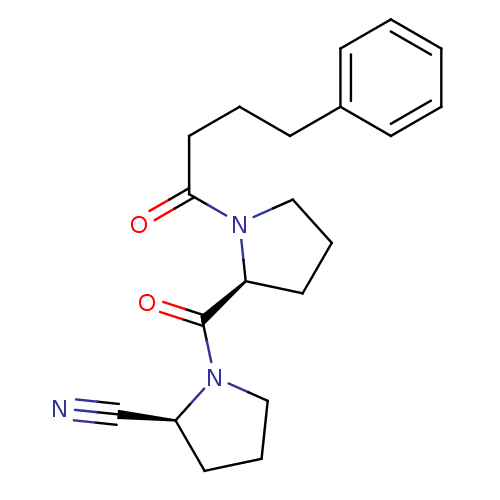
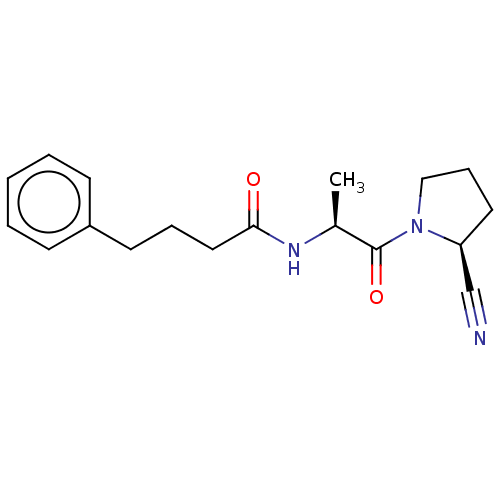
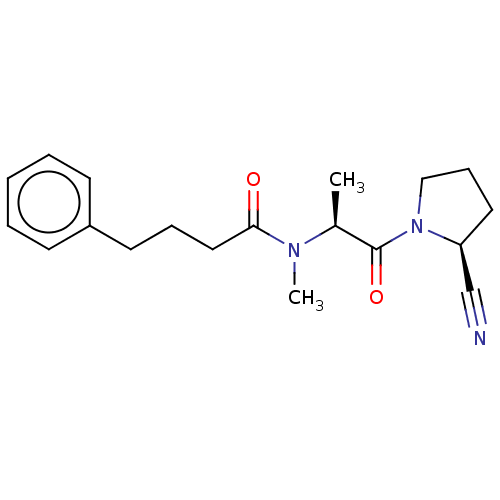
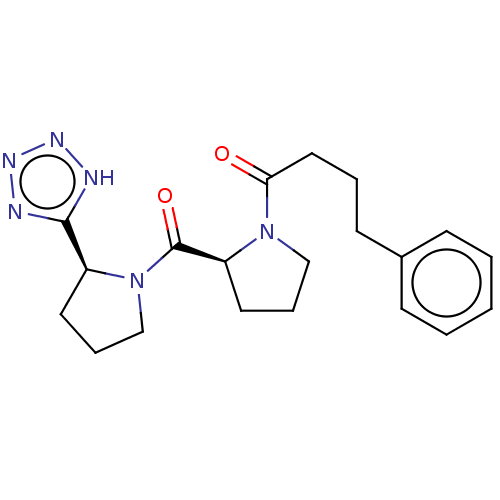
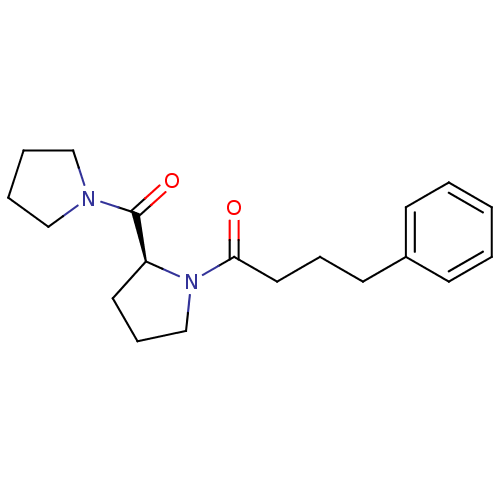
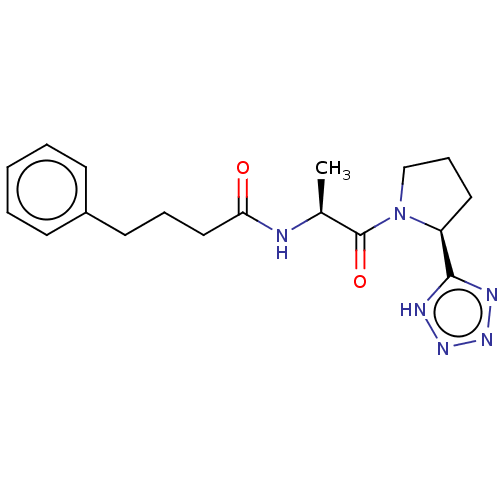
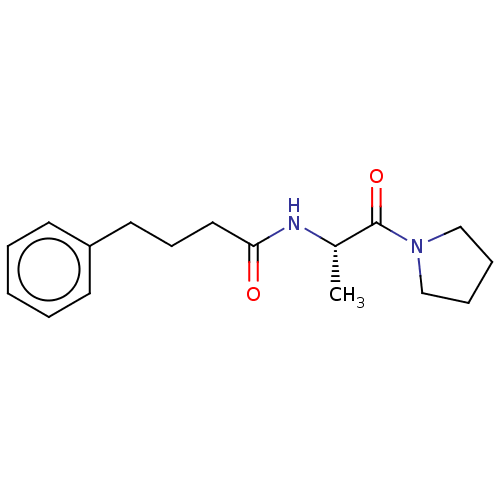
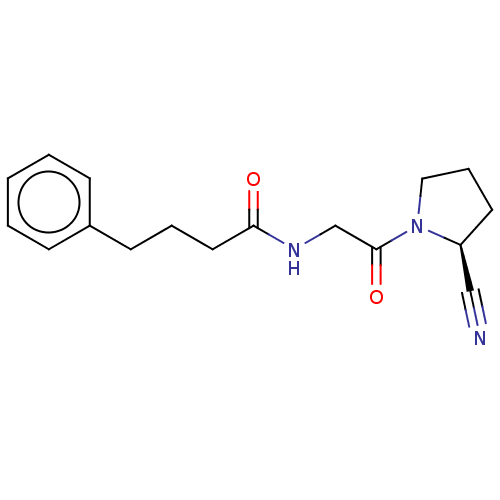
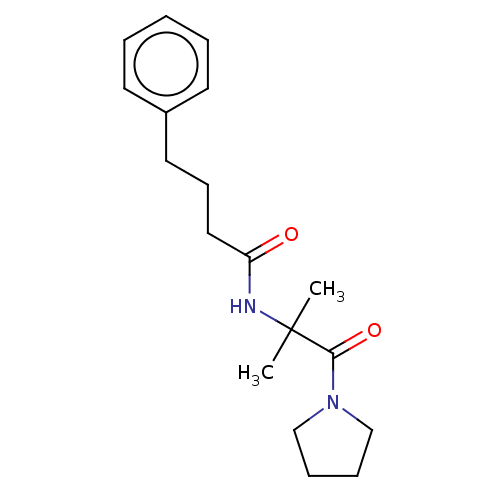
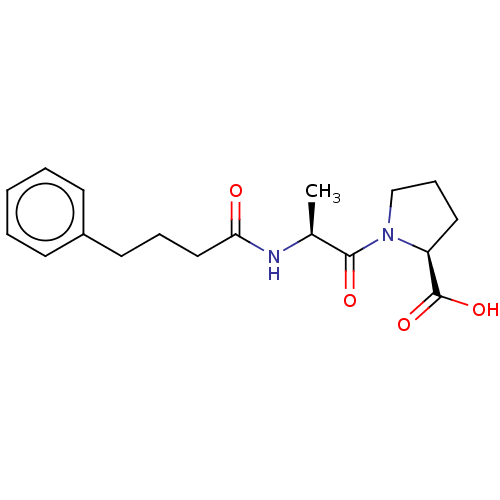
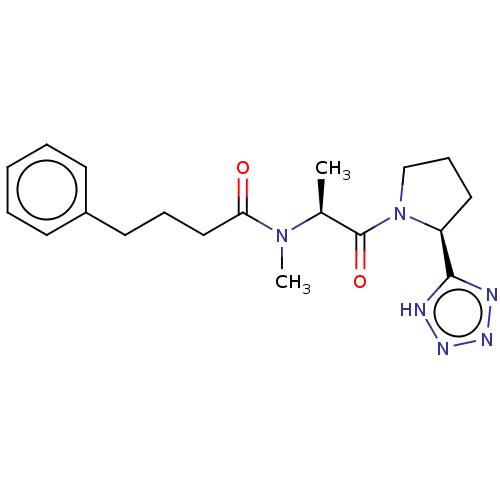
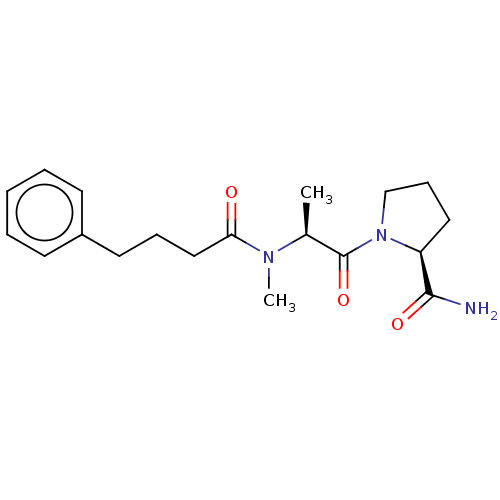
Report error Found 28 Enz. Inhib. hit(s) with all data for entry = 50009512
Affinity DataIC50: 0.300nMAssay Description:Inhibition of PREP in C57BL/6 mouse cortex homogenate assessed as formation of 7-amido-4-methylcoumarin using Suc-Gly-Pro-amido-4-methylcoumarin as s...More data for this Ligand-Target Pair
Affinity DataIC50: 0.860nMAssay Description:Inhibition of recombinant porcine brain PREP expressed in Escherichia coli assessed as formation of 7-amido-4-methylcoumarin using Suc-Gly-Pro-amido-...More data for this Ligand-Target Pair
Affinity DataIC50: 3.30nMAssay Description:Inhibition of PREP in C57BL/6 mouse cortex homogenate assessed as formation of 7-amido-4-methylcoumarin using Suc-Gly-Pro-amido-4-methylcoumarin as s...More data for this Ligand-Target Pair
Affinity DataIC50: 4.10nMAssay Description:Inhibition of recombinant porcine brain PREP expressed in Escherichia coli assessed as formation of 7-amido-4-methylcoumarin using Suc-Gly-Pro-amido-...More data for this Ligand-Target Pair
Affinity DataIC50: 5.40nMAssay Description:Inhibition of recombinant porcine brain PREP expressed in Escherichia coli assessed as formation of 7-amido-4-methylcoumarin using Suc-Gly-Pro-amido-...More data for this Ligand-Target Pair
