Report error Found 23 Enz. Inhib. hit(s) with all data for entry = 50010333
Target5'-AMP-activated protein kinase catalytic subunit alpha-2/subunit beta-1/subunit gamma-1(Human)
Nanjing University of Chinese Medicine
Curated by ChEMBL
Nanjing University of Chinese Medicine
Curated by ChEMBL
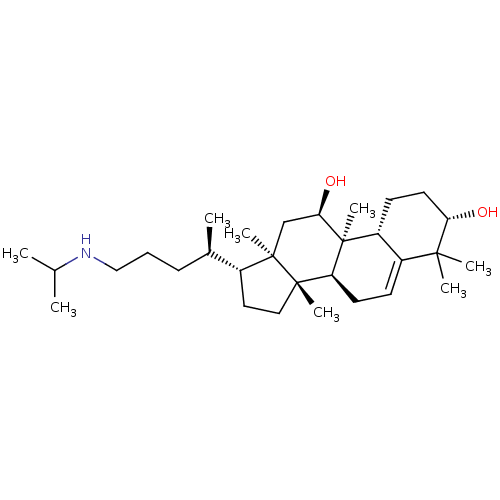
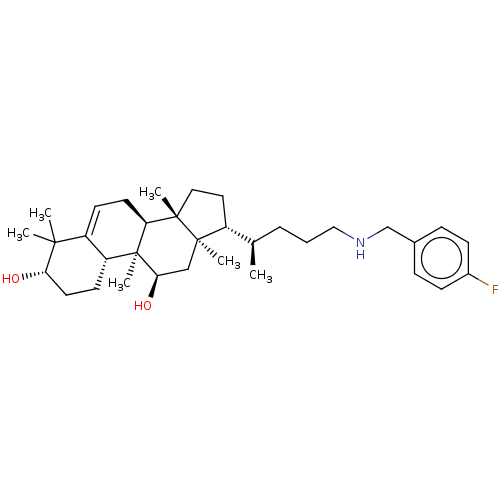
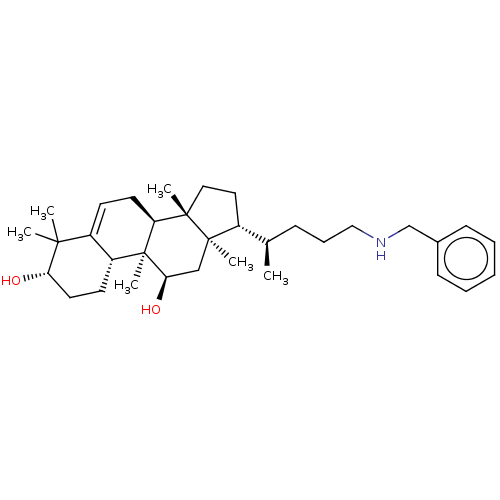
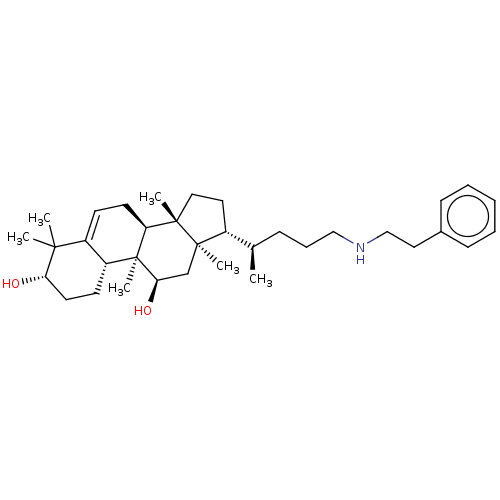
Affinity DataEC50: 140nMAssay Description:Allosteric activation of AMPK alpha2/beta1/gamma1 in human HepG2 cells by scintillation proximity assayMore data for this Ligand-Target Pair
Target5'-AMP-activated protein kinase catalytic subunit alpha-2/subunit beta-1/subunit gamma-1(Human)
Nanjing University of Chinese Medicine
Curated by ChEMBL
Nanjing University of Chinese Medicine
Curated by ChEMBL
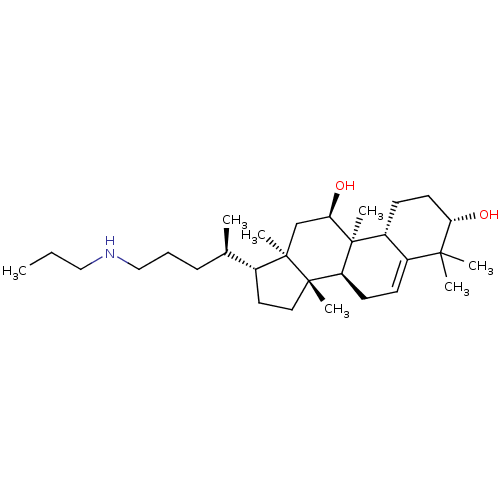
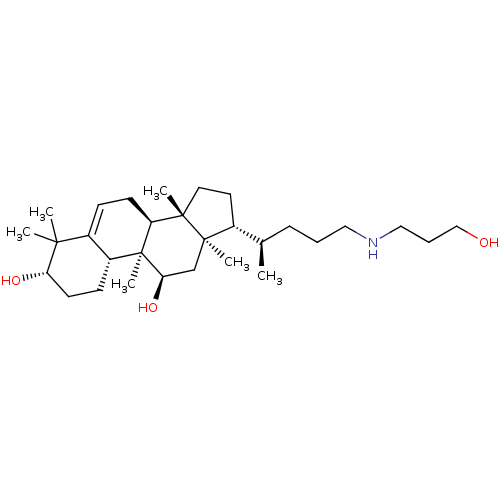
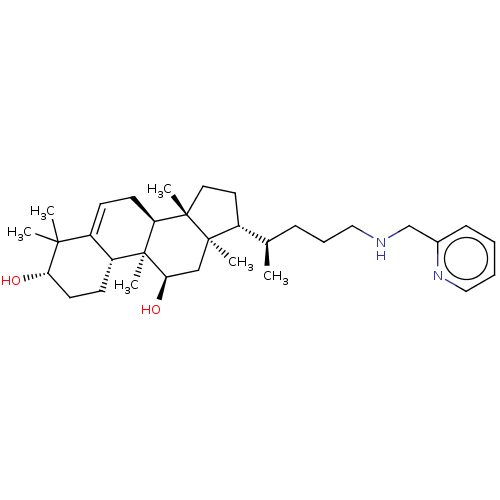
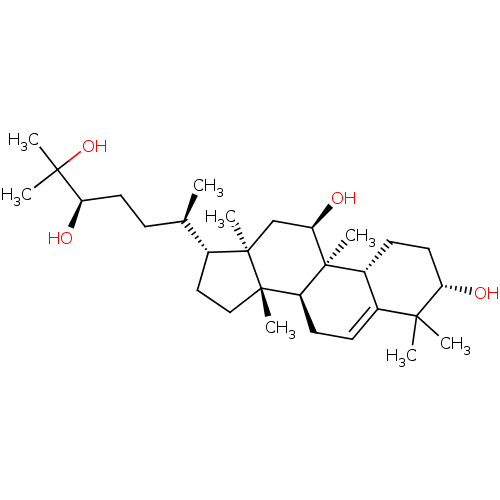
Affinity DataEC50: 150nMAssay Description:Allosteric activation of AMPK alpha2/beta1/gamma1 in human HepG2 cells by scintillation proximity assayMore data for this Ligand-Target Pair
Target5'-AMP-activated protein kinase catalytic subunit alpha-2/subunit beta-1/subunit gamma-1(Human)
Nanjing University of Chinese Medicine
Curated by ChEMBL
Nanjing University of Chinese Medicine
Curated by ChEMBL
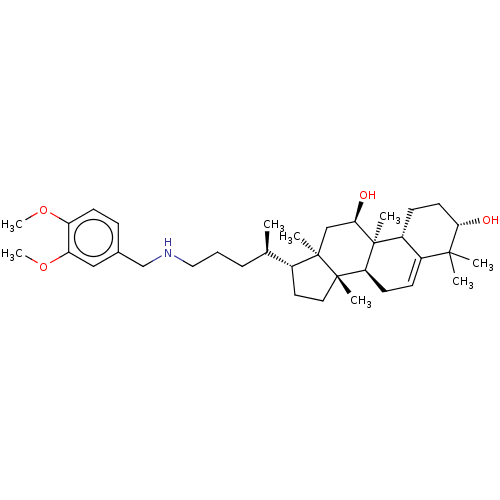
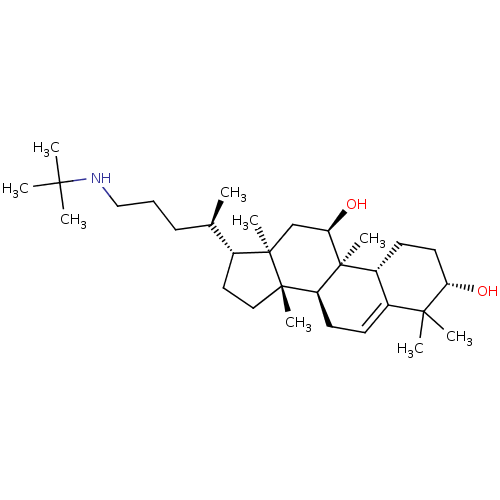
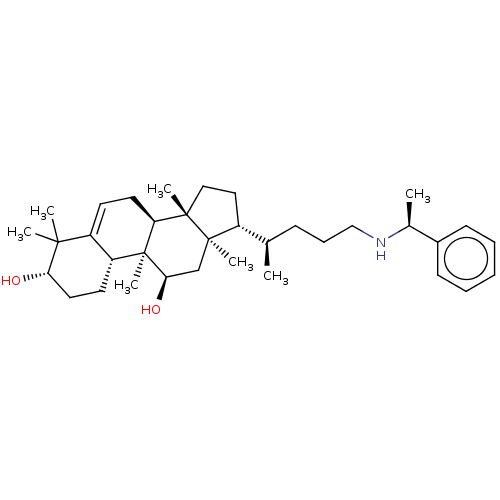
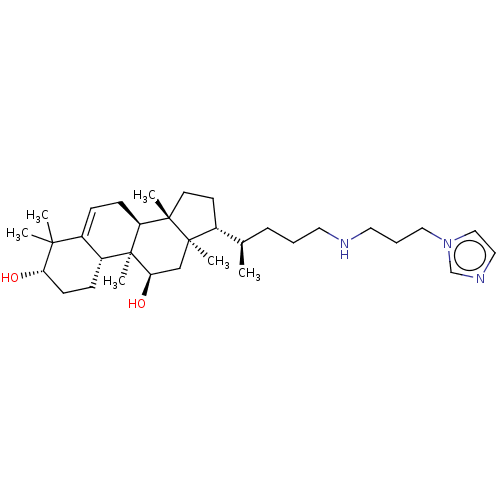
