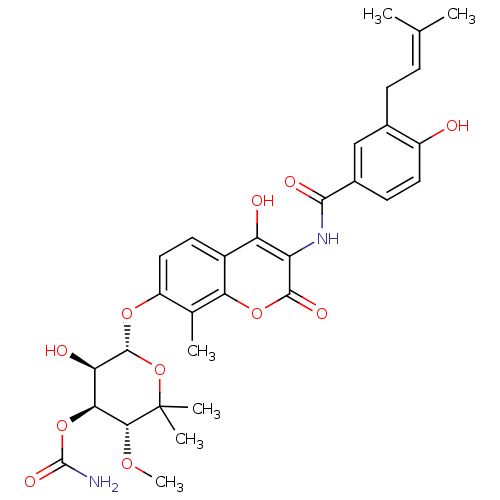
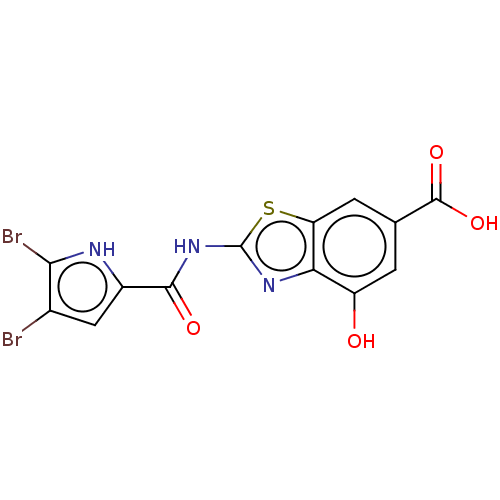
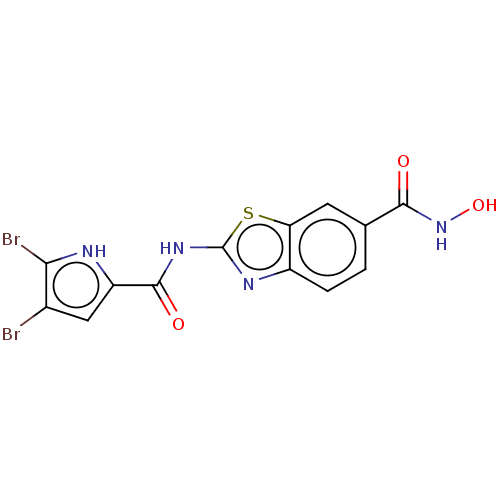
Report error Found 70 Enz. Inhib. hit(s) with all data for entry = 50013705
TargetDNA gyrase A/A/B/B tetramer(Escherichia coli (strain K12))
University of Ljubljana
Curated by ChEMBL
University of Ljubljana
Curated by ChEMBL
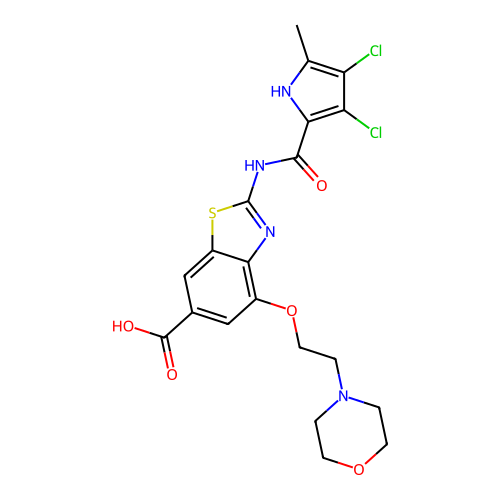
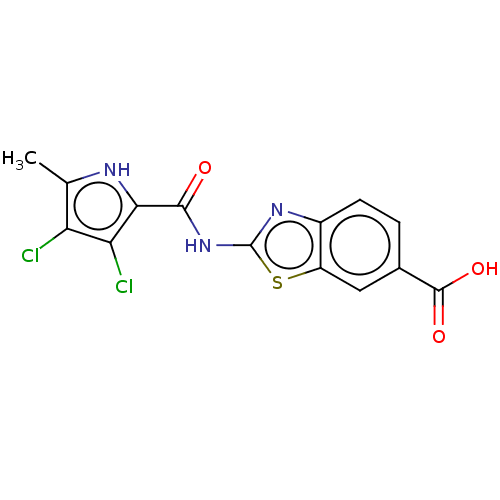
Affinity DataIC50: 10nMAssay Description:Inhibition of Escherichia coli DNA gyrase assessed as reduction in relaxation of supercoiled pNO1 DNA measured after 30 mins by fluorescence assayMore data for this Ligand-Target Pair
Affinity DataIC50: 10nMAssay Description:Inhibition of Staphylococcus aureus DNA gyrase assessed reduction in relaxation of supercoiled pNO1 DNA measured after 30 mins by fluorescence assayMore data for this Ligand-Target Pair
TargetDNA gyrase A/A/B/B tetramer(Escherichia coli (strain K12))
University of Ljubljana
Curated by ChEMBL
University of Ljubljana
Curated by ChEMBL
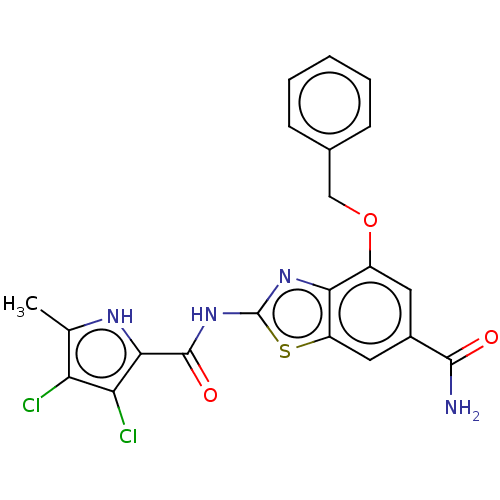
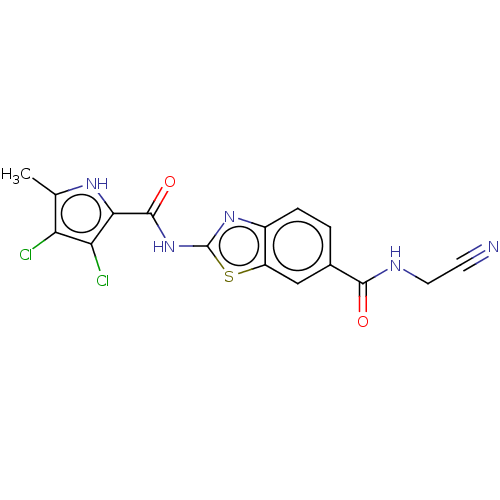
Affinity DataIC50: 10nMAssay Description:Inhibition of Escherichia coli DNA gyrase assessed as reduction in relaxation of supercoiled pNO1 DNA measured after 30 mins by fluorescence assayMore data for this Ligand-Target Pair
Affinity DataIC50: 10nMAssay Description:Inhibition of Staphylococcus aureus DNA gyrase assessed reduction in relaxation of supercoiled pNO1 DNA measured after 30 mins by fluorescence assayMore data for this Ligand-Target Pair
TargetDNA gyrase A/A/B/B tetramer(Escherichia coli (strain K12))
University of Ljubljana
Curated by ChEMBL
University of Ljubljana
Curated by ChEMBL
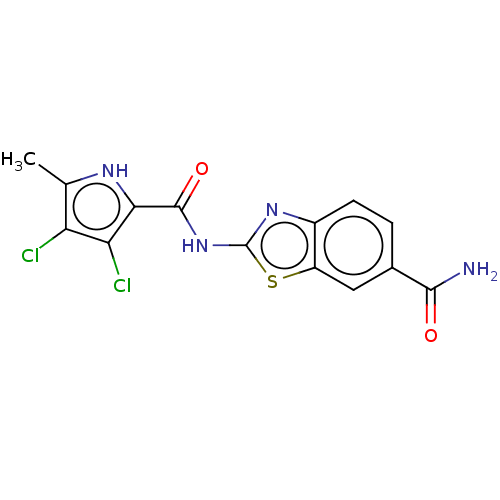
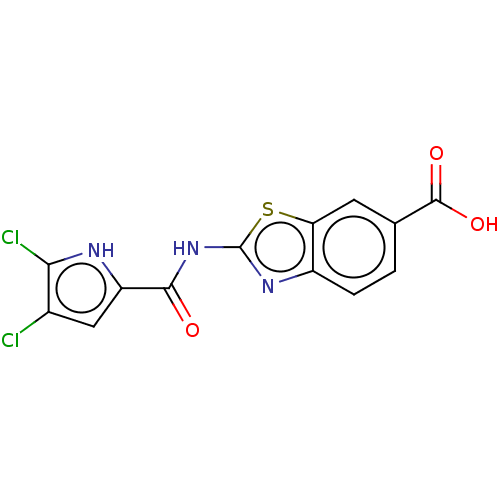
Affinity DataIC50: 10nMAssay Description:Inhibition of Escherichia coli DNA gyrase assessed as reduction in relaxation of supercoiled pNO1 DNA measured after 30 mins by fluorescence assayMore data for this Ligand-Target Pair
