Report error Found 27 Enz. Inhib. hit(s) with all data for entry = 50018507
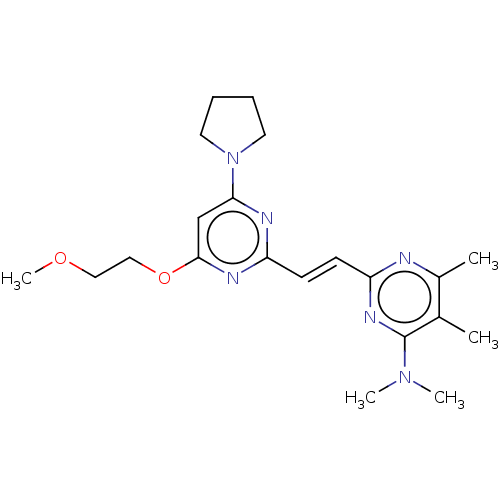
TargetcAMP and cAMP-inhibited cGMP 3',5'-cyclic phosphodiesterase 10A(Human)
Nirma University
Curated by ChEMBL
Nirma University
Curated by ChEMBL
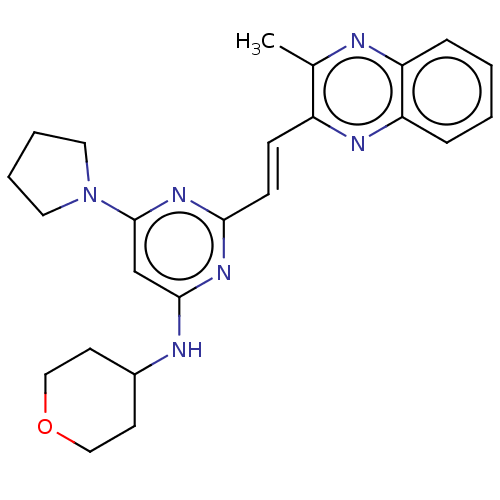
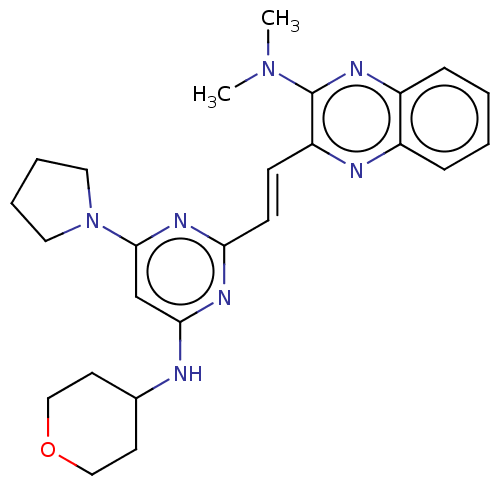
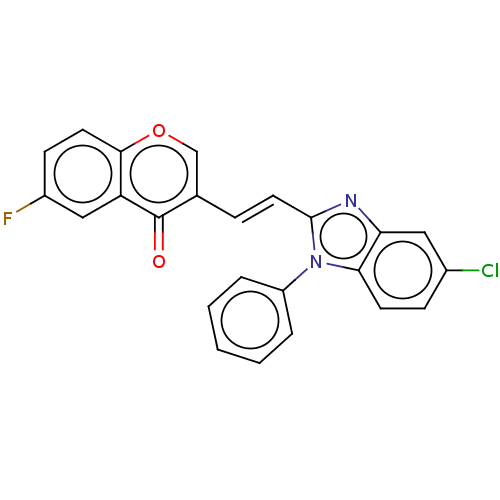
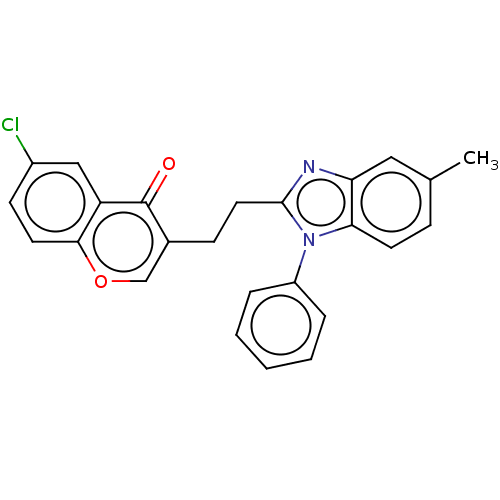
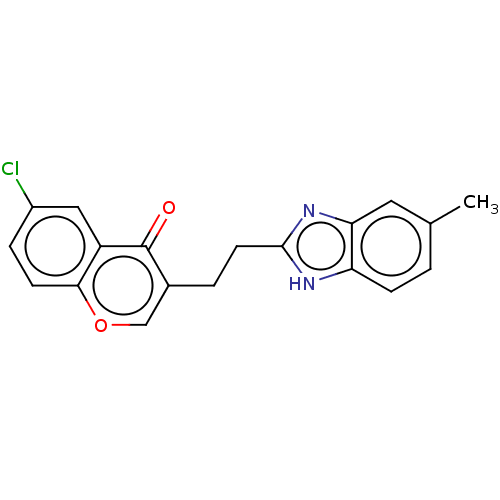
Affinity DataIC50: 0.0600nMAssay Description:Inhibition of PDE10A (unknown origin)More data for this Ligand-Target Pair
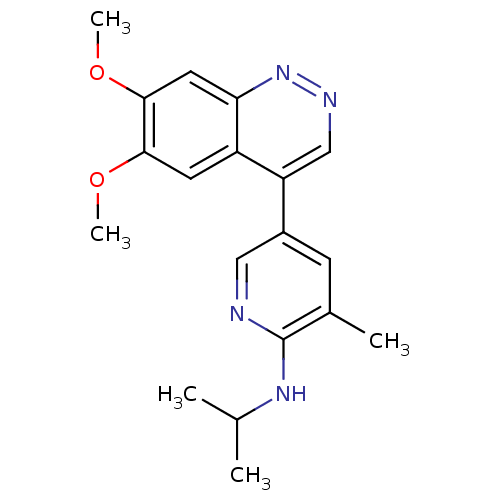
TargetcAMP and cAMP-inhibited cGMP 3',5'-cyclic phosphodiesterase 10A(Human)
Nirma University
Curated by ChEMBL
Nirma University
Curated by ChEMBL
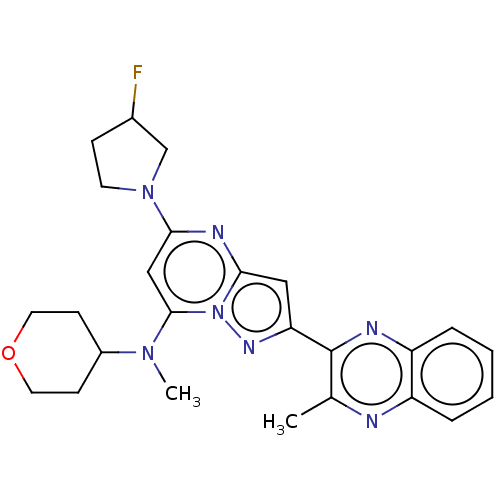
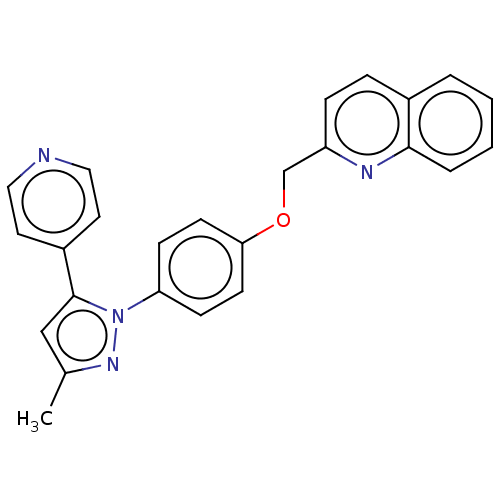
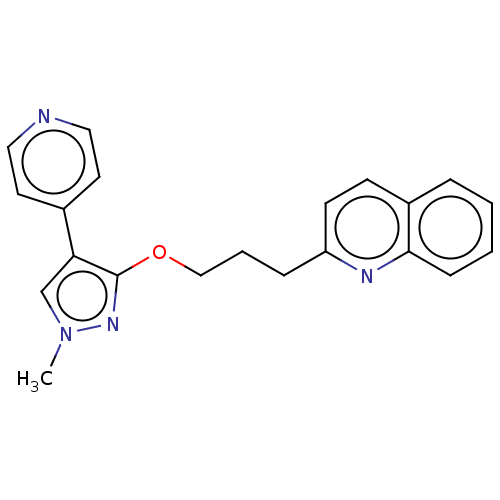
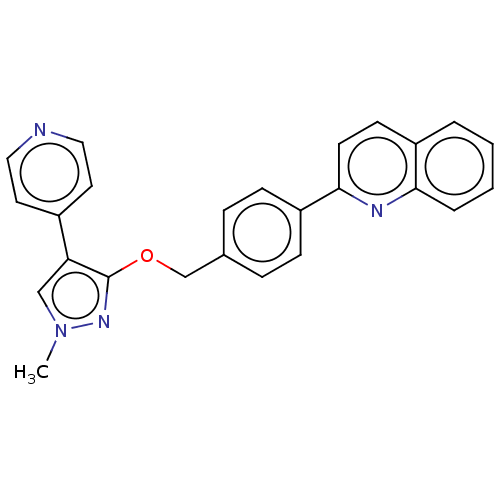
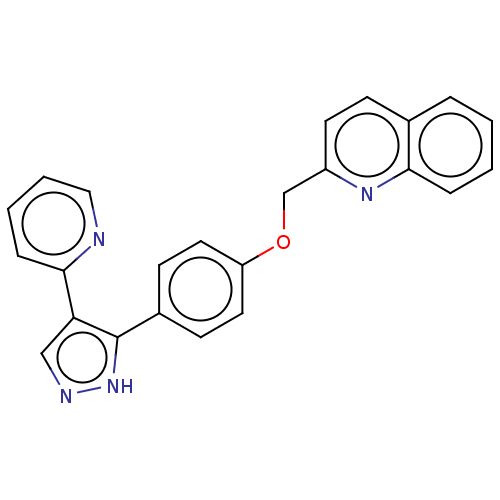
Affinity DataIC50: 0.0600nMAssay Description:Inhibition of PDE10A (unknown origin)More data for this Ligand-Target Pair
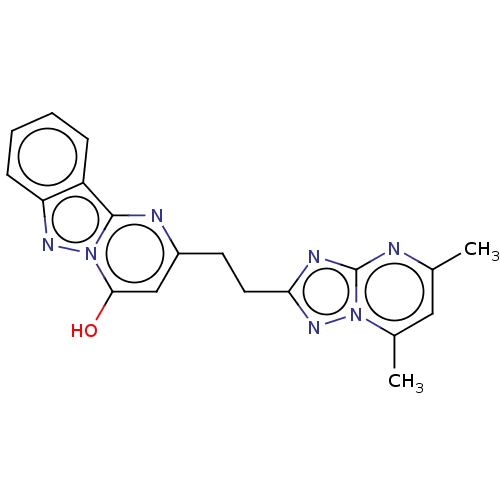
TargetcAMP and cAMP-inhibited cGMP 3',5'-cyclic phosphodiesterase 10A(Human)
