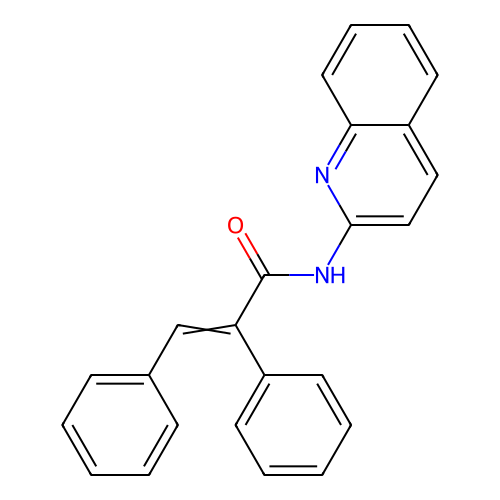
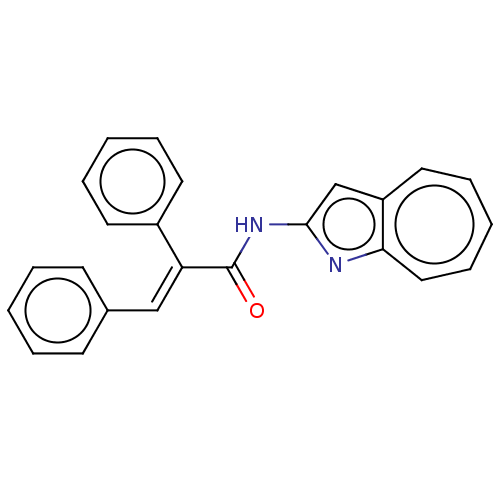
Report error Found 19 Enz. Inhib. hit(s) with all data for entry = 50007486
TargetcAMP and cAMP-inhibited cGMP 3',5'-cyclic phosphodiesterase 10A(Human)
Asubio Pharma
Curated by ChEMBL
Asubio Pharma
Curated by ChEMBL
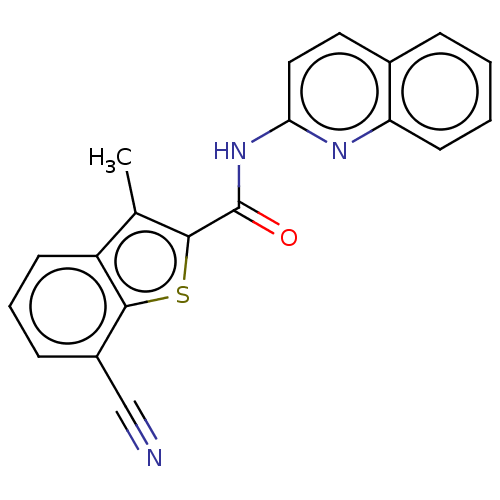
Affinity DataIC50: 2.60nMAssay Description:Inhibition of human MOLT4 cell-derived PDE10A using [3H]cAMP as substrate measured after 2 hrs in presence of rolipram/milrinone by topcount methodMore data for this Ligand-Target Pair
TargetcAMP and cAMP-inhibited cGMP 3',5'-cyclic phosphodiesterase 10A(Human)
Asubio Pharma
Curated by ChEMBL
Asubio Pharma
Curated by ChEMBL
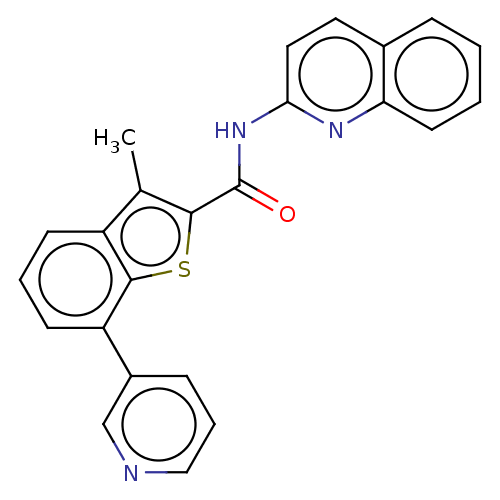
Affinity DataIC50: 4.70nMAssay Description:Inhibition of human MOLT4 cell-derived PDE10A using [3H]cAMP as substrate measured after 2 hrs in presence of rolipram/milrinone by topcount methodMore data for this Ligand-Target Pair
TargetcAMP and cAMP-inhibited cGMP 3',5'-cyclic phosphodiesterase 10A(Human)
Asubio Pharma
Curated by ChEMBL
Asubio Pharma
Curated by ChEMBL
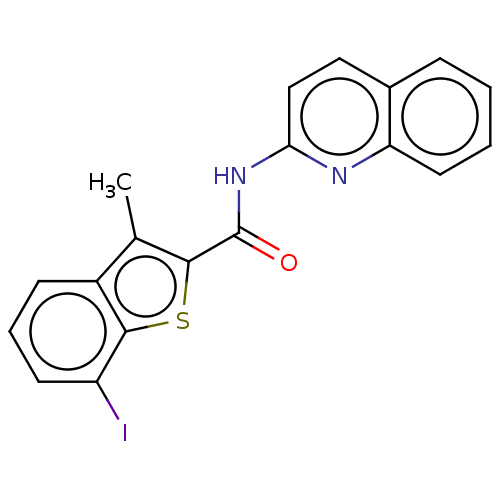
Affinity DataIC50: 5.20nMAssay Description:Inhibition of human MOLT4 cell-derived PDE10A using [3H]cAMP as substrate measured after 2 hrs in presence of rolipram/milrinone by topcount methodMore data for this Ligand-Target Pair
TargetcAMP and cAMP-inhibited cGMP 3',5'-cyclic phosphodiesterase 10A(Human)
Asubio Pharma
Curated by ChEMBL
Asubio Pharma
Curated by ChEMBL
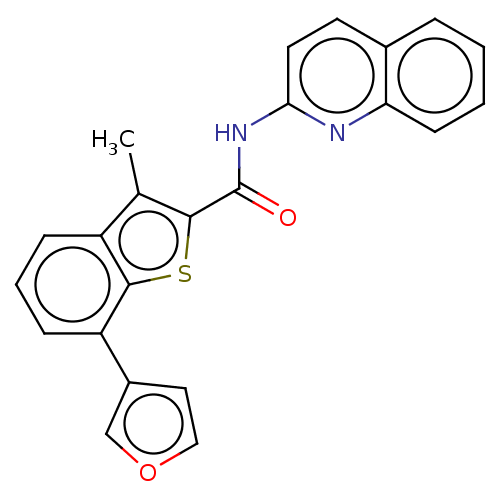