Affinity DataIC50: 10nMAssay Description:Inhibition of Staphylococcus aureus DNA gyrase assessed reduction in relaxation of supercoiled pNO1 DNA measured after 30 mins by fluorescence assayMore data for this Ligand-Target Pair
TargetDNA gyrase A/A/B/B tetramer(Escherichia coli (strain K12))
University of Ljubljana
Curated by ChEMBL
University of Ljubljana
Curated by ChEMBL
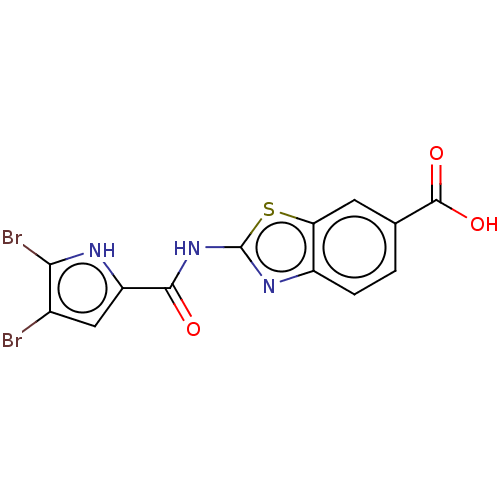
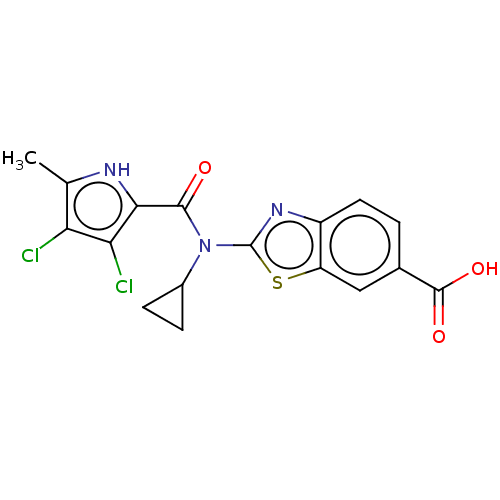
Affinity DataIC50: 10nMAssay Description:Inhibition of Escherichia coli DNA gyrase assessed as reduction in relaxation of supercoiled pNO1 DNA measured after 30 mins by fluorescence assayMore data for this Ligand-Target Pair
Affinity DataIC50: 10nMAssay Description:Inhibition of Staphylococcus aureus DNA gyrase assessed reduction in relaxation of supercoiled pNO1 DNA measured after 30 mins by fluorescence assayMore data for this Ligand-Target Pair
TargetDNA gyrase A/A/B/B tetramer(Escherichia coli (strain K12))
University of Ljubljana
Curated by ChEMBL
University of Ljubljana
Curated by ChEMBL
Affinity DataIC50: 10nMAssay Description:Inhibition of Escherichia coli DNA gyrase assessed as reduction in relaxation of supercoiled pNO1 DNA measured after 30 mins by fluorescence assayMore data for this Ligand-Target Pair
Affinity DataIC50: 10nMAssay Description:Inhibition of Staphylococcus aureus DNA gyrase assessed reduction in relaxation of supercoiled pNO1 DNA measured after 30 mins by fluorescence assayMore data for this Ligand-Target Pair
TargetDNA gyrase A/A/B/B tetramer(Escherichia coli (strain K12))
University of Ljubljana
Curated by ChEMBL
University of Ljubljana
Curated by ChEMBL
Affinity DataIC50: 10nMAssay Description:Inhibition of Escherichia coli DNA gyrase assessed as reduction in relaxation of supercoiled pNO1 DNA measured after 30 mins by fluorescence assayMore data for this Ligand-Target Pair
TargetDNA gyrase A/A/B/B tetramer(Escherichia coli (strain K12))
University of Ljubljana
Curated by ChEMBL
University of Ljubljana
Curated by ChEMBL