Nirma University
Curated by ChEMBL
Nirma University
Curated by ChEMBL
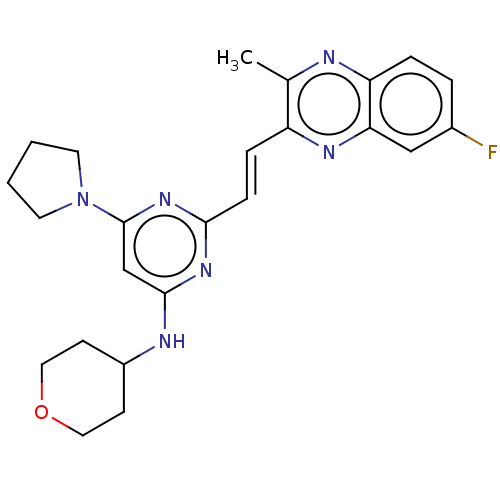
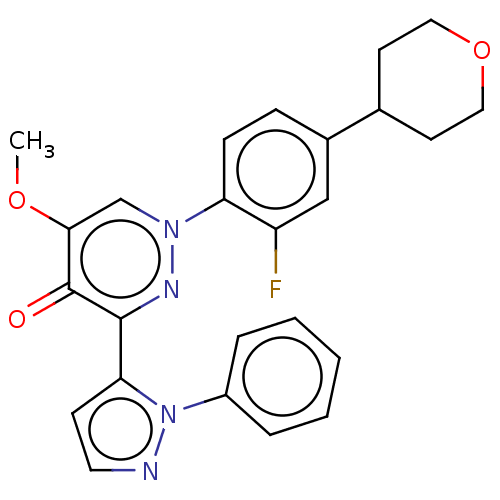
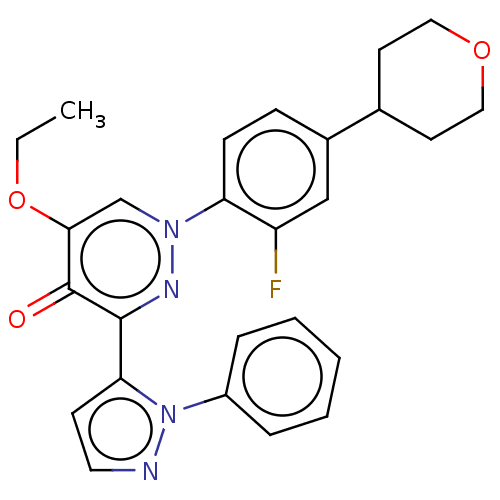
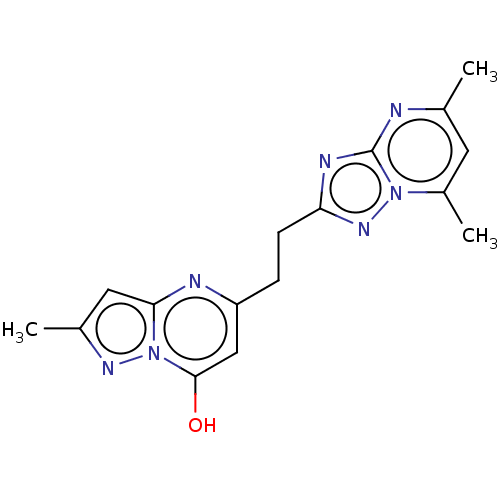
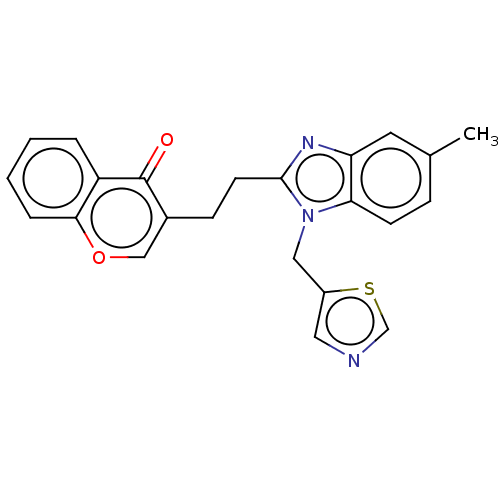
Affinity DataIC50: 0.140nMAssay Description:Inhibition of PDE10A (unknown origin)More data for this Ligand-Target Pair
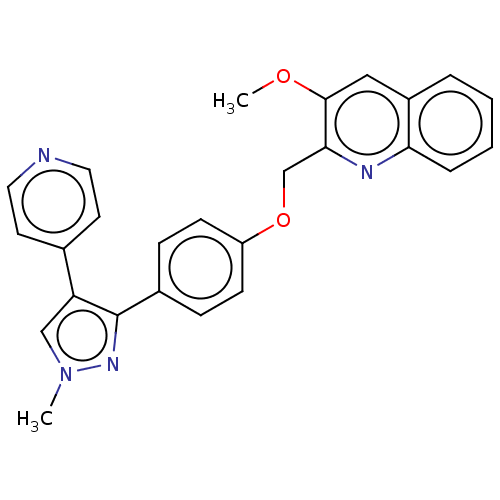
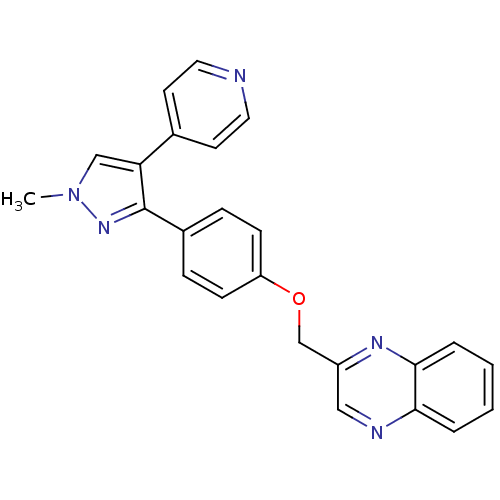
Affinity DataIC50: 0.160nMAssay Description:Inhibition of PDE5 (unknown origin)More data for this Ligand-Target Pair
TargetcAMP and cAMP-inhibited cGMP 3',5'-cyclic phosphodiesterase 10A(Human)
Nirma University
Curated by ChEMBL
Nirma University
Curated by ChEMBL
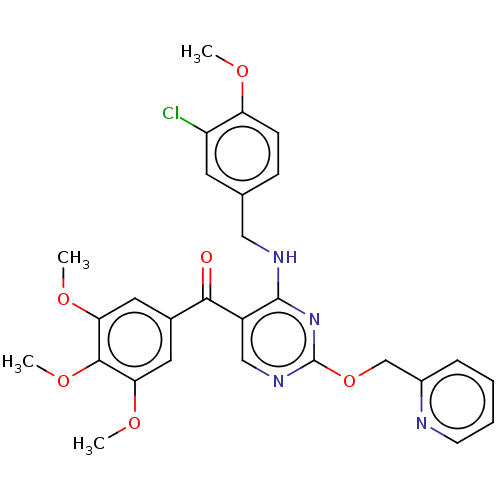
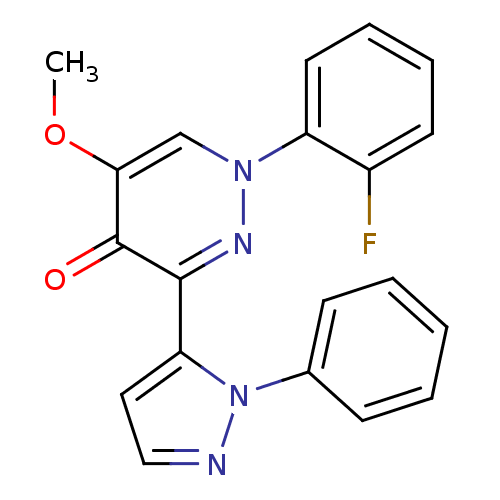
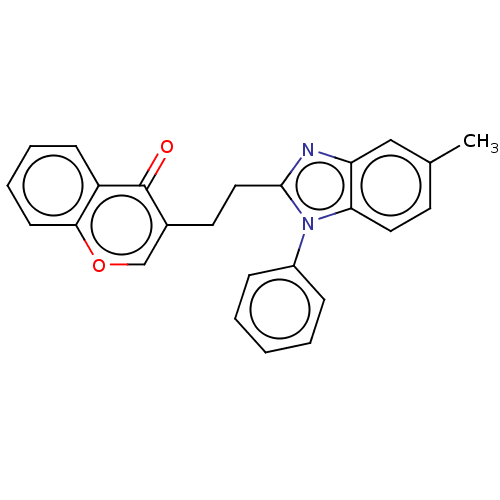
Affinity DataIC50: 0.220nMAssay Description:Inhibition of PDE10A (unknown origin)More data for this Ligand-Target Pair
TargetcAMP and cAMP-inhibited cGMP 3',5'-cyclic phosphodiesterase 10A(Human)
Nirma University
Curated by ChEMBL
Nirma University
Curated by ChEMBL
