Report error Found 22 Enz. Inhib. hit(s) with all data for entry = 50008387
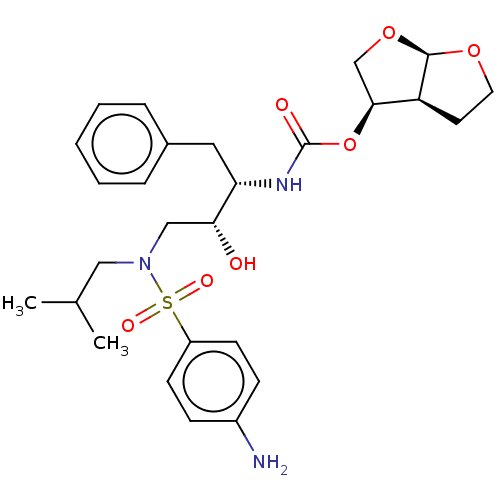
Affinity DataIC50: 0.460nMAssay Description:Inhibition of HIV1 protease expressed in Escherichia coli using Arg-Glu (EDANS)-Ser-Gln-Asn-Tyr-Pro-Ile-Val-Gln-Lys(DABCYL)-Arg as substrate preincub...More data for this Ligand-Target Pair
Affinity DataIC50: 120nMAssay Description:Inhibition of HIV1 protease expressed in Escherichia coli using Arg-Glu (EDANS)-Ser-Gln-Asn-Tyr-Pro-Ile-Val-Gln-Lys(DABCYL)-Arg as substrate preincub...More data for this Ligand-Target Pair
Affinity DataIC50: 150nMAssay Description:Inhibition of HIV1 protease expressed in Escherichia coli using Arg-Glu (EDANS)-Ser-Gln-Asn-Tyr-Pro-Ile-Val-Gln-Lys(DABCYL)-Arg as substrate preincub...More data for this Ligand-Target Pair
Affinity DataIC50: 150nMAssay Description:Inhibition of HIV1 protease expressed in Escherichia coli using Arg-Glu (EDANS)-Ser-Gln-Asn-Tyr-Pro-Ile-Val-Gln-Lys(DABCYL)-Arg as substrate preincub...More data for this Ligand-Target Pair
Affinity DataIC50: 170nMAssay Description:Inhibition of HIV1 protease expressed in Escherichia coli using Arg-Glu (EDANS)-Ser-Gln-Asn-Tyr-Pro-Ile-Val-Gln-Lys(DABCYL)-Arg as substrate preincub...More data for this Ligand-Target Pair
Affinity DataIC50: 170nMAssay Description:Inhibition of HIV1 protease expressed in Escherichia coli using Arg-Glu (EDANS)-Ser-Gln-Asn-Tyr-Pro-Ile-Val-Gln-Lys(DABCYL)-Arg as substrate preincub...More data for this Ligand-Target Pair
Affinity DataIC50: 210nMAssay Description:Inhibition of HIV1 protease expressed in Escherichia coli using Arg-Glu (EDANS)-Ser-Gln-Asn-Tyr-Pro-Ile-Val-Gln-Lys(DABCYL)-Arg as substrate preincub...More data for this Ligand-Target Pair
Affinity DataIC50: 220nMAssay Description:Inhibition of HIV1 protease expressed in Escherichia coli using Arg-Glu (EDANS)-Ser-Gln-Asn-Tyr-Pro-Ile-Val-Gln-Lys(DABCYL)-Arg as substrate preincub...More data for this Ligand-Target Pair
Affinity DataIC50: 240nMAssay Description:Inhibition of HIV1 protease expressed in Escherichia coli using Arg-Glu (EDANS)-Ser-Gln-Asn-Tyr-Pro-Ile-Val-Gln-Lys(DABCYL)-Arg as substrate preincub...More data for this Ligand-Target Pair
Affinity DataIC50: 300nMAssay Description:Inhibition of HIV1 protease expressed in Escherichia coli using Arg-Glu (EDANS)-Ser-Gln-Asn-Tyr-Pro-Ile-Val-Gln-Lys(DABCYL)-Arg as substrate preincub...More data for this Ligand-Target Pair
Affinity DataIC50: 370nMAssay Description:Inhibition of HIV1 protease expressed in Escherichia coli using Arg-Glu (EDANS)-Ser-Gln-Asn-Tyr-Pro-Ile-Val-Gln-Lys(DABCYL)-Arg as substrate preincub...More data for this Ligand-Target Pair