Affinity DataIC50: 10nMAssay Description:Inhibition of Escherichia coli DNA gyrase assessed as reduction in relaxation of supercoiled pNO1 DNA measured after 30 mins by fluorescence assayMore data for this Ligand-Target Pair
TargetDNA gyrase A/A/B/B tetramer(Escherichia coli (strain K12))
University of Ljubljana
Curated by ChEMBL
University of Ljubljana
Curated by ChEMBL
Affinity DataIC50: 10nMAssay Description:Inhibition of Escherichia coli DNA gyrase assessed as reduction in relaxation of supercoiled pNO1 DNA measured after 30 mins by fluorescence assayMore data for this Ligand-Target Pair
TargetDNA gyrase A/A/B/B tetramer(Escherichia coli (strain K12))
University of Ljubljana
Curated by ChEMBL
University of Ljubljana
Curated by ChEMBL
Affinity DataIC50: 10nMAssay Description:Inhibition of Escherichia coli DNA gyrase assessed as reduction in relaxation of supercoiled pNO1 DNA measured after 30 mins by fluorescence assayMore data for this Ligand-Target Pair
TargetDNA gyrase A/A/B/B tetramer(Escherichia coli (strain K12))
University of Ljubljana
Curated by ChEMBL
University of Ljubljana
Curated by ChEMBL
Affinity DataIC50: 10nMAssay Description:Inhibition of Escherichia coli DNA gyrase assessed as reduction in relaxation of supercoiled pNO1 DNA measured after 30 mins by fluorescence assayMore data for this Ligand-Target Pair
TargetDNA gyrase A/A/B/B tetramer(Escherichia coli (strain K12))
University of Ljubljana
Curated by ChEMBL
University of Ljubljana
Curated by ChEMBL
Affinity DataIC50: 10nMAssay Description:Inhibition of Escherichia coli DNA gyrase assessed as reduction in relaxation of supercoiled pNO1 DNA measured after 30 mins by fluorescence assayMore data for this Ligand-Target Pair
TargetDNA gyrase A/A/B/B tetramer(Escherichia coli (strain K12))
University of Ljubljana
Curated by ChEMBL
University of Ljubljana
Curated by ChEMBL
Affinity DataIC50: 10nMAssay Description:Inhibition of Escherichia coli DNA gyrase assessed as reduction in relaxation of supercoiled pNO1 DNA measured after 30 mins by fluorescence assayMore data for this Ligand-Target Pair
TargetDNA gyrase A/A/B/B tetramer(Escherichia coli (strain K12))
University of Ljubljana