Affinity DataIC50: 12nMAssay Description:Inhibition of recombinant porcine brain PREP expressed in Escherichia coli assessed as formation of 7-amido-4-methylcoumarin using Suc-Gly-Pro-amido-...More data for this Ligand-Target Pair
Affinity DataIC50: 12nMAssay Description:Inhibition of recombinant porcine brain PREP expressed in Escherichia coli assessed as formation of 7-amido-4-methylcoumarin using Suc-Gly-Pro-amido-...More data for this Ligand-Target Pair
Affinity DataIC50: 91nMAssay Description:Inhibition of PREP in C57BL/6 mouse cortex homogenate assessed as formation of 7-amido-4-methylcoumarin using Suc-Gly-Pro-amido-4-methylcoumarin as s...More data for this Ligand-Target Pair
Affinity DataIC50: 129nMAssay Description:Inhibition of recombinant porcine brain PREP expressed in Escherichia coli assessed as formation of 7-amido-4-methylcoumarin using Suc-Gly-Pro-amido-...More data for this Ligand-Target Pair
Affinity DataIC50: 147nMAssay Description:Inhibition of recombinant porcine brain PREP expressed in Escherichia coli assessed as formation of 7-amido-4-methylcoumarin using Suc-Gly-Pro-amido-...More data for this Ligand-Target Pair
Affinity DataIC50: 220nMAssay Description:Inhibition of recombinant porcine brain PREP expressed in Escherichia coli assessed as formation of 7-amido-4-methylcoumarin using Suc-Gly-Pro-amido-...More data for this Ligand-Target Pair
Affinity DataIC50: 264nMAssay Description:Inhibition of PREP in C57BL/6 mouse cortex homogenate assessed as formation of 7-amido-4-methylcoumarin using Suc-Gly-Pro-amido-4-methylcoumarin as s...More data for this Ligand-Target Pair
Affinity DataIC50: 269nMAssay Description:Inhibition of recombinant porcine brain PREP expressed in Escherichia coli assessed as formation of 7-amido-4-methylcoumarin using Suc-Gly-Pro-amido-...More data for this Ligand-Target Pair