Affinity DataIC50: 7.60nMAssay Description:Inhibition of human MOLT4 cell-derived PDE10A using [3H]cAMP as substrate measured after 2 hrs in presence of rolipram/milrinone by topcount methodMore data for this Ligand-Target Pair
TargetcAMP and cAMP-inhibited cGMP 3',5'-cyclic phosphodiesterase 10A(Human)
Asubio Pharma
Curated by ChEMBL
Asubio Pharma
Curated by ChEMBL
Affinity DataIC50: 13nMAssay Description:Inhibition of human MOLT4 cell-derived PDE10A using [3H]cAMP as substrate measured after 2 hrs in presence of rolipram/milrinone by topcount methodMore data for this Ligand-Target Pair
TargetcAMP and cAMP-inhibited cGMP 3',5'-cyclic phosphodiesterase 10A(Human)
Asubio Pharma
Curated by ChEMBL
Asubio Pharma
Curated by ChEMBL
Affinity DataIC50: 23nMAssay Description:Inhibition of human MOLT4 cell-derived PDE10A using [3H]cAMP as substrate measured after 2 hrs in presence of rolipram/milrinone by topcount methodMore data for this Ligand-Target Pair
TargetcAMP and cAMP-inhibited cGMP 3',5'-cyclic phosphodiesterase 10A(Human)
Asubio Pharma
Curated by ChEMBL
Asubio Pharma
Curated by ChEMBL
Affinity DataIC50: 23nMAssay Description:Inhibition of human MOLT4 cell-derived PDE10A using [3H]cAMP as substrate measured after 2 hrs in presence of rolipram/milrinone by topcount methodMore data for this Ligand-Target Pair
TargetcAMP and cAMP-inhibited cGMP 3',5'-cyclic phosphodiesterase 10A(Rat)
Asubio Pharma
Curated by ChEMBL
Asubio Pharma
Curated by ChEMBL