Curated by ChEMBL
University of Ljubljana
Curated by ChEMBL
Affinity DataIC50: 10nMAssay Description:Inhibition of Escherichia coli DNA gyrase assessed as reduction in relaxation of supercoiled pNO1 DNA measured after 30 mins by fluorescence assayMore data for this Ligand-Target Pair
TargetDNA gyrase A/A/B/B tetramer(Escherichia coli (strain K12))
University of Ljubljana
Curated by ChEMBL
University of Ljubljana
Curated by ChEMBL
Affinity DataIC50: 10nMAssay Description:Inhibition of Escherichia coli DNA gyrase assessed as reduction in relaxation of supercoiled pNO1 DNA measured after 30 mins by fluorescence assayMore data for this Ligand-Target Pair
TargetDNA gyrase A/A/B/B tetramer(Escherichia coli (strain K12))
University of Ljubljana
Curated by ChEMBL
University of Ljubljana
Curated by ChEMBL
Affinity DataIC50: 10nMAssay Description:Inhibition of Escherichia coli DNA gyrase assessed as reduction in relaxation of supercoiled pNO1 DNA measured after 30 mins by fluorescence assayMore data for this Ligand-Target Pair
TargetDNA gyrase A/A/B/B tetramer(Escherichia coli (strain K12))
University of Ljubljana
Curated by ChEMBL
University of Ljubljana
Curated by ChEMBL
Affinity DataIC50: 10nMAssay Description:Inhibition of Escherichia coli DNA gyrase assessed as reduction in relaxation of supercoiled pNO1 DNA measured after 30 mins by fluorescence assayMore data for this Ligand-Target Pair
TargetDNA gyrase A/A/B/B tetramer(Escherichia coli (strain K12))
University of Ljubljana
Curated by ChEMBL
University of Ljubljana
Curated by ChEMBL
Affinity DataIC50: 10nMAssay Description:Inhibition of Escherichia coli DNA gyrase assessed as reduction in relaxation of supercoiled pNO1 DNA measured after 30 mins by fluorescence assayMore data for this Ligand-Target Pair
TargetDNA gyrase A/A/B/B tetramer(Escherichia coli (strain K12))
University of Ljubljana
Curated by ChEMBL
University of Ljubljana
Curated by ChEMBL
Affinity DataIC50: 10nMAssay Description:Inhibition of Escherichia coli DNA gyrase assessed as reduction in relaxation of supercoiled pNO1 DNA measured after 30 mins by fluorescence assayMore data for this Ligand-Target Pair