Affinity DataEC50: 200nMAssay Description:Allosteric activation of AMPK alpha2/beta1/gamma1 in human HepG2 cells by scintillation proximity assayMore data for this Ligand-Target Pair
Target5'-AMP-activated protein kinase catalytic subunit alpha-2/subunit beta-1/subunit gamma-1(Human)
Nanjing University of Chinese Medicine
Curated by ChEMBL
Nanjing University of Chinese Medicine
Curated by ChEMBL
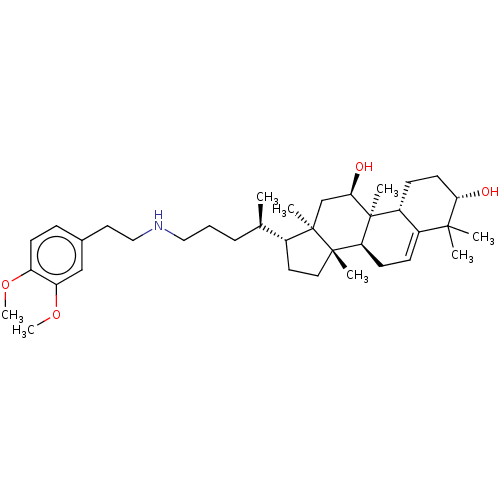
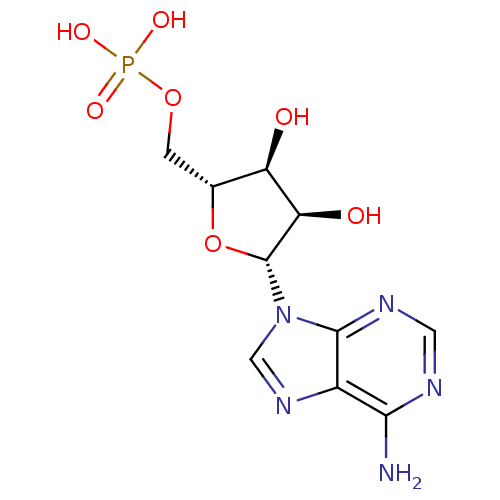
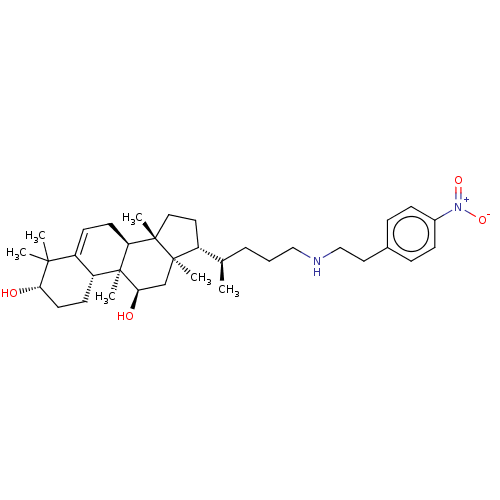
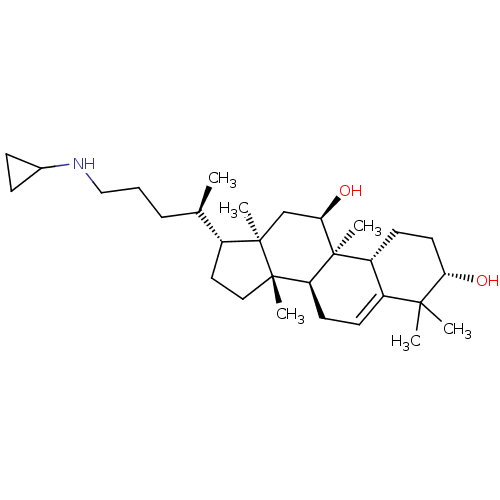
Affinity DataEC50: 500nMAssay Description:Allosteric activation of AMPK alpha2/beta1/gamma1 in human HepG2 cells by scintillation proximity assayMore data for this Ligand-Target Pair
Target5'-AMP-activated protein kinase catalytic subunit alpha-2/subunit beta-1/subunit gamma-1(Human)
Nanjing University of Chinese Medicine
Curated by ChEMBL
Nanjing University of Chinese Medicine
Curated by ChEMBL
Affinity DataEC50: 500nMAssay Description:Allosteric activation of AMPK alpha2/beta1/gamma1 in human HepG2 cells by scintillation proximity assayMore data for this Ligand-Target Pair
Target5'-AMP-activated protein kinase catalytic subunit alpha-2/subunit beta-1/subunit gamma-1(Human)
Nanjing University of Chinese Medicine
Curated by ChEMBL
Nanjing University of Chinese Medicine
Curated by ChEMBL
Affinity DataEC50: 500nMAssay Description:Allosteric activation of AMPK alpha2/beta1/gamma1 in human HepG2 cells by scintillation proximity assayMore data for this Ligand-Target Pair
Target5'-AMP-activated protein kinase catalytic subunit alpha-2/subunit beta-1/subunit gamma-1(Human)
Nanjing University of Chinese Medicine
Curated by ChEMBL
Nanjing University of Chinese Medicine