Affinity DataIC50: 36nMAssay Description:Inhibition of recombinant rat PDE10A expressed in Sf9 cells using cGMP/cAMP as substrate by scintillation proximity assayMore data for this Ligand-Target Pair
TargetcAMP and cAMP-inhibited cGMP 3',5'-cyclic phosphodiesterase 10A(Human)
Asubio Pharma
Curated by ChEMBL
Asubio Pharma
Curated by ChEMBL
Affinity DataIC50: 44nMAssay Description:Inhibition of human MOLT4 cell-derived PDE10A using [3H]cAMP as substrate measured after 2 hrs in presence of rolipram/milrinone by topcount methodMore data for this Ligand-Target Pair
TargetcAMP and cAMP-inhibited cGMP 3',5'-cyclic phosphodiesterase 10A(Human)
Asubio Pharma
Curated by ChEMBL
Asubio Pharma
Curated by ChEMBL
Affinity DataIC50: 47nMAssay Description:Inhibition of human MOLT4 cell-derived PDE10A using [3H]cAMP as substrate measured after 2 hrs in presence of rolipram/milrinone by topcount methodMore data for this Ligand-Target Pair
TargetcAMP and cAMP-inhibited cGMP 3',5'-cyclic phosphodiesterase 10A(Human)
Asubio Pharma
Curated by ChEMBL
Asubio Pharma
Curated by ChEMBL
Affinity DataIC50: 62nMAssay Description:Inhibition of human MOLT4 cell-derived PDE10A using [3H]cAMP as substrate measured after 2 hrs in presence of rolipram/milrinone by topcount methodMore data for this Ligand-Target Pair
TargetcAMP and cAMP-inhibited cGMP 3',5'-cyclic phosphodiesterase 10A(Human)
Asubio Pharma
Curated by ChEMBL
Asubio Pharma
Curated by ChEMBL
Affinity DataIC50: 97nMAssay Description:Inhibition of human MOLT4 cell-derived PDE10A using [3H]cAMP as substrate measured after 2 hrs in presence of rolipram/milrinone by topcount methodMore data for this Ligand-Target Pair
