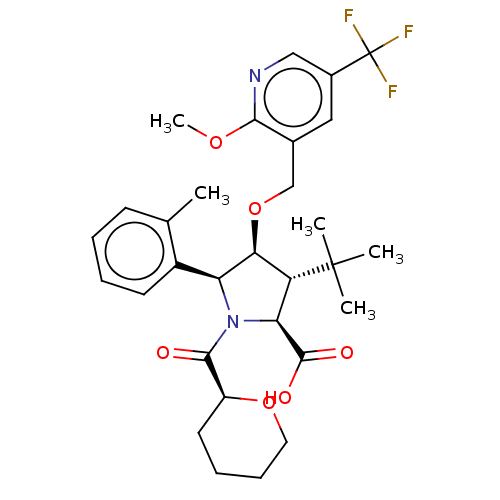
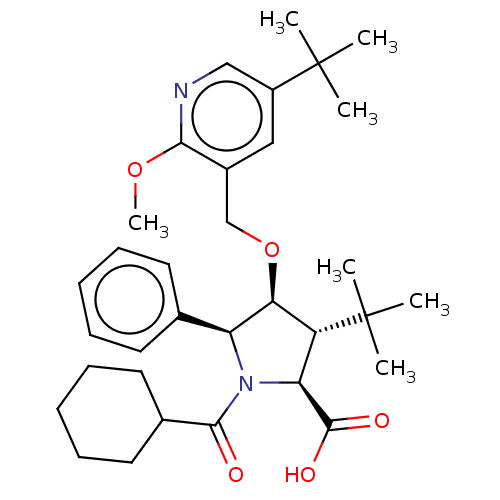
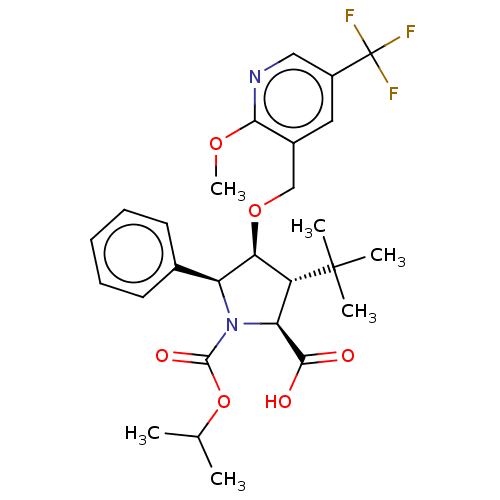
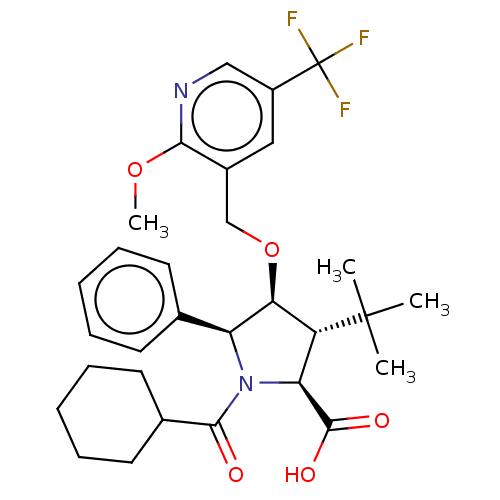
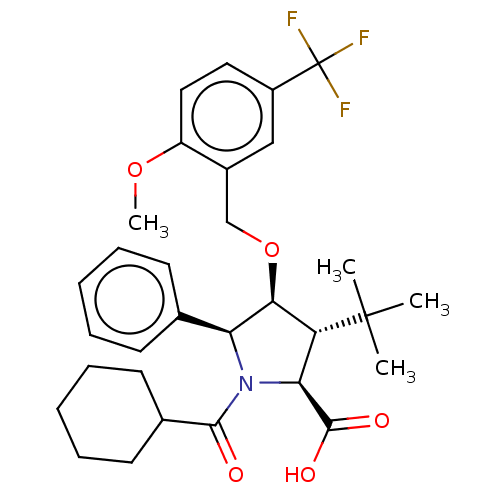
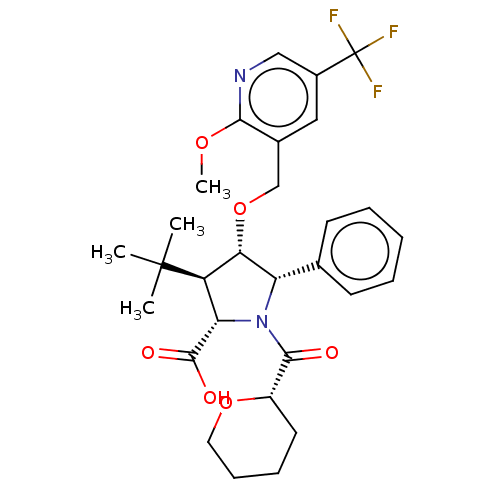
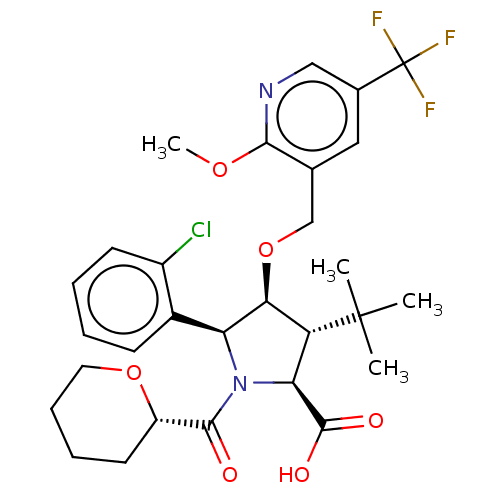
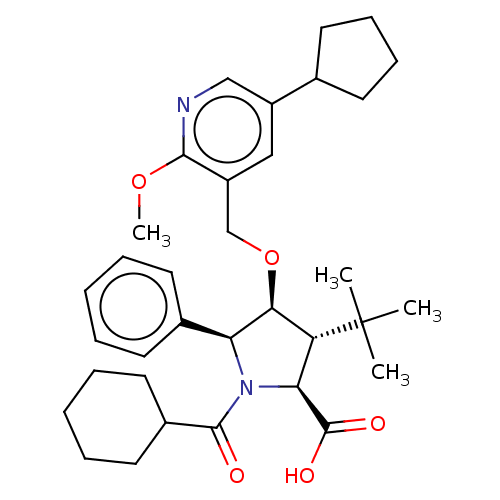
Report error Found 25 Enz. Inhib. hit(s) with all data for entry = 50006946
Affinity DataEC50: 35nMAssay Description:Corrector activity at forskolin-stimulated CFTR F508del mutant in human NHBE cells after 18 to 24 hrs in presence of GLPG1873 by voltage-clamp based ...More data for this Ligand-Target Pair
Affinity DataEC50: 48nMAssay Description:Corrector activity at forskolin-stimulated CFTR F508del mutant in human NHBE cells after 18 to 24 hrs in presence of GLPG1873 by voltage-clamp based ...More data for this Ligand-Target Pair
Affinity DataEC50: 52nMAssay Description:Corrector activity at forskolin-stimulated CFTR F508del mutant in human NHBE cells after 18 to 24 hrs in presence of GLPG1873 by voltage-clamp based ...More data for this Ligand-Target Pair
Affinity DataEC50: 66nMAssay Description:Corrector activity at forskolin-stimulated CFTR F508del mutant in human NHBE cells after 18 to 24 hrs in presence of GLPG1873 by voltage-clamp based ...More data for this Ligand-Target Pair
Affinity DataEC50: 105nMAssay Description:Corrector activity at forskolin-stimulated CFTR F508del mutant in human NHBE cells after 18 to 24 hrs in presence of GLPG1873 by voltage-clamp based ...More data for this Ligand-Target Pair
Affinity DataEC50: 132nMAssay Description:Corrector activity at forskolin-stimulated CFTR F508del mutant in human NHBE cells after 18 to 24 hrs in presence of GLPG1873 by voltage-clamp based ...More data for this Ligand-Target Pair
Affinity DataEC50: 137nMAssay Description:Corrector activity at forskolin-stimulated CFTR F508del mutant in human NHBE cells after 18 to 24 hrs in presence of GLPG1873 by voltage-clamp based ...More data for this Ligand-Target Pair
