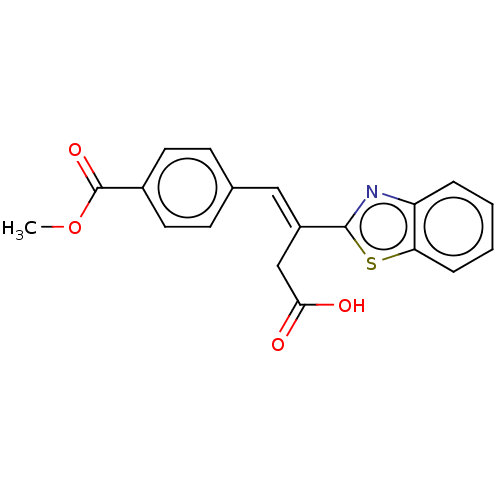
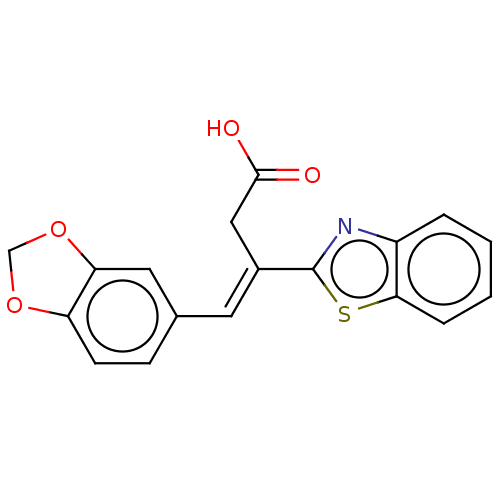
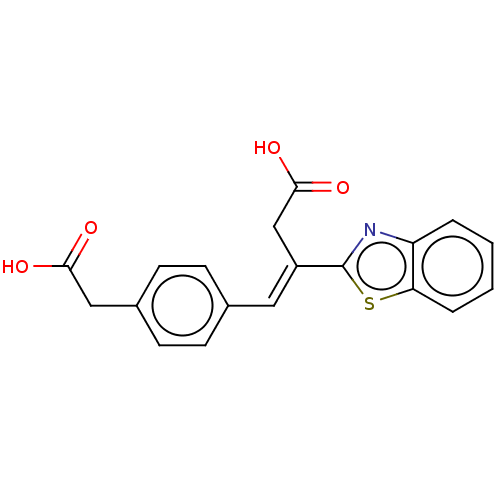
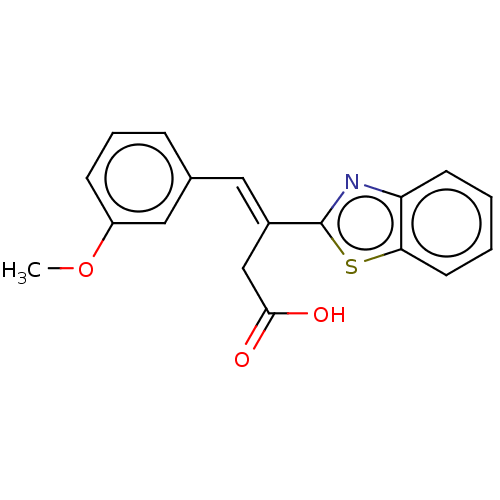
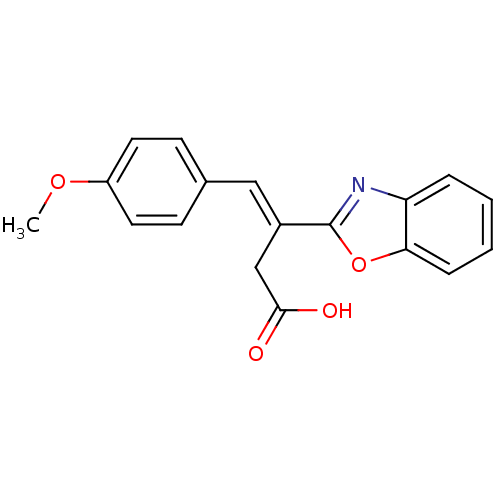
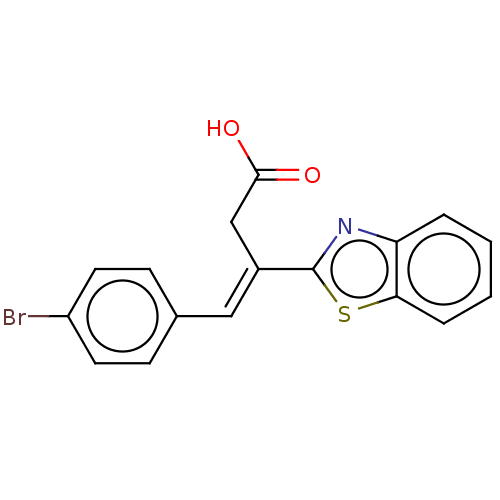
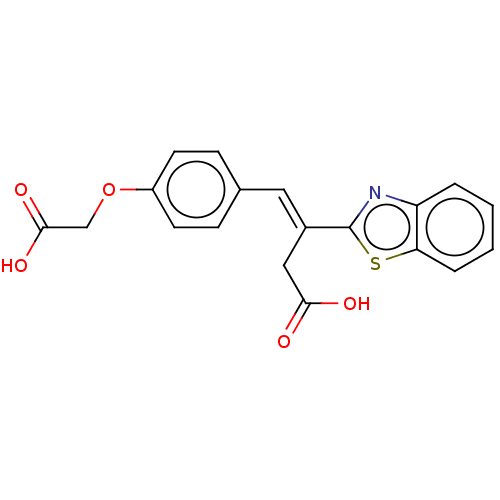
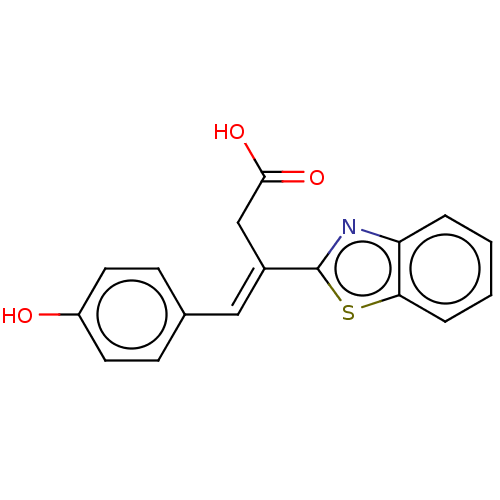
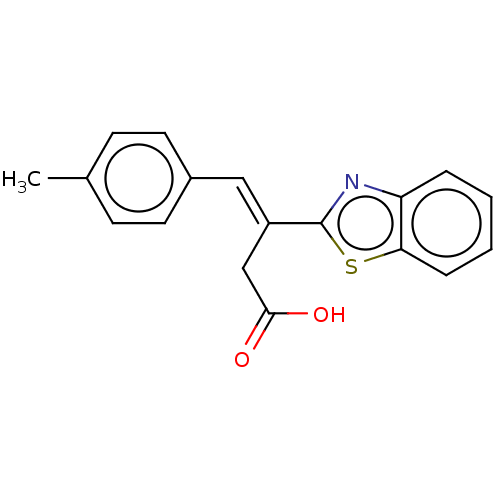
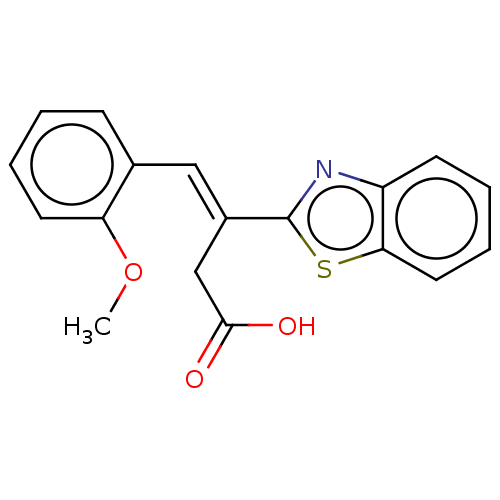
Report error Found 16 Enz. Inhib. hit(s) with all data for entry = 50013736
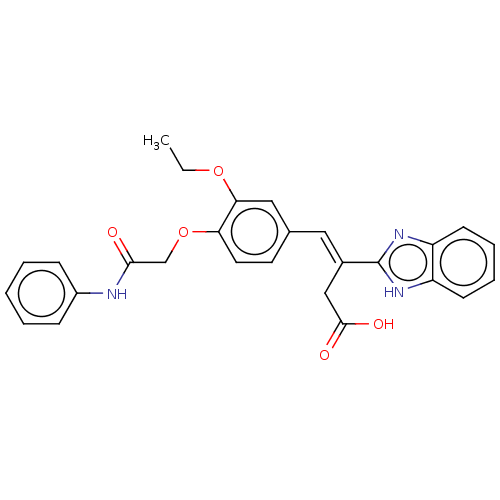
Affinity DataIC50: 2.30E+3nMAssay Description:Inhibition of Glyoxalase-1 (unknown origin) assessed as inhibition of S-D-lactoylglutathione formation using MG and reduced glutathione as substrate ...More data for this Ligand-Target Pair
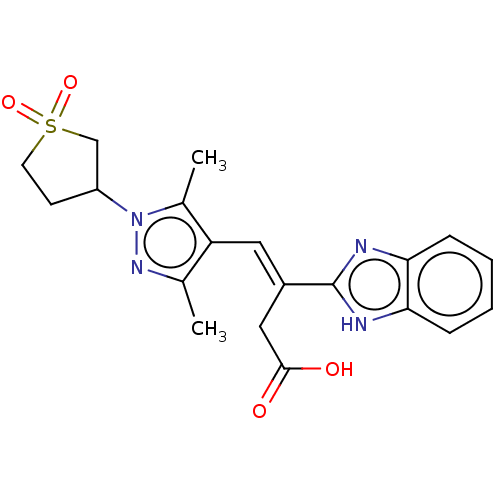
Affinity DataIC50: 2.30E+3nMAssay Description:Inhibition of Glyoxalase-1 (unknown origin) assessed as inhibition of S-D-lactoylglutathione formation using MG and reduced glutathione as substrate ...More data for this Ligand-Target Pair
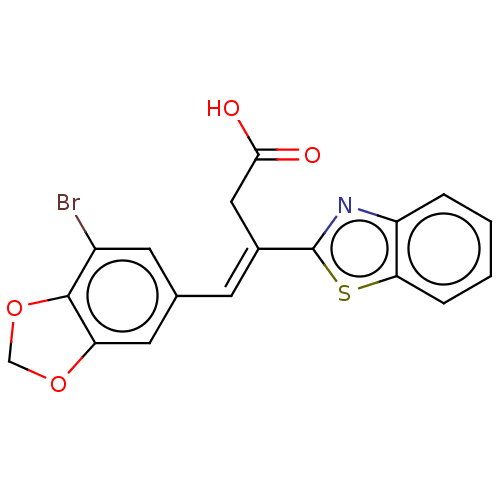