TargetcAMP and cAMP-inhibited cGMP 3',5'-cyclic phosphodiesterase 10A(Human)
Asubio Pharma
Curated by ChEMBL
Asubio Pharma
Curated by ChEMBL
Affinity DataIC50: 110nMAssay Description:Inhibition of human MOLT4 cell-derived PDE10A using [3H]cAMP as substrate measured after 2 hrs in presence of rolipram/milrinone by topcount methodMore data for this Ligand-Target Pair
TargetcAMP and cAMP-inhibited cGMP 3',5'-cyclic phosphodiesterase 10A(Human)
Asubio Pharma
Curated by ChEMBL
Asubio Pharma
Curated by ChEMBL
Affinity DataIC50: 120nMAssay Description:Inhibition of human MOLT4 cell-derived PDE10A using [3H]cAMP as substrate measured after 2 hrs in presence of rolipram/milrinone by topcount methodMore data for this Ligand-Target Pair
TargetcAMP and cAMP-inhibited cGMP 3',5'-cyclic phosphodiesterase 10A(Human)
Asubio Pharma
Curated by ChEMBL
Asubio Pharma
Curated by ChEMBL
Affinity DataIC50: 200nMAssay Description:Inhibition of human MOLT4 cell-derived PDE10A using [3H]cAMP as substrate measured after 2 hrs in presence of rolipram/milrinone by topcount methodMore data for this Ligand-Target Pair
TargetcAMP and cAMP-inhibited cGMP 3',5'-cyclic phosphodiesterase 10A(Human)
Asubio Pharma
Curated by ChEMBL
Asubio Pharma
Curated by ChEMBL
Affinity DataIC50: 320nMAssay Description:Inhibition of human MOLT4 cell-derived PDE10A using [3H]cAMP as substrate measured after 2 hrs in presence of rolipram/milrinone by topcount methodMore data for this Ligand-Target Pair
TargetcAMP and cAMP-inhibited cGMP 3',5'-cyclic phosphodiesterase 10A(Human)