Affinity DataIC50: 0.5nMAssay Description:Inhibition of PDE10A (unknown origin)More data for this Ligand-Target Pair
TargetcAMP and cAMP-inhibited cGMP 3',5'-cyclic phosphodiesterase 10A(Human)
Nirma University
Curated by ChEMBL
Nirma University
Curated by ChEMBL
Affinity DataIC50: 0.770nMAssay Description:Inhibition of PDE10A (unknown origin) by scintillation proximity assayMore data for this Ligand-Target Pair
TargetcAMP and cAMP-inhibited cGMP 3',5'-cyclic phosphodiesterase 10A(Rat)
Nirma University
Curated by ChEMBL
Nirma University
Curated by ChEMBL
Affinity DataKd: 1.30nMAssay Description:Binding affinity to PDE10A in rat striatum homogenateMore data for this Ligand-Target Pair
TargetcAMP and cAMP-inhibited cGMP 3',5'-cyclic phosphodiesterase 10A(Human)
Nirma University
Curated by ChEMBL
Nirma University
Curated by ChEMBL
Affinity DataIC50: 1.80nMAssay Description:Inhibition of PDE10A (unknown origin)More data for this Ligand-Target Pair
TargetcAMP and cAMP-inhibited cGMP 3',5'-cyclic phosphodiesterase 10A(Human)
Nirma University
Curated by ChEMBL
Nirma University
Curated by ChEMBL
Affinity DataIC50: 1.80nMAssay Description:Inhibition of PDE10A2 (unknown origin)More data for this Ligand-Target Pair