TargetDNA gyrase A/A/B/B tetramer(Escherichia coli (strain K12))
University of Ljubljana
Curated by ChEMBL
University of Ljubljana
Curated by ChEMBL
Affinity DataIC50: 10nMAssay Description:Inhibition of Escherichia coli DNA gyrase assessed as reduction in relaxation of supercoiled pNO1 DNA measured after 30 mins by fluorescence assayMore data for this Ligand-Target Pair
TargetDNA gyrase A/A/B/B tetramer(Escherichia coli (strain K12))
University of Ljubljana
Curated by ChEMBL
University of Ljubljana
Curated by ChEMBL
Affinity DataIC50: 11nMAssay Description:Inhibition of Escherichia coli DNA gyrase assessed as reduction in relaxation of supercoiled pNO1 DNA measured after 30 mins by fluorescence assayMore data for this Ligand-Target Pair
TargetDNA gyrase A/A/B/B tetramer(Escherichia coli (strain K12))
University of Ljubljana
Curated by ChEMBL
University of Ljubljana
Curated by ChEMBL
Affinity DataIC50: 11nMAssay Description:Inhibition of Escherichia coli DNA gyrase assessed as reduction in relaxation of supercoiled pNO1 DNA measured after 30 mins by fluorescence assayMore data for this Ligand-Target Pair
TargetDNA gyrase A/A/B/B tetramer(Escherichia coli (strain K12))
University of Ljubljana
Curated by ChEMBL
University of Ljubljana
Curated by ChEMBL
Affinity DataIC50: 12nMAssay Description:Inhibition of Escherichia coli DNA gyrase assessed as reduction in relaxation of supercoiled pNO1 DNA measured after 30 mins by fluorescence assayMore data for this Ligand-Target Pair
TargetDNA gyrase A/A/B/B tetramer(Escherichia coli (strain K12))
University of Ljubljana
Curated by ChEMBL
University of Ljubljana
Curated by ChEMBL
Affinity DataIC50: 13nMAssay Description:Inhibition of Escherichia coli DNA gyrase assessed as reduction in relaxation of supercoiled pNO1 DNA measured after 30 mins by fluorescence assayMore data for this Ligand-Target Pair
TargetDNA gyrase A/A/B/B tetramer(Escherichia coli (strain K12))
University of Ljubljana
Curated by ChEMBL
University of Ljubljana