Affinity DataIC50: 298nMAssay Description:Inhibition of PREP in C57BL/6 mouse cortex homogenate assessed as formation of 7-amido-4-methylcoumarin using Suc-Gly-Pro-amido-4-methylcoumarin as s...More data for this Ligand-Target Pair
Affinity DataIC50: 3.63E+3nMAssay Description:Inhibition of recombinant porcine brain PREP expressed in Escherichia coli assessed as formation of 7-amido-4-methylcoumarin using Suc-Gly-Pro-amido-...More data for this Ligand-Target Pair
Affinity DataIC50: 4.37E+3nMAssay Description:Inhibition of recombinant porcine brain PREP expressed in Escherichia coli assessed as formation of 7-amido-4-methylcoumarin using Suc-Gly-Pro-amido-...More data for this Ligand-Target Pair
Affinity DataIC50: 5.14E+3nMAssay Description:Inhibition of PREP in C57BL/6 mouse cortex homogenate assessed as formation of 7-amido-4-methylcoumarin using Suc-Gly-Pro-amido-4-methylcoumarin as s...More data for this Ligand-Target Pair
Affinity DataIC50: 6.21E+3nMAssay Description:Inhibition of PREP in C57BL/6 mouse cortex homogenate assessed as formation of 7-amido-4-methylcoumarin using Suc-Gly-Pro-amido-4-methylcoumarin as s...More data for this Ligand-Target Pair
Affinity DataIC50: 1.06E+4nMAssay Description:Inhibition of recombinant porcine brain PREP expressed in Escherichia coli assessed as formation of 7-amido-4-methylcoumarin using Suc-Gly-Pro-amido-...More data for this Ligand-Target Pair
Affinity DataIC50: 1.21E+4nMAssay Description:Inhibition of PREP in C57BL/6 mouse cortex homogenate assessed as formation of 7-amido-4-methylcoumarin using Suc-Gly-Pro-amido-4-methylcoumarin as s...More data for this Ligand-Target Pair