TargetcAMP and cAMP-inhibited cGMP 3',5'-cyclic phosphodiesterase 10A(Human)
Nirma University
Curated by ChEMBL
Nirma University
Curated by ChEMBL
Affinity DataIC50: 1.90nMAssay Description:Inhibition of PDE10A (unknown origin)More data for this Ligand-Target Pair
TargetcAMP and cAMP-inhibited cGMP 3',5'-cyclic phosphodiesterase 10A(Human)
Nirma University
Curated by ChEMBL
Nirma University
Curated by ChEMBL
Affinity DataIC50: 2nMAssay Description:Inhibition of human PDE10A (449 to 789 residues) expressed in Escherichia coli BL21(DE3) using cAMP as substrate incubated for 30 mins by HTRF analys...More data for this Ligand-Target Pair
TargetcAMP and cAMP-inhibited cGMP 3',5'-cyclic phosphodiesterase 10A(Rat)
Nirma University
Curated by ChEMBL
Nirma University
Curated by ChEMBL
Affinity DataKd: 4.70nMAssay Description:Binding affinity to PDE10A in rat brain nucleus accumbens by saturation binding assayMore data for this Ligand-Target Pair
TargetcAMP and cAMP-inhibited cGMP 3',5'-cyclic phosphodiesterase 10A(Human)
Nirma University
Curated by ChEMBL
Nirma University
Curated by ChEMBL
Affinity DataIC50: 5.10nMAssay Description:Inhibition of PDE10A (unknown origin)More data for this Ligand-Target Pair