Curated by ChEMBL
Affinity DataEC50: 600nMAssay Description:Allosteric activation of AMPK alpha2/beta1/gamma1 in human HepG2 cells by scintillation proximity assayMore data for this Ligand-Target Pair
Target5'-AMP-activated protein kinase catalytic subunit alpha-2/subunit beta-1/subunit gamma-1(Human)
Nanjing University of Chinese Medicine
Curated by ChEMBL
Nanjing University of Chinese Medicine
Curated by ChEMBL
Affinity DataEC50: 700nMAssay Description:Allosteric activation of AMPK alpha2/beta1/gamma1 in human HepG2 cells by scintillation proximity assayMore data for this Ligand-Target Pair
Target5'-AMP-activated protein kinase catalytic subunit alpha-2/subunit beta-1/subunit gamma-1(Human)
Nanjing University of Chinese Medicine
Curated by ChEMBL
Nanjing University of Chinese Medicine
Curated by ChEMBL
Affinity DataEC50: 700nMAssay Description:Allosteric activation of AMPK alpha2/beta1/gamma1 in human HepG2 cells by scintillation proximity assayMore data for this Ligand-Target Pair
Target5'-AMP-activated protein kinase catalytic subunit alpha-2/subunit beta-1/subunit gamma-1(Human)
Nanjing University of Chinese Medicine
Curated by ChEMBL
Nanjing University of Chinese Medicine
Curated by ChEMBL
Affinity DataEC50: 1.40E+3nMAssay Description:Allosteric activation of AMPK alpha2/beta1/gamma1 in human HepG2 cells by scintillation proximity assayMore data for this Ligand-Target Pair
Target5'-AMP-activated protein kinase catalytic subunit alpha-2/subunit beta-1/subunit gamma-1(Human)
Nanjing University of Chinese Medicine
Curated by ChEMBL
Nanjing University of Chinese Medicine
Curated by ChEMBL
