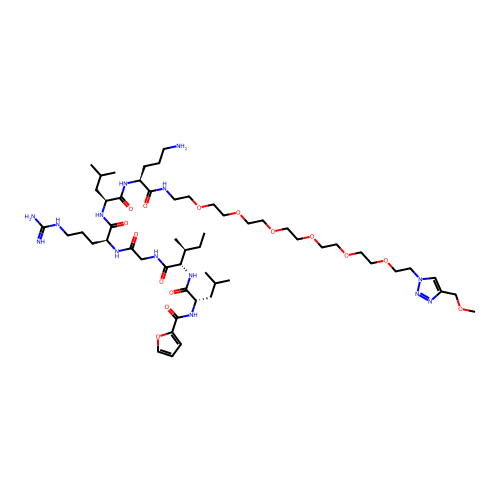
Report error Found 16 Enz. Inhib. hit(s) with all data for entry = 50012010
Affinity DataEC50: 62nMAssay Description:Agonist activity at PAR2 in human EA.hy926 cells assessed as stimulation of calcium mobilization by Fluo-4-AM dye-based FLIPR assayMore data for this Ligand-Target Pair
Affinity DataEC50: 73nMAssay Description:Agonist activity at PAR2 in human EA.hy926 cells assessed as stimulation of calcium mobilization by Fluo-4-AM dye-based FLIPR assayMore data for this Ligand-Target Pair
Affinity DataEC50: 84nMAssay Description:Agonist activity at PAR2 in human EA.hy926 cells assessed as stimulation of calcium mobilization by Fluo-4-AM dye-based FLIPR assayMore data for this Ligand-Target Pair
Affinity DataEC50: 90nMAssay Description:Agonist activity at PAR2 in HUVEC cells assessed as stimulation of calcium mobilization by Fluo-4-AM dye-based FLIPR assayMore data for this Ligand-Target Pair
Affinity DataEC50: 130nMAssay Description:Agonist activity at PAR2 in human EA.hy926 cells assessed as stimulation of calcium mobilization by Fluo-4-AM dye-based FLIPR assayMore data for this Ligand-Target Pair
Affinity DataEC50: 270nMAssay Description:Agonist activity at PAR2 in human EA.hy926 cells assessed as stimulation of calcium mobilization by Fluo-4-AM dye-based FLIPR assayMore data for this Ligand-Target Pair
Affinity DataEC50: 350nMAssay Description:Agonist activity at PAR2 in human EA.hy926 cells assessed as stimulation of calcium mobilization by Fluo-4-AM dye-based FLIPR assayMore data for this Ligand-Target Pair
Affinity DataEC50: 560nMAssay Description:Agonist activity at PAR2 in human EA.hy926 cells assessed as stimulation of calcium mobilization by Fluo-4-AM dye-based FLIPR assayMore data for this Ligand-Target Pair
Affinity DataEC50: 880nMAssay Description:Agonist activity at PAR2 in human EA.hy926 cells assessed as stimulation of calcium mobilization by Fluo-4-AM dye-based FLIPR assayMore data for this Ligand-Target Pair
Affinity DataEC50: 890nMAssay Description:Agonist activity at PAR2 in human EA.hy926 cells assessed as stimulation of calcium mobilization by Fluo-4-AM dye-based FLIPR assayMore data for this Ligand-Target Pair
Affinity DataEC50: 1.00E+3nMAssay Description:Agonist activity at PAR2 in HUVEC cells assessed as stimulation of calcium mobilization by Fluo-4-AM dye-based FLIPR assayMore data for this Ligand-Target Pair
Affinity DataEC50: 1.20E+3nMAssay Description:Agonist activity at PAR2 in human EA.hy926 cells assessed as stimulation of calcium mobilization treated with cRGD for 30 mins prior to compound addi...More data for this Ligand-Target Pair
Affinity DataEC50: 1.20E+3nMAssay Description:Agonist activity at PAR2 in human EA.hy926 cells assessed as stimulation of calcium mobilization preincubated for 2 hrs with LPS and 30 mins with cRG...More data for this Ligand-Target Pair
Affinity DataEC50: 1.20E+3nMAssay Description:Agonist activity at PAR2 in human EA.hy926 cells assessed as stimulation of calcium mobilization by Fluo-4-AM dye-based FLIPR assayMore data for this Ligand-Target Pair
Affinity DataEC50: 2.50E+3nMAssay Description:Agonist activity at PAR2 in human EA.hy926 cells assessed as stimulation of calcium mobilization by Fluo-4-AM dye-based FLIPR assayMore data for this Ligand-Target Pair
Affinity DataEC50: 3.50E+3nMAssay Description:Agonist activity at PAR2 in human EA.hy926 cells assessed as stimulation of calcium mobilization preincubated for 2 hrs with LPS followed by compound...More data for this Ligand-Target Pair