TargetcAMP and cAMP-inhibited cGMP 3',5'-cyclic phosphodiesterase 10A(Human)
Nirma University
Curated by ChEMBL
Nirma University
Curated by ChEMBL
Affinity DataIC50: 6.5nMAssay Description:Inhibition of PDE10A (unknown origin)More data for this Ligand-Target Pair
TargetcAMP and cAMP-inhibited cGMP 3',5'-cyclic phosphodiesterase 10A(Human)
Nirma University
Curated by ChEMBL
Nirma University
Curated by ChEMBL
Affinity DataIC50: 8.90nMAssay Description:Inhibition of PDE10A (unknown origin)More data for this Ligand-Target Pair
TargetcAMP and cAMP-inhibited cGMP 3',5'-cyclic phosphodiesterase 10A(Human)
Nirma University
Curated by ChEMBL
Nirma University
Curated by ChEMBL
Affinity DataIC50: 9.70nMAssay Description:Inhibition of PDE10A2 (unknown origin)More data for this Ligand-Target Pair
TargetcAMP and cAMP-inhibited cGMP 3',5'-cyclic phosphodiesterase 10A(Rat)
Nirma University
Curated by ChEMBL
Nirma University
Curated by ChEMBL
Affinity DataKd: 12nMAssay Description:Binding affinity to PDE10A in rat brain caudate-putamen by saturation binding assayMore data for this Ligand-Target Pair
TargetcAMP and cAMP-inhibited cGMP 3',5'-cyclic phosphodiesterase 10A(Human)
Nirma University