Affinity DataEC50: 1.40E+3nMAssay Description:Allosteric activation of AMPK alpha2/beta1/gamma1 in human HepG2 cells by scintillation proximity assayMore data for this Ligand-Target Pair
Target5'-AMP-activated protein kinase catalytic subunit alpha-2/subunit beta-1/subunit gamma-1(Human)
Nanjing University of Chinese Medicine
Curated by ChEMBL
Nanjing University of Chinese Medicine
Curated by ChEMBL
Affinity DataEC50: 1.40E+3nMAssay Description:Allosteric activation of AMPK alpha2/beta1/gamma1 in human HepG2 cells by scintillation proximity assayMore data for this Ligand-Target Pair
Target5'-AMP-activated protein kinase catalytic subunit alpha-2/subunit beta-1/subunit gamma-1(Human)
Nanjing University of Chinese Medicine
Curated by ChEMBL
Nanjing University of Chinese Medicine
Curated by ChEMBL
Affinity DataEC50: 1.50E+3nMAssay Description:Allosteric activation of AMPK alpha2/beta1/gamma1 in human HepG2 cells by scintillation proximity assayMore data for this Ligand-Target Pair
Target5'-AMP-activated protein kinase catalytic subunit alpha-2/subunit beta-1/subunit gamma-1(Human)
Nanjing University of Chinese Medicine
Curated by ChEMBL
Nanjing University of Chinese Medicine
Curated by ChEMBL
Affinity DataEC50: 1.60E+3nMAssay Description:Allosteric activation of AMPK alpha2/beta1/gamma1 in human HepG2 cells by scintillation proximity assayMore data for this Ligand-Target Pair
Target5'-AMP-activated protein kinase catalytic subunit alpha-2/subunit beta-1/subunit gamma-1(Human)
Nanjing University of Chinese Medicine
Curated by ChEMBL
Nanjing University of Chinese Medicine
Curated by ChEMBL