Affinity DataEC50: 150nMAssay Description:Corrector activity at forskolin-stimulated CFTR F508del mutant in human NHBE cells after 18 to 24 hrs in presence of GLPG1873 by voltage-clamp based ...More data for this Ligand-Target Pair
Affinity DataEC50: 173nMAssay Description:Corrector activity at forskolin-stimulated CFTR F508del mutant in human NHBE cells after 18 to 24 hrs in presence of GLPG1873 by voltage-clamp based ...More data for this Ligand-Target Pair
Affinity DataEC50: 213nMAssay Description:Corrector activity at forskolin-stimulated CFTR F508del mutant in human NHBE cells after 18 to 24 hrs in presence of GLPG1873 by voltage-clamp based ...More data for this Ligand-Target Pair
Affinity DataEC50: 222nMAssay Description:Corrector activity at forskolin-stimulated CFTR F508del mutant in human NHBE cells after 18 to 24 hrs in presence of GLPG1873 by voltage-clamp based ...More data for this Ligand-Target Pair
Affinity DataEC50: 247nMAssay Description:Corrector activity at forskolin-stimulated CFTR F508del mutant in human NHBE cells after 18 to 24 hrs in presence of GLPG1873 by voltage-clamp based ...More data for this Ligand-Target Pair
Affinity DataEC50: 346nMAssay Description:Corrector activity at forskolin-stimulated CFTR F508del mutant in human NHBE cells after 18 to 24 hrs in presence of GLPG1873 by voltage-clamp based ...More data for this Ligand-Target Pair
Affinity DataEC50: 580nMAssay Description:Corrector activity at CFTR F508del mutant (unknown origin) expressed in CFBE41o- cells measured at 18 to 24 hrs in presence of 3-((2R,4S)-4-(1-(2,2-d...More data for this Ligand-Target Pair
Affinity DataEC50: 820nMAssay Description:Corrector activity at CFTR F508del mutant (unknown origin) expressed in CFBE41o- cells measured after 18 to 24 hrs by HRP based luminescence assayMore data for this Ligand-Target Pair
Affinity DataEC50: 1.13E+3nMAssay Description:Corrector activity at CFTR F508del mutant (unknown origin) expressed in CFBE41o- cells measured at 18 to 24 hrs in presence of 3-((2R,4S)-4-(1-(2,2-d...More data for this Ligand-Target Pair
