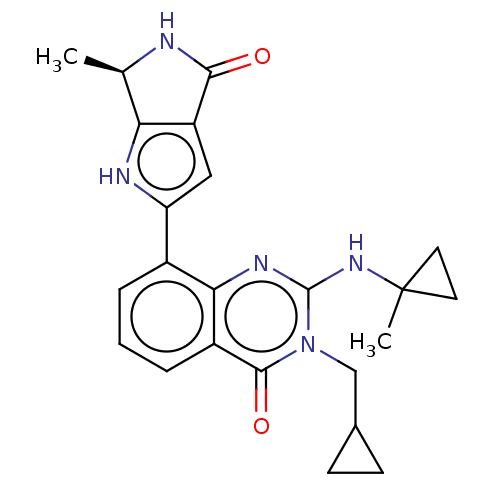
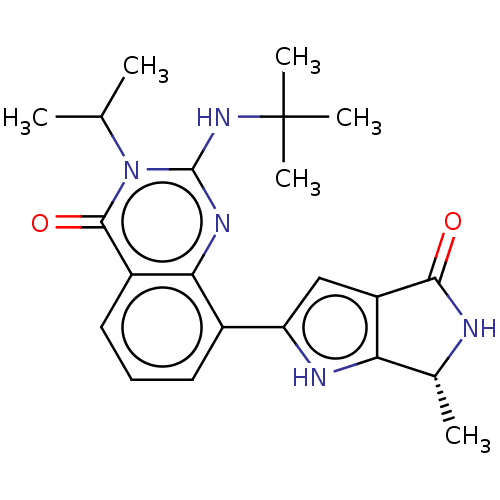
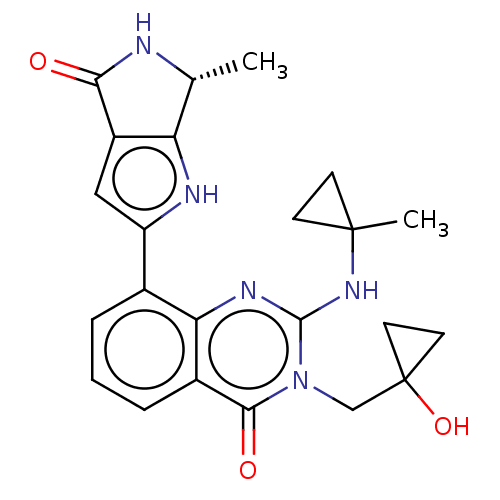
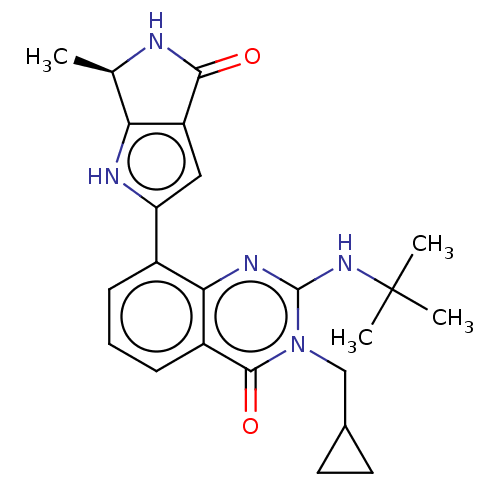
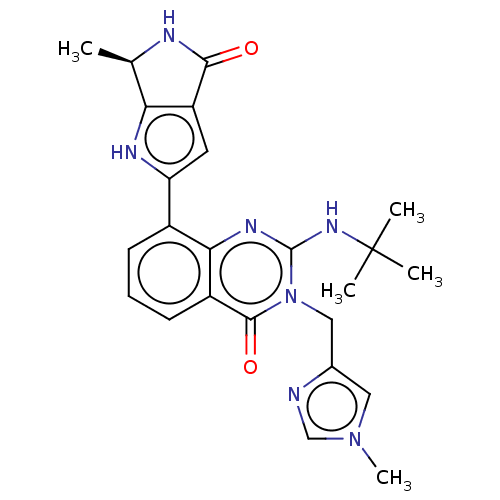
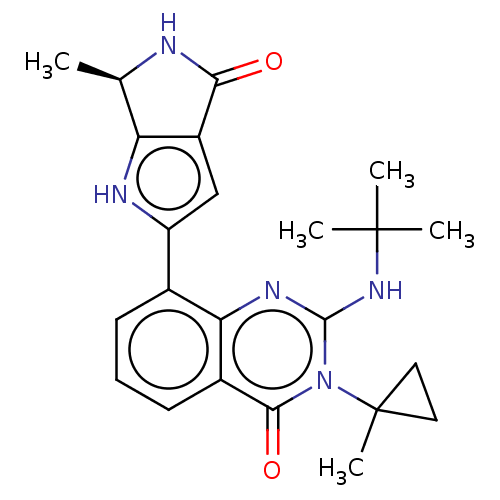
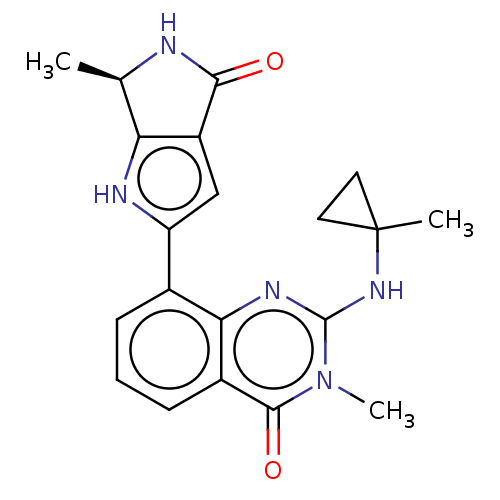
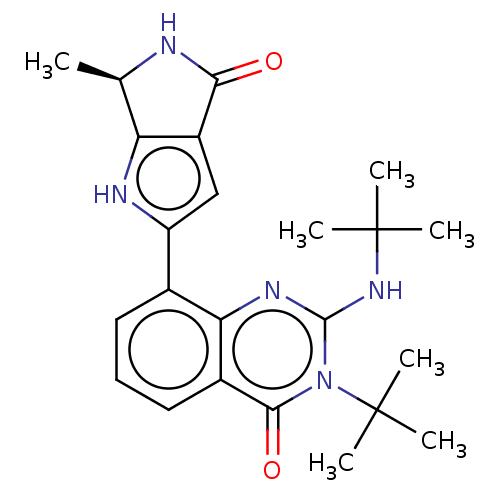
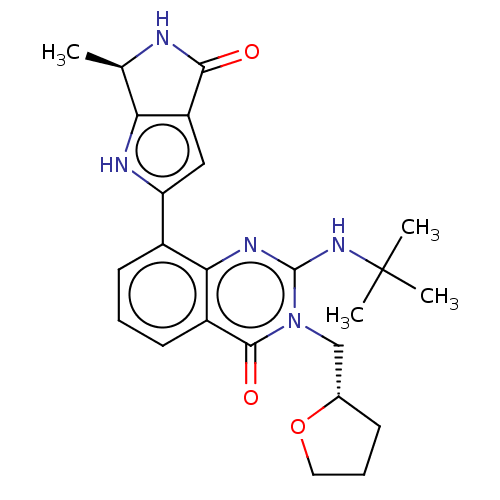
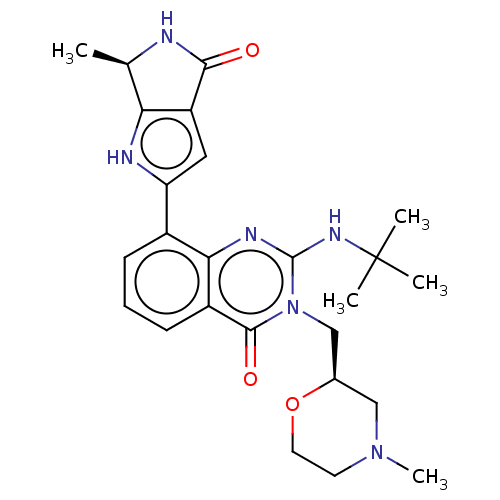
Report error Found 55 Enz. Inhib. hit(s) with all data for entry = 50008283
Affinity DataIC50: 0.0200nMAssay Description:Inhibition of full length recombinant PIM1 (unknown origin) assessed as reduction in BAD phosphorylation at Ser112 residues preincubated for 30 mins ...More data for this Ligand-Target Pair
Affinity DataIC50: 0.0300nMAssay Description:Inhibition of full length recombinant PIM1 (unknown origin) assessed as reduction in BAD phosphorylation at Ser112 residues preincubated for 30 mins ...More data for this Ligand-Target Pair
Affinity DataIC50: 0.0300nMAssay Description:Inhibition of full length recombinant PIM1 (unknown origin) assessed as reduction in BAD phosphorylation at Ser112 residues preincubated for 30 mins ...More data for this Ligand-Target Pair
Affinity DataIC50: 0.0500nMAssay Description:Inhibition of full length recombinant PIM2 (unknown origin) assessed as reduction in BAD phosphorylation at Ser112 residues preincubated for 30 mins ...More data for this Ligand-Target Pair
Affinity DataIC50: 0.0500nMAssay Description:Inhibition of full length recombinant PIM1 (unknown origin) assessed as reduction in BAD phosphorylation at Ser112 residues preincubated for 30 mins ...More data for this Ligand-Target Pair
Affinity DataIC50: 0.0600nMAssay Description:Inhibition of full length recombinant PIM1 (unknown origin) assessed as reduction in BAD phosphorylation at Ser112 residues preincubated for 30 mins ...More data for this Ligand-Target Pair
Affinity DataIC50: 0.0700nMAssay Description:Inhibition of full length recombinant PIM1 (unknown origin) assessed as reduction in BAD phosphorylation at Ser112 residues preincubated for 30 mins ...More data for this Ligand-Target Pair
Affinity DataIC50: 0.0800nMAssay Description:Inhibition of full length recombinant PIM1 (unknown origin) assessed as reduction in BAD phosphorylation at Ser112 residues preincubated for 30 mins ...More data for this Ligand-Target Pair