Affinity DataEC50: 1.70E+3nMAssay Description:Allosteric activation of AMPK alpha2/beta1/gamma1 in human HepG2 cells by scintillation proximity assayMore data for this Ligand-Target Pair
Target5'-AMP-activated protein kinase catalytic subunit alpha-2/subunit beta-1/subunit gamma-1(Human)
Nanjing University of Chinese Medicine
Curated by ChEMBL
Nanjing University of Chinese Medicine
Curated by ChEMBL
Affinity DataEC50: 1.70E+3nMAssay Description:Allosteric activation of AMPK alpha2/beta1/gamma1 in human HepG2 cells by scintillation proximity assayMore data for this Ligand-Target Pair
Target5'-AMP-activated protein kinase catalytic subunit alpha-2/subunit beta-1/subunit gamma-1(Human)
Nanjing University of Chinese Medicine
Curated by ChEMBL
Nanjing University of Chinese Medicine
Curated by ChEMBL
Affinity DataEC50: 2.00E+3nMAssay Description:Allosteric activation of AMPK alpha2/beta1/gamma1 in human HepG2 cells by scintillation proximity assayMore data for this Ligand-Target Pair
Target5'-AMP-activated protein kinase catalytic subunit alpha-2/subunit beta-1/subunit gamma-1(Human)
Nanjing University of Chinese Medicine
Curated by ChEMBL
Nanjing University of Chinese Medicine
Curated by ChEMBL
Affinity DataEC50: 3.00E+3nMAssay Description:Allosteric activation of AMPK alpha2/beta1/gamma1 in human HepG2 cells by scintillation proximity assayMore data for this Ligand-Target Pair
Target5'-AMP-activated protein kinase catalytic subunit alpha-2/subunit beta-1/subunit gamma-1(Human)
Nanjing University of Chinese Medicine
Curated by ChEMBL
Nanjing University of Chinese Medicine
Curated by ChEMBL
Affinity DataEC50: 3.50E+3nMAssay Description:Allosteric activation of AMPK alpha2/beta1/gamma1 in human HepG2 cells by scintillation proximity assayMore data for this Ligand-Target Pair