Asubio Pharma
Curated by ChEMBL
Asubio Pharma
Curated by ChEMBL
Affinity DataIC50: 440nMAssay Description:Inhibition of human MOLT4 cell-derived PDE10A using [3H]cAMP as substrate measured after 2 hrs in presence of rolipram/milrinone by topcount methodMore data for this Ligand-Target Pair
TargetcAMP and cAMP-inhibited cGMP 3',5'-cyclic phosphodiesterase 10A(Human)
Asubio Pharma
Curated by ChEMBL
Asubio Pharma
Curated by ChEMBL
Affinity DataIC50: 510nMAssay Description:Inhibition of human MOLT4 cell-derived PDE10A using [3H]cAMP as substrate measured after 2 hrs in presence of rolipram/milrinone by topcount methodMore data for this Ligand-Target Pair
TargetHigh affinity cAMP-specific 3',5'-cyclic phosphodiesterase 7A/cAMP-specific 3',5'-cyclic phosphodiesterase 7B(Human)
Asubio Pharma
Curated by ChEMBL
Asubio Pharma
Curated by ChEMBL
Affinity DataIC50: 1.00E+4nMAssay Description:Inhibition of human MOLT4 cell-derived PDE7 using [3H]cAMP as substrate measured after 2 hrs in presence of rolipram/milrinone by topcount methodMore data for this Ligand-Target Pair
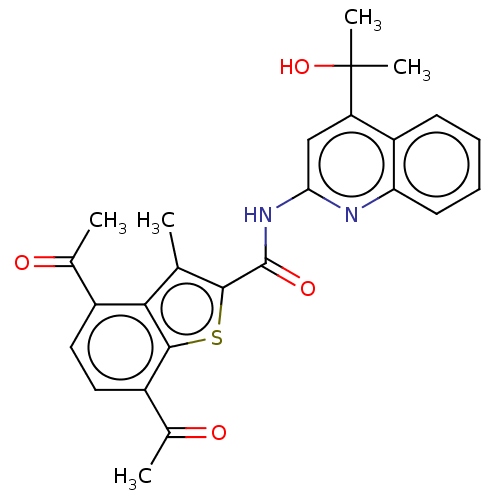
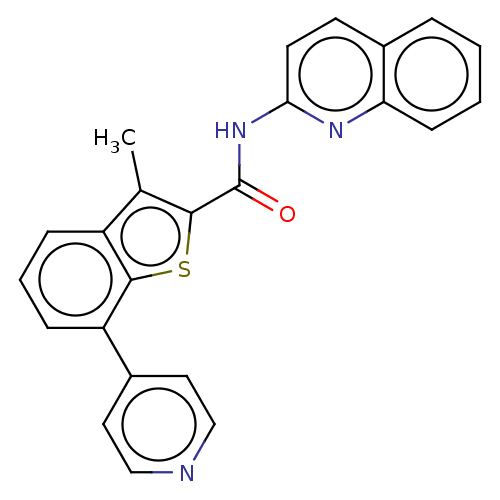
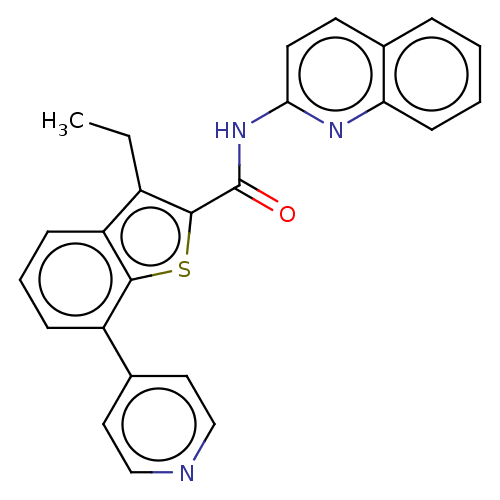
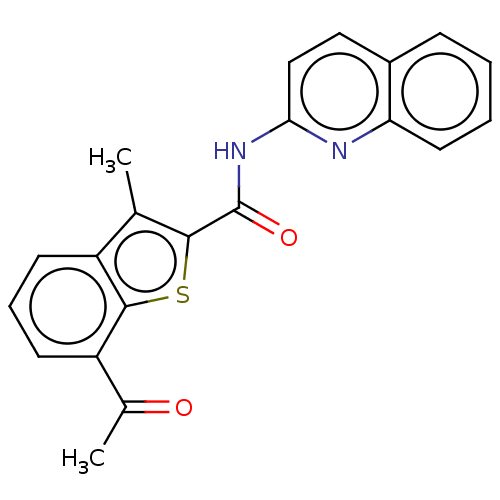
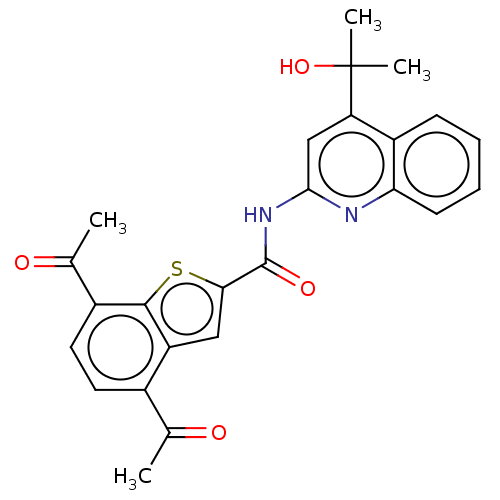
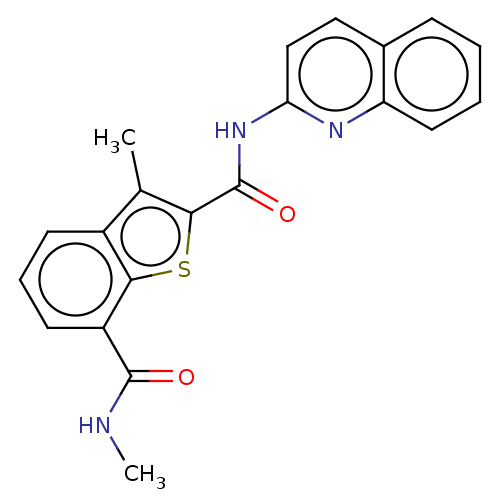
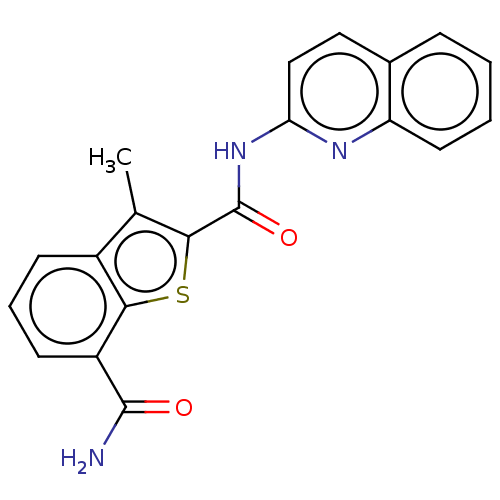
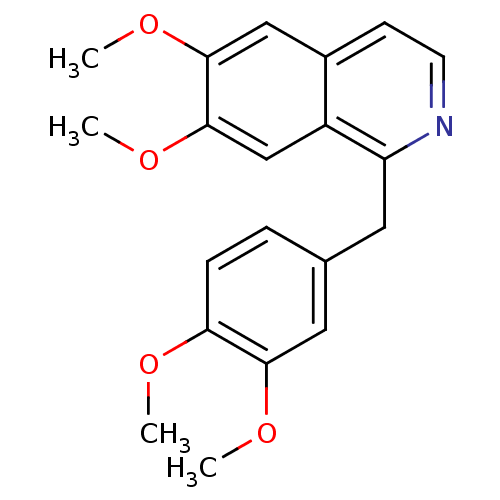
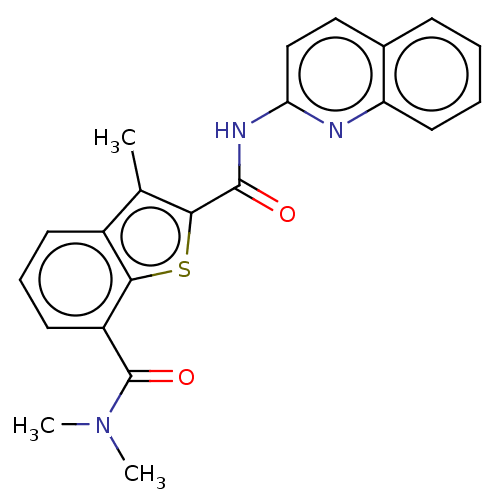
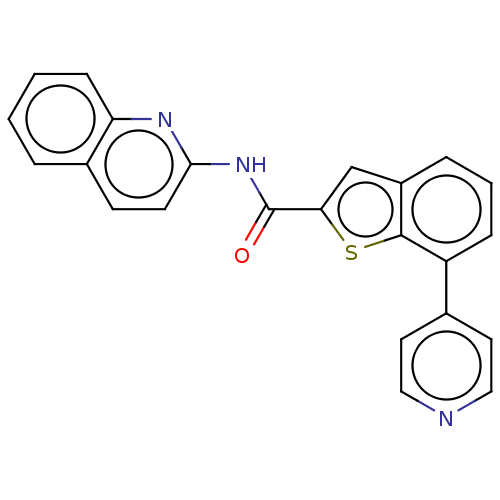
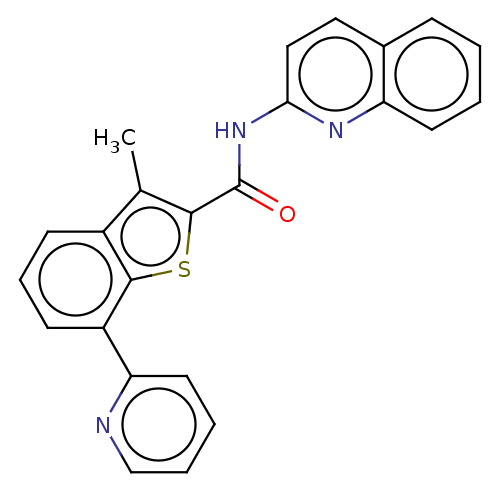
 3D Structure (crystal)
3D Structure (crystal)