Curated by ChEMBL
Nirma University
Curated by ChEMBL
Affinity DataIC50: 12nMAssay Description:Inhibition of PDE10A (unknown origin)More data for this Ligand-Target Pair
TargetcAMP and cAMP-inhibited cGMP 3',5'-cyclic phosphodiesterase 10A(Human)
Nirma University
Curated by ChEMBL
Nirma University
Curated by ChEMBL
Affinity DataIC50: 22nMAssay Description:Inhibition of PDE10A (unknown origin)More data for this Ligand-Target Pair
TargetcAMP and cAMP-inhibited cGMP 3',5'-cyclic phosphodiesterase 10A(Human)
Nirma University
Curated by ChEMBL
Nirma University
Curated by ChEMBL
Affinity DataIC50: 29nMAssay Description:Inhibition of PDE10A (unknown origin)More data for this Ligand-Target Pair
TargetcAMP and cAMP-inhibited cGMP 3',5'-cyclic phosphodiesterase 10A(Human)
Nirma University
Curated by ChEMBL
Nirma University
Curated by ChEMBL
Affinity DataIC50: 44nMAssay Description:Inhibition of human PDE10A (449 to 789 residues) expressed in Escherichia coli BL21(DE3) using cAMP as substrate incubated for 30 mins by HTRF analys...More data for this Ligand-Target Pair
TargetcAMP and cAMP-inhibited cGMP 3',5'-cyclic phosphodiesterase 10A(Human)
Nirma University
Curated by ChEMBL
Nirma University