Curated by ChEMBL
Affinity DataIC50: 16nMAssay Description:Inhibition of Escherichia coli DNA gyrase assessed as reduction in relaxation of supercoiled pNO1 DNA measured after 30 mins by fluorescence assayMore data for this Ligand-Target Pair
Affinity DataIC50: 16nMAssay Description:Inhibition of Staphylococcus aureus DNA gyrase assessed reduction in relaxation of supercoiled pNO1 DNA measured after 30 mins by fluorescence assayMore data for this Ligand-Target Pair
Affinity DataIC50: 17nMAssay Description:Inhibition of Staphylococcus aureus DNA gyrase assessed reduction in relaxation of supercoiled pNO1 DNA measured after 30 mins by fluorescence assayMore data for this Ligand-Target Pair
Affinity DataIC50: 17nMAssay Description:Inhibition of Staphylococcus aureus DNA gyrase assessed reduction in relaxation of supercoiled pNO1 DNA measured after 30 mins by fluorescence assayMore data for this Ligand-Target Pair
TargetDNA gyrase A/A/B/B tetramer(Escherichia coli (strain K12))
University of Ljubljana
Curated by ChEMBL
University of Ljubljana
Curated by ChEMBL
Affinity DataIC50: 19nMAssay Description:Inhibition of Escherichia coli DNA gyrase assessed as reduction in relaxation of supercoiled pNO1 DNA measured after 30 mins by fluorescence assayMore data for this Ligand-Target Pair
Affinity DataIC50: 22nMAssay Description:Inhibition of Staphylococcus aureus DNA gyrase assessed reduction in relaxation of supercoiled pNO1 DNA measured after 30 mins by fluorescence assayMore data for this Ligand-Target Pair
TargetDNA gyrase A/A/B/B tetramer(Escherichia coli (strain K12))
University of Ljubljana
Curated by ChEMBL
University of Ljubljana
Curated by ChEMBL
Affinity DataIC50: 24nMAssay Description:Inhibition of Escherichia coli DNA gyrase assessed as reduction in relaxation of supercoiled pNO1 DNA measured after 30 mins by fluorescence assayMore data for this Ligand-Target Pair
Affinity DataIC50: 24nMAssay Description:Inhibition of Staphylococcus aureus DNA gyrase assessed reduction in relaxation of supercoiled pNO1 DNA measured after 30 mins by fluorescence assayMore data for this Ligand-Target Pair
