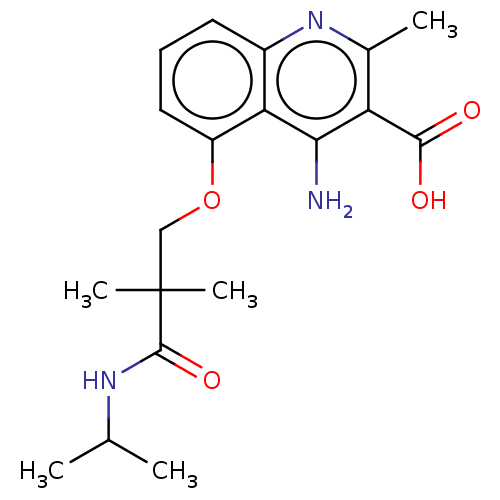
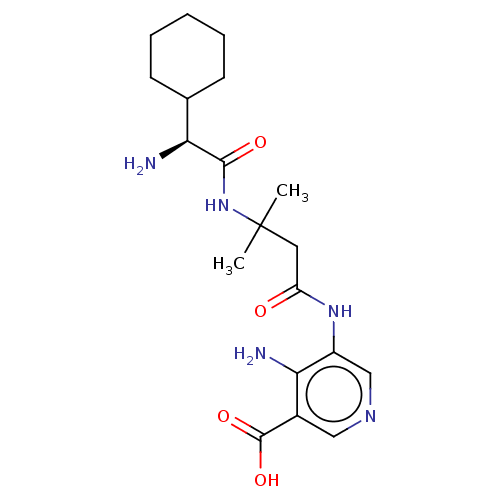
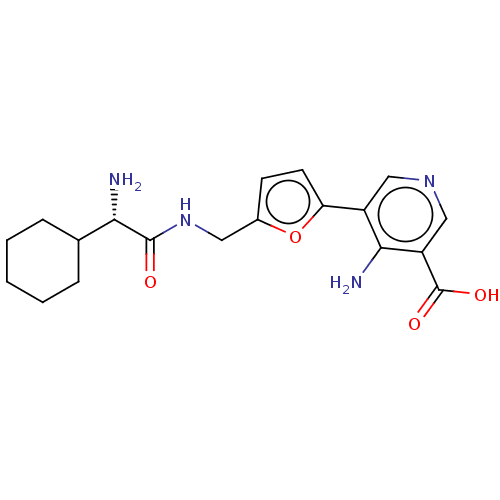
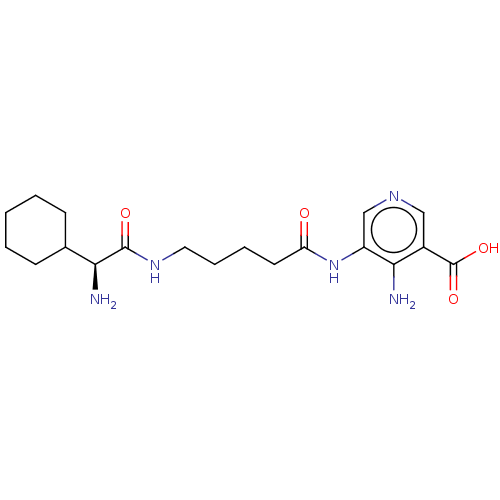
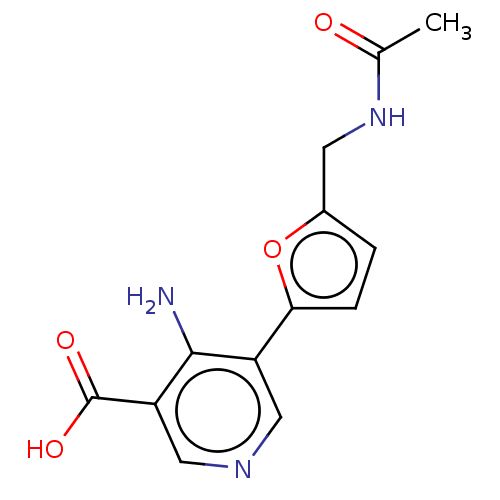
Report error Found 25 Enz. Inhib. hit(s) with all data for entry = 50010138
Affinity DataEC50: 6.00E+5nMAssay Description:Positive allosteric modulator activity at human T1R2/T1R3 expressed in PEAKrapid cells assessed as potentiation of sucrose-induced intracellular calc...More data for this Ligand-Target Pair
Affinity DataEC50: 2.20E+6nMAssay Description:Positive allosteric modulator activity at human T1R2/T1R3 expressed in PEAKrapid cells assessed as potentiation of sucrose-induced intracellular calc...More data for this Ligand-Target Pair
Affinity DataEC50: 5.80E+6nMAssay Description:Positive allosteric modulator activity at human T1R2/T1R3 expressed in PEAKrapid cells assessed as potentiation of sucrose-induced intracellular calc...More data for this Ligand-Target Pair
Affinity DataEC50: 7.80E+6nMAssay Description:Positive allosteric modulator activity at human T1R2/T1R3 expressed in PEAKrapid cells assessed as potentiation of sucrose-induced intracellular calc...More data for this Ligand-Target Pair
Affinity DataEC50: 8.50E+6nMAssay Description:Positive allosteric modulator activity at human T1R2/T1R3 expressed in PEAKrapid cells assessed as potentiation of sucrose-induced intracellular calc...More data for this Ligand-Target Pair
Affinity DataEC50: 8.90E+6nMAssay Description:Positive allosteric modulator activity at human T1R2/T1R3 expressed in PEAKrapid cells assessed as potentiation of sucrose-induced intracellular calc...More data for this Ligand-Target Pair
Affinity DataEC50: 1.00E+7nMAssay Description:Positive allosteric modulator activity at human T1R2/T1R3 expressed in PEAKrapid cells assessed as potentiation of sucrose-induced intracellular calc...More data for this Ligand-Target Pair
Affinity DataEC50: 1.02E+7nMAssay Description:Positive allosteric modulator activity at human T1R2/T1R3 expressed in PEAKrapid cells assessed as potentiation of sucrose-induced intracellular calc...More data for this Ligand-Target Pair
Affinity DataEC50: 1.05E+7nMAssay Description:Positive allosteric modulator activity at human T1R2/T1R3 expressed in PEAKrapid cells assessed as potentiation of sucrose-induced intracellular calc...More data for this Ligand-Target Pair
Affinity DataEC50: 1.11E+7nMAssay Description:Positive allosteric modulator activity at human T1R2/T1R3 expressed in PEAKrapid cells assessed as potentiation of sucrose-induced intracellular calc...More data for this Ligand-Target Pair
Affinity DataEC50: 1.46E+7nMAssay Description:Positive allosteric modulator activity at human T1R2/T1R3 expressed in PEAKrapid cells assessed as potentiation of sucrose-induced intracellular calc...More data for this Ligand-Target Pair
Affinity DataEC50: 1.59E+7nMAssay Description:Positive allosteric modulator activity at human T1R2/T1R3 expressed in PEAKrapid cells assessed as potentiation of sucrose-induced intracellular calc...More data for this Ligand-Target Pair