Affinity DataIC50: 0.0800nMAssay Description:Inhibition of full length recombinant PIM2 (unknown origin) assessed as reduction in BAD phosphorylation at Ser112 residues preincubated for 30 mins ...More data for this Ligand-Target Pair
Affinity DataIC50: 0.0900nMAssay Description:Inhibition of full length recombinant PIM2 (unknown origin) assessed as reduction in BAD phosphorylation at Ser112 residues preincubated for 30 mins ...More data for this Ligand-Target Pair
Affinity DataIC50: 0.0900nMAssay Description:Inhibition of full length recombinant PIM1 (unknown origin) assessed as reduction in BAD phosphorylation at Ser112 residues preincubated for 30 mins ...More data for this Ligand-Target Pair
Affinity DataIC50: 0.100nMAssay Description:Inhibition of full length recombinant PIM1 (unknown origin) assessed as reduction in BAD phosphorylation at Ser112 residues preincubated for 30 mins ...More data for this Ligand-Target Pair
Affinity DataIC50: 0.110nMAssay Description:Inhibition of full length recombinant PIM2 (unknown origin) assessed as reduction in BAD phosphorylation at Ser112 residues preincubated for 30 mins ...More data for this Ligand-Target Pair
Affinity DataIC50: 0.110nMAssay Description:Inhibition of full length recombinant PIM1 (unknown origin) assessed as reduction in BAD phosphorylation at Ser112 residues preincubated for 30 mins ...More data for this Ligand-Target Pair
Affinity DataIC50: 0.120nMAssay Description:Inhibition of full length recombinant PIM1 (unknown origin) assessed as reduction in BAD phosphorylation at Ser112 residues preincubated for 30 mins ...More data for this Ligand-Target Pair
Affinity DataIC50: 0.120nMAssay Description:Inhibition of full length recombinant PIM2 (unknown origin) assessed as reduction in BAD phosphorylation at Ser112 residues preincubated for 30 mins ...More data for this Ligand-Target Pair
Affinity DataIC50: 0.120nMAssay Description:Inhibition of full length recombinant PIM2 (unknown origin) assessed as reduction in BAD phosphorylation at Ser112 residues preincubated for 30 mins ...More data for this Ligand-Target Pair
Affinity DataIC50: 0.130nMAssay Description:Inhibition of full length recombinant PIM2 (unknown origin) assessed as reduction in BAD phosphorylation at Ser112 residues preincubated for 30 mins ...More data for this Ligand-Target Pair
Affinity DataIC50: 0.130nMAssay Description:Inhibition of full length recombinant PIM1 (unknown origin) assessed as reduction in BAD phosphorylation at Ser112 residues preincubated for 30 mins ...More data for this Ligand-Target Pair