Affinity DataIC50: 25nMAssay Description:Inhibition of Staphylococcus aureus DNA gyrase assessed reduction in relaxation of supercoiled pNO1 DNA measured after 30 mins by fluorescence assayMore data for this Ligand-Target Pair
Affinity DataIC50: 26nMAssay Description:Inhibition of Staphylococcus aureus DNA gyrase assessed reduction in relaxation of supercoiled pNO1 DNA measured after 30 mins by fluorescence assayMore data for this Ligand-Target Pair
Affinity DataIC50: 32nMAssay Description:Inhibition of Staphylococcus aureus DNA gyrase assessed reduction in relaxation of supercoiled pNO1 DNA measured after 30 mins by fluorescence assayMore data for this Ligand-Target Pair
Affinity DataIC50: 34nMAssay Description:Inhibition of Staphylococcus aureus DNA gyrase assessed reduction in relaxation of supercoiled pNO1 DNA measured after 30 mins by fluorescence assayMore data for this Ligand-Target Pair
Affinity DataIC50: 44nMAssay Description:Inhibition of Staphylococcus aureus DNA gyrase assessed reduction in relaxation of supercoiled pNO1 DNA measured after 30 mins by fluorescence assayMore data for this Ligand-Target Pair
TargetDNA gyrase A/A/B/B tetramer(Escherichia coli (strain K12))
University of Ljubljana
Curated by ChEMBL
University of Ljubljana
Curated by ChEMBL
Affinity DataIC50: 46nMAssay Description:Inhibition of Escherichia coli DNA gyrase assessed as reduction in relaxation of supercoiled pNO1 DNA measured after 30 mins by fluorescence assayMore data for this Ligand-Target Pair
Affinity DataIC50: 48nMAssay Description:Inhibition of Staphylococcus aureus DNA gyrase assessed reduction in relaxation of supercoiled pNO1 DNA measured after 30 mins by fluorescence assayMore data for this Ligand-Target Pair
Affinity DataIC50: 59nMAssay Description:Inhibition of Staphylococcus aureus DNA gyrase assessed reduction in relaxation of supercoiled pNO1 DNA measured after 30 mins by fluorescence assayMore data for this Ligand-Target Pair