Affinity DataIC50: 400nMAssay Description:Inhibition of HIV1 protease expressed in Escherichia coli using Arg-Glu (EDANS)-Ser-Gln-Asn-Tyr-Pro-Ile-Val-Gln-Lys(DABCYL)-Arg as substrate preincub...More data for this Ligand-Target Pair
Affinity DataIC50: 460nMAssay Description:Inhibition of HIV1 protease expressed in Escherichia coli using Arg-Glu (EDANS)-Ser-Gln-Asn-Tyr-Pro-Ile-Val-Gln-Lys(DABCYL)-Arg as substrate preincub...More data for this Ligand-Target Pair
Affinity DataIC50: 480nMAssay Description:Inhibition of HIV1 protease expressed in Escherichia coli using Arg-Glu (EDANS)-Ser-Gln-Asn-Tyr-Pro-Ile-Val-Gln-Lys(DABCYL)-Arg as substrate preincub...More data for this Ligand-Target Pair
Affinity DataIC50: 490nMAssay Description:Inhibition of HIV1 protease expressed in Escherichia coli using Arg-Glu (EDANS)-Ser-Gln-Asn-Tyr-Pro-Ile-Val-Gln-Lys(DABCYL)-Arg as substrate preincub...More data for this Ligand-Target Pair
Affinity DataIC50: 500nMAssay Description:Inhibition of HIV1 protease expressed in Escherichia coli using Arg-Glu (EDANS)-Ser-Gln-Asn-Tyr-Pro-Ile-Val-Gln-Lys(DABCYL)-Arg as substrate preincub...More data for this Ligand-Target Pair
Affinity DataIC50: 580nMAssay Description:Inhibition of HIV1 protease expressed in Escherichia coli using Arg-Glu (EDANS)-Ser-Gln-Asn-Tyr-Pro-Ile-Val-Gln-Lys(DABCYL)-Arg as substrate preincub...More data for this Ligand-Target Pair
Affinity DataIC50: 610nMAssay Description:Inhibition of HIV1 protease expressed in Escherichia coli using Arg-Glu (EDANS)-Ser-Gln-Asn-Tyr-Pro-Ile-Val-Gln-Lys(DABCYL)-Arg as substrate preincub...More data for this Ligand-Target Pair
Affinity DataIC50: 1.05E+3nMAssay Description:Inhibition of HIV1 protease expressed in Escherichia coli using Arg-Glu (EDANS)-Ser-Gln-Asn-Tyr-Pro-Ile-Val-Gln-Lys(DABCYL)-Arg as substrate preincub...More data for this Ligand-Target Pair
Affinity DataIC50: 1.08E+3nMAssay Description:Inhibition of HIV1 protease expressed in Escherichia coli using Arg-Glu (EDANS)-Ser-Gln-Asn-Tyr-Pro-Ile-Val-Gln-Lys(DABCYL)-Arg as substrate preincub...More data for this Ligand-Target Pair
Affinity DataIC50: 1.85E+3nMAssay Description:Inhibition of HIV1 protease expressed in Escherichia coli using Arg-Glu (EDANS)-Ser-Gln-Asn-Tyr-Pro-Ile-Val-Gln-Lys(DABCYL)-Arg as substrate preincub...More data for this Ligand-Target Pair
Affinity DataIC50: 2.37E+3nMAssay Description:Inhibition of HIV1 protease expressed in Escherichia coli using Arg-Glu (EDANS)-Ser-Gln-Asn-Tyr-Pro-Ile-Val-Gln-Lys(DABCYL)-Arg as substrate preincub...More data for this Ligand-Target Pair