Affinity DataIC50: 1.35E+4nMAssay Description:Inhibition of PREP in C57BL/6 mouse cortex homogenate assessed as formation of 7-amido-4-methylcoumarin using Suc-Gly-Pro-amido-4-methylcoumarin as s...More data for this Ligand-Target Pair
Affinity DataIC50: 1.75E+4nMAssay Description:Inhibition of recombinant porcine brain PREP expressed in Escherichia coli assessed as formation of 7-amido-4-methylcoumarin using Suc-Gly-Pro-amido-...More data for this Ligand-Target Pair
Affinity DataIC50: 2.72E+4nMAssay Description:Inhibition of recombinant porcine brain PREP expressed in Escherichia coli assessed as formation of 7-amido-4-methylcoumarin using Suc-Gly-Pro-amido-...More data for this Ligand-Target Pair
Affinity DataIC50: 2.88E+4nMAssay Description:Inhibition of PREP in C57BL/6 mouse cortex homogenate assessed as formation of 7-amido-4-methylcoumarin using Suc-Gly-Pro-amido-4-methylcoumarin as s...More data for this Ligand-Target Pair
Affinity DataIC50: 3.52E+4nMAssay Description:Inhibition of PREP in C57BL/6 mouse cortex homogenate assessed as formation of 7-amido-4-methylcoumarin using Suc-Gly-Pro-amido-4-methylcoumarin as s...More data for this Ligand-Target Pair
Affinity DataIC50: 1.54E+5nMAssay Description:Inhibition of PREP in C57BL/6 mouse cortex homogenate assessed as formation of 7-amido-4-methylcoumarin using Suc-Gly-Pro-amido-4-methylcoumarin as s...More data for this Ligand-Target Pair
Affinity DataIC50: 2.05E+5nMAssay Description:Inhibition of recombinant porcine brain PREP expressed in Escherichia coli assessed as formation of 7-amido-4-methylcoumarin using Suc-Gly-Pro-amido-...More data for this Ligand-Target Pair
Affinity DataIC50: 4.57E+5nMAssay Description:Inhibition of PREP in C57BL/6 mouse cortex homogenate assessed as formation of 7-amido-4-methylcoumarin using Suc-Gly-Pro-amido-4-methylcoumarin as s...More data for this Ligand-Target Pair