Affinity DataIC50: 61nMAssay Description:Inhibition of Staphylococcus aureus DNA gyrase assessed reduction in relaxation of supercoiled pNO1 DNA measured after 30 mins by fluorescence assayMore data for this Ligand-Target Pair
Affinity DataIC50: 71nMAssay Description:Inhibition of Staphylococcus aureus DNA gyrase assessed reduction in relaxation of supercoiled pNO1 DNA measured after 30 mins by fluorescence assayMore data for this Ligand-Target Pair
Affinity DataIC50: 72nMAssay Description:Inhibition of Staphylococcus aureus DNA gyrase assessed reduction in relaxation of supercoiled pNO1 DNA measured after 30 mins by fluorescence assayMore data for this Ligand-Target Pair
TargetDNA gyrase A/A/B/B tetramer(Escherichia coli (strain K12))
University of Ljubljana
Curated by ChEMBL
University of Ljubljana
Curated by ChEMBL
Affinity DataIC50: 89nMAssay Description:Inhibition of Escherichia coli DNA gyrase assessed as reduction in relaxation of supercoiled pNO1 DNA measured after 30 mins by fluorescence assayMore data for this Ligand-Target Pair
Affinity DataIC50: 100nMAssay Description:Inhibition of Staphylococcus aureus DNA gyrase assessed reduction in relaxation of supercoiled pNO1 DNA measured after 30 mins by fluorescence assayMore data for this Ligand-Target Pair
Affinity DataIC50: 170nMAssay Description:Inhibition of Staphylococcus aureus DNA gyrase assessed reduction in relaxation of supercoiled pNO1 DNA measured after 30 mins by fluorescence assayMore data for this Ligand-Target Pair
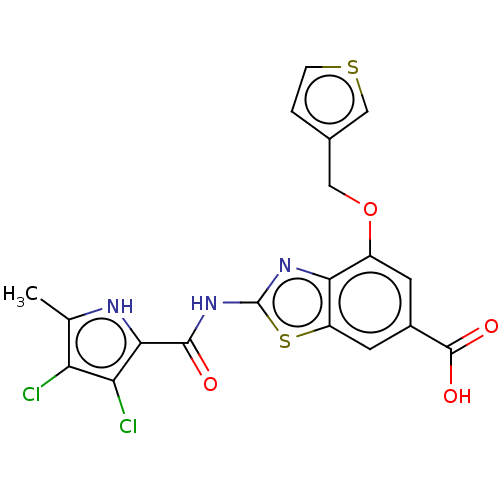
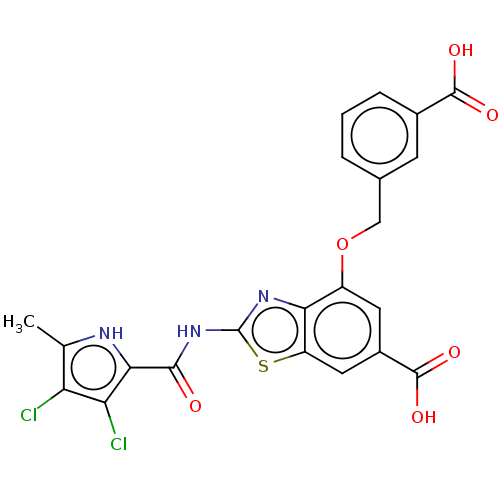
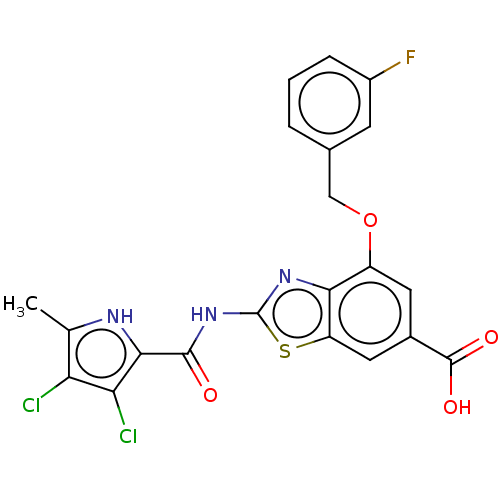
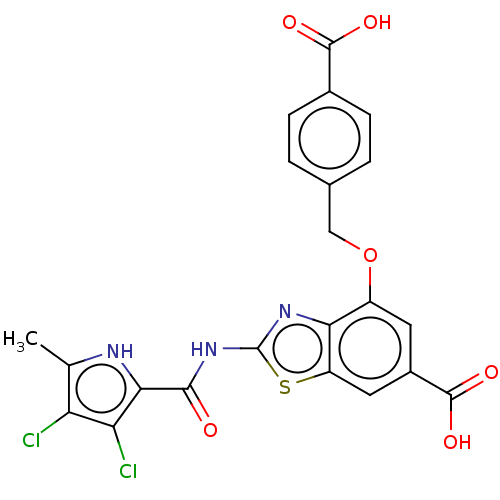
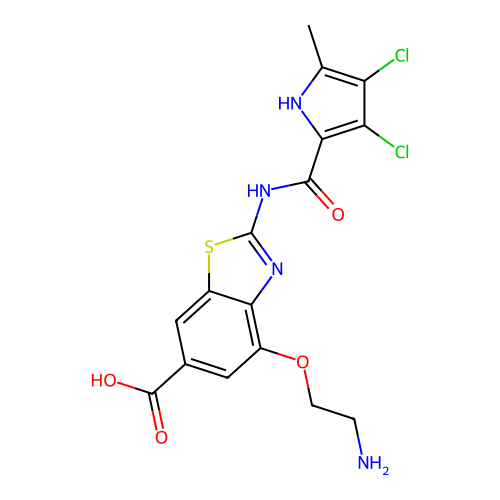
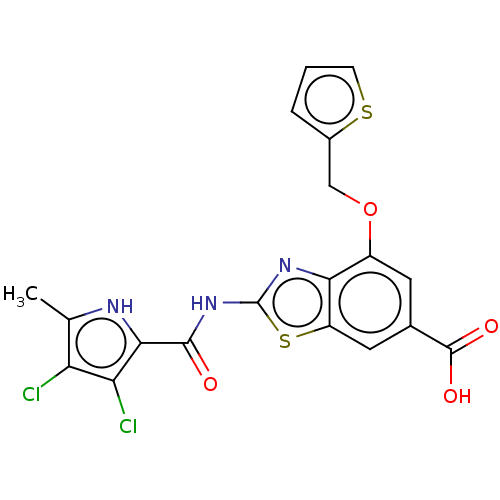
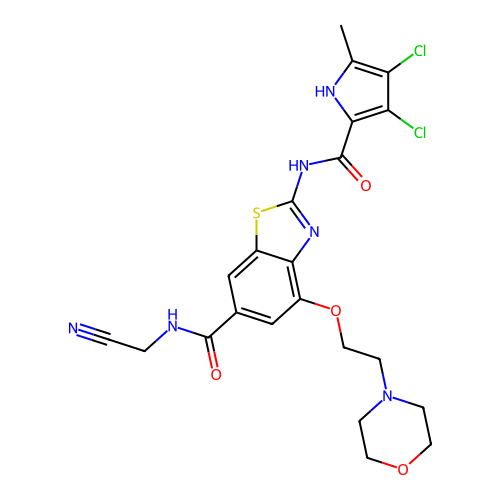
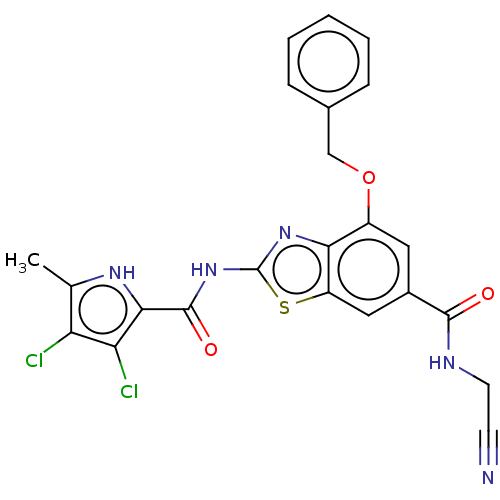
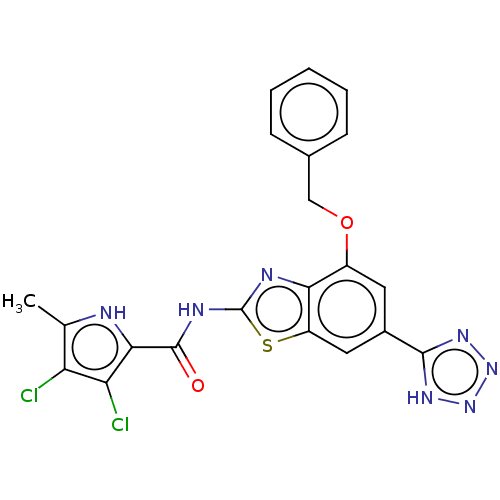
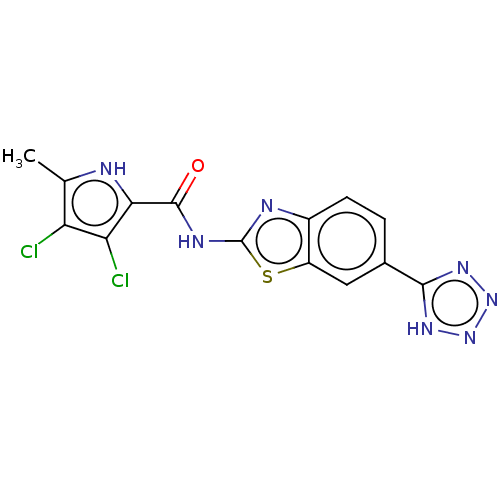
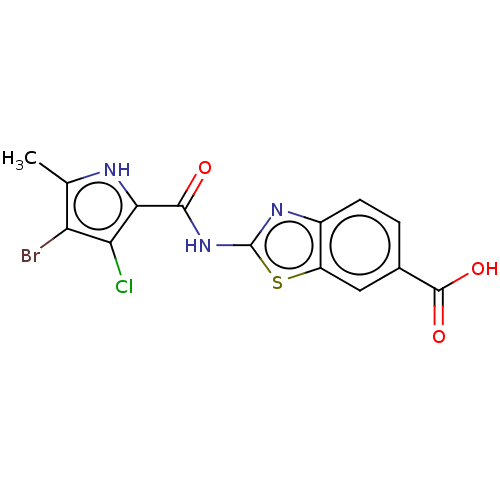
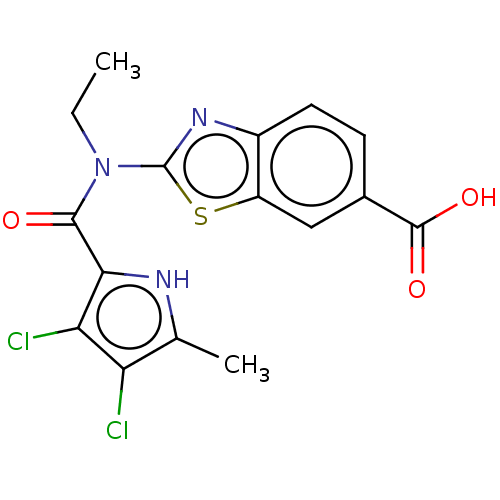
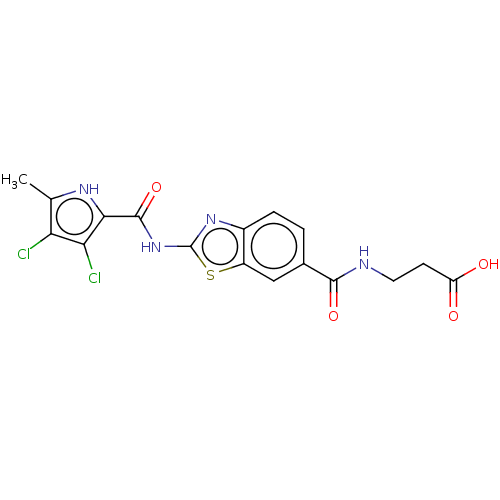
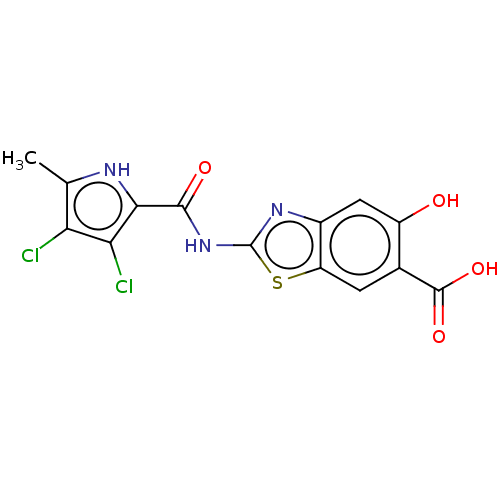
 3D Structure (crystal)
3D Structure (crystal)