Affinity DataIC50: 2.80E+3nMAssay Description:Inhibition of Glyoxalase-1 (unknown origin) assessed as inhibition of S-D-lactoylglutathione formation using MG and reduced glutathione as substrate ...More data for this Ligand-Target Pair
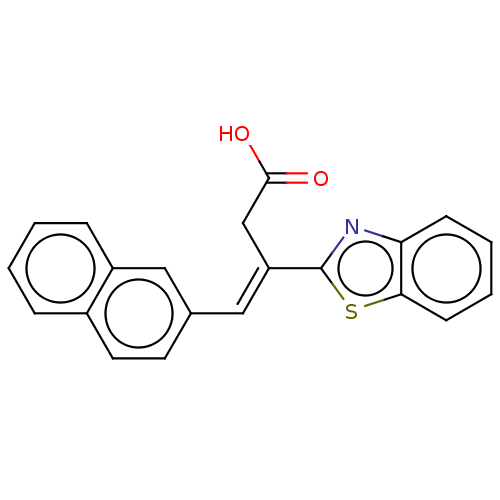
Affinity DataIC50: 2.90E+3nMAssay Description:Inhibition of Glyoxalase-1 (unknown origin) assessed as inhibition of S-D-lactoylglutathione formation using MG and reduced glutathione as substrate ...More data for this Ligand-Target Pair
Affinity DataIC50: 3.00E+3nMAssay Description:Inhibition of Glyoxalase-1 (unknown origin) assessed as inhibition of S-D-lactoylglutathione formation using MG and reduced glutathione as substrate ...More data for this Ligand-Target Pair
Affinity DataIC50: 3.20E+3nMAssay Description:Inhibition of Glyoxalase-1 (unknown origin) assessed as inhibition of S-D-lactoylglutathione formation using MG and reduced glutathione as substrate ...More data for this Ligand-Target Pair
Affinity DataIC50: 3.30E+3nMAssay Description:Inhibition of Glyoxalase-1 (unknown origin) assessed as inhibition of S-D-lactoylglutathione formation using MG and reduced glutathione as substrate ...More data for this Ligand-Target Pair
