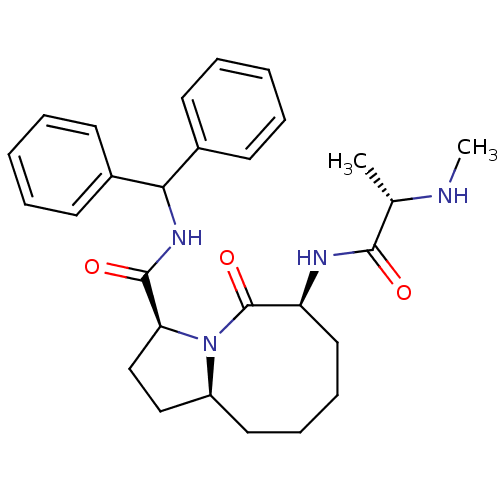
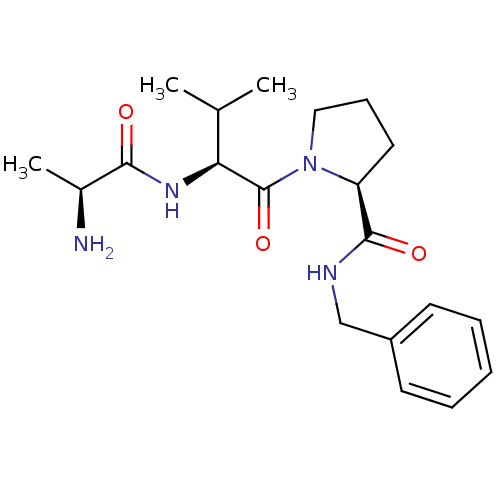
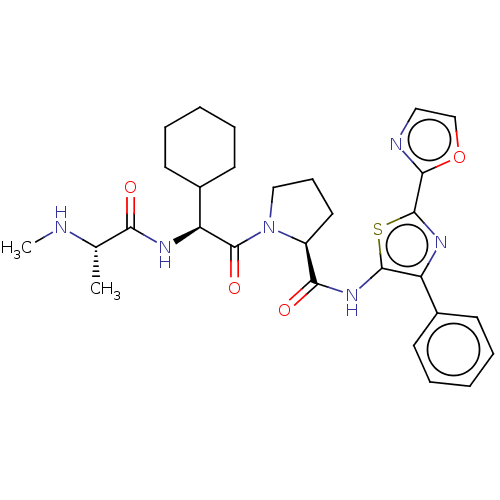
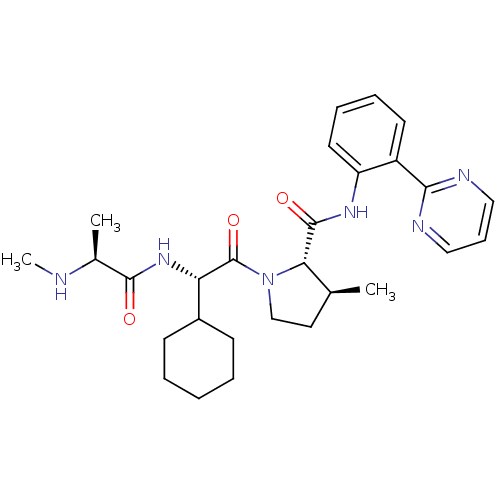
Report error Found 21 Enz. Inhib. hit(s) with all data for entry = 50010800
Affinity DataIC50: 100nMAssay Description:Inhibition of Zika virus NS2B-NS3 proteaseMore data for this Ligand-Target Pair
Affinity DataKi: 140nMAssay Description:Inhibition of Zika virus NS2B-NS3 proteaseMore data for this Ligand-Target Pair
Affinity DataIC50: 300nMAssay Description:Inhibition of Zika virus NS2B-NS3 proteaseMore data for this Ligand-Target Pair
Affinity DataKi: 430nMAssay Description:Inhibition of Zika virus NS2B-NS3 proteaseMore data for this Ligand-Target Pair
Affinity DataKi: 800nMAssay Description:Inhibition of Zika virus NS2B-NS3 proteaseMore data for this Ligand-Target Pair
Affinity DataIC50: 820nMAssay Description:Allosteric inhibition of Zika virus NS2B-NS3 proteaseMore data for this Ligand-Target Pair
Affinity DataIC50: 1.10E+3nMAssay Description:Inhibition of Zika virus NS2B-NS3 proteaseMore data for this Ligand-Target Pair
Affinity DataIC50: 1.70E+3nMAssay Description:Inhibition of Zika virus NS2B-NS3 proteaseMore data for this Ligand-Target Pair
Affinity DataIC50: 1.90E+3nMAssay Description:Inhibition of Zika virus NS2B-NS3 proteaseMore data for this Ligand-Target Pair
Affinity DataIC50: 1.90E+3nMAssay Description:Inhibition of West Nile virus NS2B-NS3 proteaseMore data for this Ligand-Target Pair
Affinity DataEC50: 1.93E+3nMAssay Description:Binding affinity to Zika virus NS2B-NS3 proteaseMore data for this Ligand-Target Pair
Affinity DataKd: 2.00E+3nMAssay Description:Binding affinity to West Nile virus NS2B-NS3 proteaseMore data for this Ligand-Target Pair
Affinity DataKd: 2.00E+3nMAssay Description:Binding affinity to Zika virus NS2B-NS3 proteaseMore data for this Ligand-Target Pair
Affinity DataIC50: 4.10E+3nMAssay Description:Inhibition of N-terminal His-tagged Zika virus NS2B-NS3 protease (1 to 187 residues) using Boc-Gly-Arg-Arg-AMC as substrateMore data for this Ligand-Target Pair
Affinity DataKd: 9.00E+3nMAssay Description:Binding affinity to N-terminal His-tagged Zika virus NS2B-NS3 protease (1 to 187 residues) using Boc-Gly-Arg-Arg-AMC as substrateMore data for this Ligand-Target Pair
Affinity DataKi: 1.35E+4nMAssay Description:Inhibition of N-terminal His-tagged Zika virus NS2B-NS3 protease (1 to 187 residues) using Boc-Gly-Arg-Arg-AMC as substrateMore data for this Ligand-Target Pair
Affinity DataIC50: 1.42E+4nMAssay Description:Inhibition of Zika virus NS2B-NS3 proteaseMore data for this Ligand-Target Pair
Affinity DataIC50: 2.16E+4nMAssay Description:Inhibition of Zika virus NS2B-NS3 proteaseMore data for this Ligand-Target Pair
Affinity DataIC50: 4.70E+4nMAssay Description:Inhibition of Zika virus NS2B-NS3 proteaseMore data for this Ligand-Target Pair
Affinity DataKi: 9.23E+4nMAssay Description:Inhibition of Zika virus NS2B-NS3 proteaseMore data for this Ligand-Target Pair
Affinity DataIC50: 1.41E+7nMAssay Description:Inhibition of Zika virus NS2B-NS3 proteaseMore data for this Ligand-Target Pair