Affinity DataIC50: 0.130nMAssay Description:Inhibition of full length recombinant PIM2 (unknown origin) assessed as reduction in BAD phosphorylation at Ser112 residues preincubated for 30 mins ...More data for this Ligand-Target Pair
Affinity DataIC50: 0.140nMAssay Description:Inhibition of full length recombinant PIM1 (unknown origin) assessed as reduction in BAD phosphorylation at Ser112 residues preincubated for 30 mins ...More data for this Ligand-Target Pair
Affinity DataIC50: 0.150nMAssay Description:Inhibition of full length recombinant PIM2 (unknown origin) assessed as reduction in BAD phosphorylation at Ser112 residues preincubated for 30 mins ...More data for this Ligand-Target Pair
Affinity DataIC50: 0.160nMAssay Description:Inhibition of full length recombinant PIM1 (unknown origin) assessed as reduction in BAD phosphorylation at Ser112 residues preincubated for 30 mins ...More data for this Ligand-Target Pair
Affinity DataIC50: 0.160nMAssay Description:Inhibition of full length recombinant PIM2 (unknown origin) assessed as reduction in BAD phosphorylation at Ser112 residues preincubated for 30 mins ...More data for this Ligand-Target Pair
Affinity DataIC50: 0.160nMAssay Description:Inhibition of full length recombinant PIM2 (unknown origin) assessed as reduction in BAD phosphorylation at Ser112 residues preincubated for 30 mins ...More data for this Ligand-Target Pair
Affinity DataIC50: 0.160nMAssay Description:Inhibition of full length recombinant PIM2 (unknown origin) assessed as reduction in BAD phosphorylation at Ser112 residues preincubated for 30 mins ...More data for this Ligand-Target Pair
Affinity DataIC50: 0.170nMAssay Description:Inhibition of full length recombinant PIM1 (unknown origin) assessed as reduction in BAD phosphorylation at Ser112 residues preincubated for 30 mins ...More data for this Ligand-Target Pair
Affinity DataIC50: 0.180nMAssay Description:Inhibition of full length recombinant PIM2 (unknown origin) assessed as reduction in BAD phosphorylation at Ser112 residues preincubated for 30 mins ...More data for this Ligand-Target Pair
Affinity DataIC50: 0.180nMAssay Description:Inhibition of full length recombinant PIM2 (unknown origin) assessed as reduction in BAD phosphorylation at Ser112 residues preincubated for 30 mins ...More data for this Ligand-Target Pair