Affinity DataIC50: 3.40E+3nMAssay Description:Inhibition of Glyoxalase-1 (unknown origin) assessed as inhibition of S-D-lactoylglutathione formation using MG and reduced glutathione as substrate ...More data for this Ligand-Target Pair
Affinity DataIC50: 4.40E+3nMAssay Description:Inhibition of Glyoxalase-1 (unknown origin) assessed as inhibition of S-D-lactoylglutathione formation using MG and reduced glutathione as substrate ...More data for this Ligand-Target Pair
Affinity DataIC50: 4.60E+3nMAssay Description:Inhibition of Glyoxalase-1 (unknown origin) assessed as inhibition of S-D-lactoylglutathione formation using MG and reduced glutathione as substrate ...More data for this Ligand-Target Pair
Affinity DataIC50: 5.30E+3nMAssay Description:Inhibition of Glyoxalase-1 (unknown origin) assessed as inhibition of S-D-lactoylglutathione formation using MG and reduced glutathione as substrate ...More data for this Ligand-Target Pair
Affinity DataIC50: 5.40E+3nMAssay Description:Inhibition of Glyoxalase-1 (unknown origin) assessed as inhibition of S-D-lactoylglutathione formation using MG and reduced glutathione as substrate ...More data for this Ligand-Target Pair