Target5'-AMP-activated protein kinase catalytic subunit alpha-2/subunit beta-1/subunit gamma-1(Human)
Nanjing University of Chinese Medicine
Curated by ChEMBL
Nanjing University of Chinese Medicine
Curated by ChEMBL
Affinity DataEC50: 4.10E+3nMAssay Description:Allosteric activation of AMPK alpha2/beta1/gamma1 in human HepG2 cells by scintillation proximity assayMore data for this Ligand-Target Pair
Target5'-AMP-activated protein kinase catalytic subunit alpha-2/subunit beta-1/subunit gamma-1(Human)
Nanjing University of Chinese Medicine
Curated by ChEMBL
Nanjing University of Chinese Medicine
Curated by ChEMBL
Affinity DataEC50: 4.70E+3nMAssay Description:Allosteric activation of AMPK alpha2/beta1/gamma1 in human HepG2 cells by scintillation proximity assayMore data for this Ligand-Target Pair
Target5'-AMP-activated protein kinase catalytic subunit alpha-2/subunit beta-1/subunit gamma-1(Human)
Nanjing University of Chinese Medicine
Curated by ChEMBL
Nanjing University of Chinese Medicine
Curated by ChEMBL
Affinity DataEC50: 5.00E+3nMAssay Description:Allosteric activation of AMPK alpha2/beta1/gamma1 in human HepG2 cells by scintillation proximity assayMore data for this Ligand-Target Pair
Target5'-AMP-activated protein kinase catalytic subunit alpha-2/subunit beta-1/subunit gamma-1(Human)
Nanjing University of Chinese Medicine
Curated by ChEMBL
Nanjing University of Chinese Medicine
Curated by ChEMBL
Affinity DataEC50: 7.40E+3nMAssay Description:Allosteric activation of AMPK alpha2/beta1/gamma1 in human HepG2 cells by scintillation proximity assayMore data for this Ligand-Target Pair