Affinity DataEC50: 1.13E+3nMAssay Description:Corrector activity at CFTR F508del mutant (unknown origin) expressed in CFBE41o- cells measured at 18 to 24 hrs in presence of 3-((2R,4S)-4-(1-(2,2-d...More data for this Ligand-Target Pair
Affinity DataEC50: 1.32E+3nMAssay Description:Corrector activity at CFTR F508del mutant (unknown origin) expressed in CFBE41o- cells measured after 18 to 24 hrs by HRP based luminescence assayMore data for this Ligand-Target Pair
Affinity DataEC50: 1.97E+3nMAssay Description:Corrector activity at CFTR F508del mutant (unknown origin) expressed in CFBE41o- cells measured at 18 to 24 hrs in presence of 3-((2R,4S)-4-(1-(2,2-d...More data for this Ligand-Target Pair
Affinity DataEC50: 2.01E+3nMAssay Description:Corrector activity at CFTR F508del mutant (unknown origin) expressed in CFBE41o- cells measured at 18 to 24 hrs in presence of 3-((2R,4S)-4-(1-(2,2-d...More data for this Ligand-Target Pair
Affinity DataEC50: 2.11E+3nMAssay Description:Corrector activity at CFTR F508del mutant (unknown origin) expressed in CFBE41o- cells measured at 18 to 24 hrs in presence of 3-((2R,4S)-4-(1-(2,2-d...More data for this Ligand-Target Pair
Affinity DataEC50: 2.52E+3nMAssay Description:Corrector activity at CFTR F508del mutant (unknown origin) expressed in CFBE41o- cells measured after 18 to 24 hrs by HRP based luminescence assayMore data for this Ligand-Target Pair
Affinity DataEC50: 2.66E+3nMAssay Description:Corrector activity at CFTR F508del mutant (unknown origin) expressed in CFBE41o- cells measured after 18 to 24 hrs by HRP based luminescence assayMore data for this Ligand-Target Pair
Affinity DataEC50: 2.72E+3nMAssay Description:Corrector activity at CFTR F508del mutant (unknown origin) expressed in CFBE41o- cells measured after 18 to 24 hrs by HRP based luminescence assayMore data for this Ligand-Target Pair
Affinity DataEC50: >2.00E+4nMAssay Description:Corrector activity at CFTR F508del mutant (unknown origin) expressed in CFBE41o- cells measured after 18 to 24 hrs by HRP based luminescence assayMore data for this Ligand-Target Pair