...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+5nMAssay Description:Inhibition of recombinant human PHD2 (181 to 426 residues) using DLDLEMLAPYIPMDDDFQL peptide as substrate preincubated for 15 mins followed by substr...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+5nMAssay Description:Inhibition of recombinant human PHD2 (181 to 426 residues) using DLDLEMLAPYIPMDDDFQL peptide as substrate preincubated for 15 mins followed by substr...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+5nMAssay Description:Inhibition of recombinant human PHD2 (181 to 426 residues) using DLDLEMLAPYIPMDDDFQL peptide as substrate preincubated for 15 mins followed by substr...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+5nMAssay Description:Inhibition of recombinant human PHD2 (181 to 426 residues) using DLDLEMLAPYIPMDDDFQL peptide as substrate preincubated for 15 mins followed by substr...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+5nMAssay Description:Inhibition of recombinant human PHD2 (181 to 426 residues) using DLDLEMLAPYIPMDDDFQL peptide as substrate preincubated for 15 mins followed by substr...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+5nMAssay Description:Inhibition of recombinant human PHD2 (181 to 426 residues) using DLDLEMLAPYIPMDDDFQL peptide as substrate preincubated for 15 mins followed by substr...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+5nMAssay Description:Inhibition of recombinant human PHD2 (181 to 426 residues) using DLDLEMLAPYIPMDDDFQL peptide as substrate preincubated for 15 mins followed by substr...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+5nMAssay Description:Inhibition of recombinant human PHD2 (181 to 426 residues) using DLDLEMLAPYIPMDDDFQL peptide as substrate preincubated for 15 mins followed by substr...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+5nMAssay Description:Inhibition of recombinant human PHD2 (181 to 426 residues) using DLDLEMLAPYIPMDDDFQL peptide as substrate preincubated for 15 mins followed by substr...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+5nMAssay Description:Inhibition of recombinant human PHD2 (181 to 426 residues) using DLDLEMLAPYIPMDDDFQL peptide as substrate preincubated for 15 mins followed by substr...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+5nMAssay Description:Inhibition of recombinant human PHD2 (181 to 426 residues) using DLDLEMLAPYIPMDDDFQL peptide as substrate preincubated for 15 mins followed by substr...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+5nMAssay Description:Inhibition of recombinant human PHD2 (181 to 426 residues) using DLDLEMLAPYIPMDDDFQL peptide as substrate preincubated for 15 mins followed by substr...More data for this Ligand-Target Pair