Affinity DataEC50: 1.62E+7nMAssay Description:Positive allosteric modulator activity at human T1R2/T1R3 expressed in PEAKrapid cells assessed as potentiation of sucrose-induced intracellular calc...More data for this Ligand-Target Pair
Affinity DataEC50: 1.68E+7nMAssay Description:Positive allosteric modulator activity at human T1R2/T1R3 expressed in PEAKrapid cells assessed as potentiation of sucrose-induced intracellular calc...More data for this Ligand-Target Pair
Affinity DataEC50: 1.83E+7nMAssay Description:Positive allosteric modulator activity at human T1R2/T1R3 expressed in PEAKrapid cells assessed as potentiation of sucrose-induced intracellular calc...More data for this Ligand-Target Pair
Affinity DataEC50: 1.84E+7nMAssay Description:Positive allosteric modulator activity at human T1R2/T1R3 expressed in PEAKrapid cells assessed as potentiation of sucrose-induced intracellular calc...More data for this Ligand-Target Pair
Affinity DataEC50: 2.02E+7nMAssay Description:Positive allosteric modulator activity at human T1R2/T1R3 expressed in PEAKrapid cells assessed as potentiation of sucrose-induced intracellular calc...More data for this Ligand-Target Pair
Affinity DataEC50: 2.10E+7nMAssay Description:Positive allosteric modulator activity at human T1R2/T1R3 expressed in PEAKrapid cells assessed as potentiation of sucrose-induced intracellular calc...More data for this Ligand-Target Pair
Affinity DataEC50: 2.15E+7nMAssay Description:Positive allosteric modulator activity at human T1R2/T1R3 expressed in PEAKrapid cells assessed as potentiation of sucrose-induced intracellular calc...More data for this Ligand-Target Pair
Affinity DataEC50: 2.16E+7nMAssay Description:Positive allosteric modulator activity at human T1R2/T1R3 expressed in PEAKrapid cells assessed as potentiation of sucrose-induced intracellular calc...More data for this Ligand-Target Pair
Affinity DataEC50: 2.21E+7nMAssay Description:Positive allosteric modulator activity at human T1R2/T1R3 expressed in PEAKrapid cells assessed as potentiation of sucrose-induced intracellular calc...More data for this Ligand-Target Pair
Affinity DataEC50: 2.27E+7nMAssay Description:Positive allosteric modulator activity at human T1R2/T1R3 expressed in PEAKrapid cells assessed as potentiation of sucrose-induced intracellular calc...More data for this Ligand-Target Pair
Affinity DataEC50: 2.45E+7nMAssay Description:Positive allosteric modulator activity at human T1R2/T1R3 expressed in PEAKrapid cells assessed as potentiation of sucrose-induced intracellular calc...More data for this Ligand-Target Pair
Affinity DataEC50: 2.65E+7nMAssay Description:Positive allosteric modulator activity at human T1R2/T1R3 expressed in PEAKrapid cells assessed as potentiation of sucrose-induced intracellular calc...More data for this Ligand-Target Pair
Affinity DataEC50: 2.71E+7nMAssay Description:Positive allosteric modulator activity at human T1R2/T1R3 expressed in PEAKrapid cells assessed as potentiation of sucrose-induced intracellular calc...More data for this Ligand-Target Pair