Curated by ChEMBL
Affinity DataIC50: 52nMAssay Description:Inhibition of PDE10A (unknown origin)More data for this Ligand-Target Pair
TargetcAMP and cAMP-inhibited cGMP 3',5'-cyclic phosphodiesterase 10A(Human)
Nirma University
Curated by ChEMBL
Nirma University
Curated by ChEMBL
Affinity DataIC50: 112nMAssay Description:Inhibition of PDE10A (unknown origin)More data for this Ligand-Target Pair
TargetcAMP and cAMP-inhibited cGMP 3',5'-cyclic phosphodiesterase 10A(Human)
Nirma University
Curated by ChEMBL
Nirma University
Curated by ChEMBL
Affinity DataIC50: 280nMAssay Description:Inhibition of PDE10A (unknown origin)More data for this Ligand-Target Pair
TargetcAMP and cAMP-inhibited cGMP 3',5'-cyclic phosphodiesterase 10A(Human)
Nirma University
Curated by ChEMBL
Nirma University
Curated by ChEMBL
Affinity DataIC50: 371nMAssay Description:Inhibition of PDE10A (unknown origin)More data for this Ligand-Target Pair
TargetcAMP and cAMP-inhibited cGMP 3',5'-cyclic phosphodiesterase 10A(Human)
Nirma University
Curated by ChEMBL
Nirma University
Curated by ChEMBL
Affinity DataIC50: 500nMAssay Description:Inhibition of PDE10A (unknown origin)More data for this Ligand-Target Pair