Affinity DataIC50: 5.50E+3nMAssay Description:Inhibition of Glyoxalase-1 (unknown origin) assessed as inhibition of S-D-lactoylglutathione formation using MG and reduced glutathione as substrate ...More data for this Ligand-Target Pair
Affinity DataIC50: 8.20E+3nMAssay Description:Inhibition of Glyoxalase-1 (unknown origin) assessed as inhibition of S-D-lactoylglutathione formation using MG and reduced glutathione as substrate ...More data for this Ligand-Target Pair
Affinity DataIC50: 9.50E+3nMAssay Description:Inhibition of Glyoxalase-1 (unknown origin) assessed as inhibition of S-D-lactoylglutathione formation using MG and reduced glutathione as substrate ...More data for this Ligand-Target Pair
Affinity DataIC50: 4.66E+4nMAssay Description:Inhibition of Glyoxalase-1 (unknown origin) assessed as inhibition of S-D-lactoylglutathione formation using MG and reduced glutathione as substrate ...More data for this Ligand-Target Pair
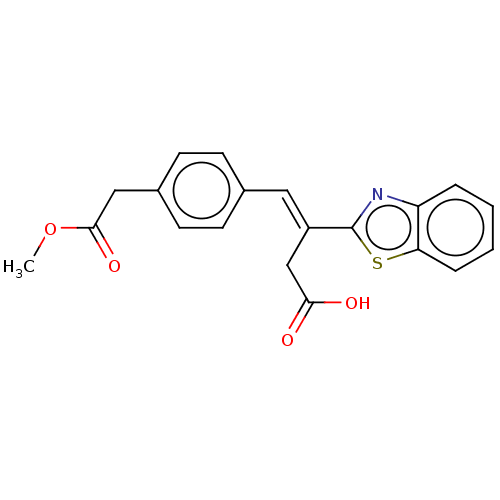
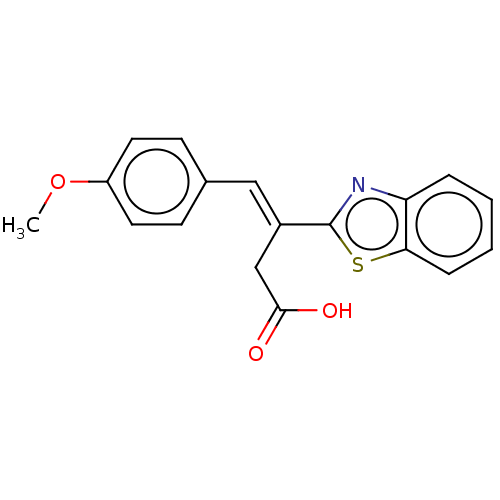
 3D Structure (crystal)
3D Structure (crystal)