Affinity DataIC50: 0.200nMAssay Description:Inhibition of full length recombinant PIM1 (unknown origin) assessed as reduction in BAD phosphorylation at Ser112 residues preincubated for 30 mins ...More data for this Ligand-Target Pair
Affinity DataIC50: 0.210nMAssay Description:Inhibition of full length recombinant PIM1 (unknown origin) assessed as reduction in BAD phosphorylation at Ser112 residues preincubated for 30 mins ...More data for this Ligand-Target Pair
Affinity DataIC50: 0.230nMAssay Description:Inhibition of full length recombinant PIM2 (unknown origin) assessed as reduction in BAD phosphorylation at Ser112 residues preincubated for 30 mins ...More data for this Ligand-Target Pair
Affinity DataIC50: 0.240nMAssay Description:Inhibition of full length recombinant PIM2 (unknown origin) assessed as reduction in BAD phosphorylation at Ser112 residues preincubated for 30 mins ...More data for this Ligand-Target Pair
Affinity DataIC50: 0.290nMAssay Description:Inhibition of full length recombinant PIM1 (unknown origin) assessed as reduction in BAD phosphorylation at Ser112 residues preincubated for 30 mins ...More data for this Ligand-Target Pair
Affinity DataIC50: 0.340nMAssay Description:Inhibition of full length recombinant PIM1 (unknown origin) assessed as reduction in BAD phosphorylation at Ser112 residues preincubated for 30 mins ...More data for this Ligand-Target Pair
Affinity DataIC50: 0.350nMAssay Description:Inhibition of full length recombinant PIM1 (unknown origin) assessed as reduction in BAD phosphorylation at Ser112 residues preincubated for 30 mins ...More data for this Ligand-Target Pair
Affinity DataIC50: 0.360nMAssay Description:Inhibition of full length recombinant PIM1 (unknown origin) assessed as reduction in BAD phosphorylation at Ser112 residues preincubated for 30 mins ...More data for this Ligand-Target Pair
Affinity DataIC50: 0.380nMAssay Description:Inhibition of full length recombinant PIM2 (unknown origin) assessed as reduction in BAD phosphorylation at Ser112 residues preincubated for 30 mins ...More data for this Ligand-Target Pair
Affinity DataIC50: 0.380nMAssay Description:Inhibition of full length recombinant PIM2 (unknown origin) assessed as reduction in BAD phosphorylation at Ser112 residues preincubated for 30 mins ...More data for this Ligand-Target Pair
Affinity DataIC50: 0.410nMAssay Description:Inhibition of full length recombinant PIM2 (unknown origin) assessed as reduction in BAD phosphorylation at Ser112 residues preincubated for 30 mins ...More data for this Ligand-Target Pair
Affinity DataIC50: 0.550nMAssay Description:Inhibition of full length recombinant PIM1 (unknown origin) assessed as reduction in BAD phosphorylation at Ser112 residues preincubated for 30 mins ...More data for this Ligand-Target Pair
Affinity DataIC50: 0.560nMAssay Description:Inhibition of full length recombinant PIM1 (unknown origin) assessed as reduction in BAD phosphorylation at Ser112 residues preincubated for 30 mins ...More data for this Ligand-Target Pair
Affinity DataIC50: 0.600nMAssay Description:Inhibition of full length recombinant PIM2 (unknown origin) assessed as reduction in BAD phosphorylation at Ser112 residues preincubated for 30 mins ...More data for this Ligand-Target Pair
Affinity DataIC50: 0.650nMAssay Description:Inhibition of full length recombinant PIM1 (unknown origin) assessed as reduction in BAD phosphorylation at Ser112 residues preincubated for 30 mins ...More data for this Ligand-Target Pair
Affinity DataIC50: 0.700nMAssay Description:Inhibition of full length recombinant PIM2 (unknown origin) assessed as reduction in BAD phosphorylation at Ser112 residues preincubated for 30 mins ...More data for this Ligand-Target Pair
Affinity DataIC50: 0.830nMAssay Description:Inhibition of full length recombinant PIM2 (unknown origin) assessed as reduction in BAD phosphorylation at Ser112 residues preincubated for 30 mins ...More data for this Ligand-Target Pair
Affinity DataIC50: 1.10nMAssay Description:Inhibition of full length recombinant PIM2 (unknown origin) assessed as reduction in BAD phosphorylation at Ser112 residues preincubated for 30 mins ...More data for this Ligand-Target Pair
Affinity DataIC50: 1.20nMAssay Description:Inhibition of full length recombinant PIM1 (unknown origin) assessed as reduction in BAD phosphorylation at Ser112 residues preincubated for 30 mins ...More data for this Ligand-Target Pair
Affinity DataIC50: 1.60nMAssay Description:Inhibition of full length recombinant PIM2 (unknown origin) assessed as reduction in BAD phosphorylation at Ser112 residues preincubated for 30 mins ...More data for this Ligand-Target Pair
Affinity DataIC50: 1.80nMAssay Description:Inhibition of full length recombinant PIM1 (unknown origin) assessed as reduction in BAD phosphorylation at Ser112 residues preincubated for 30 mins ...More data for this Ligand-Target Pair