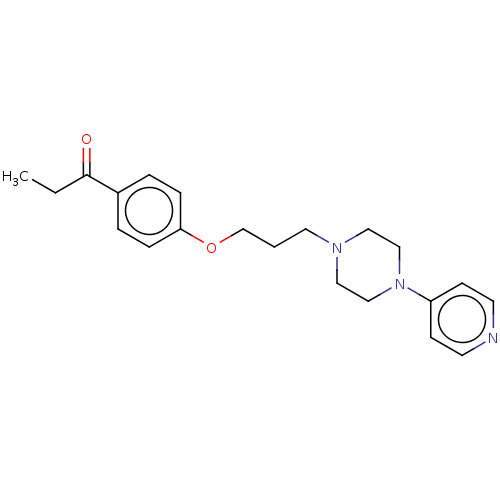
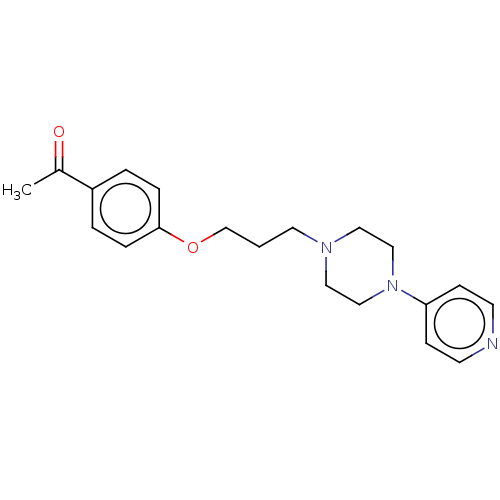
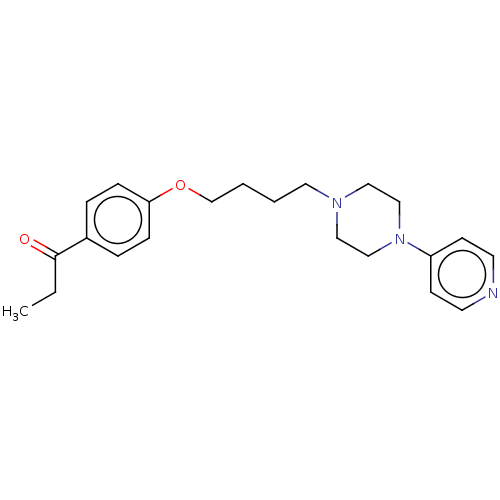
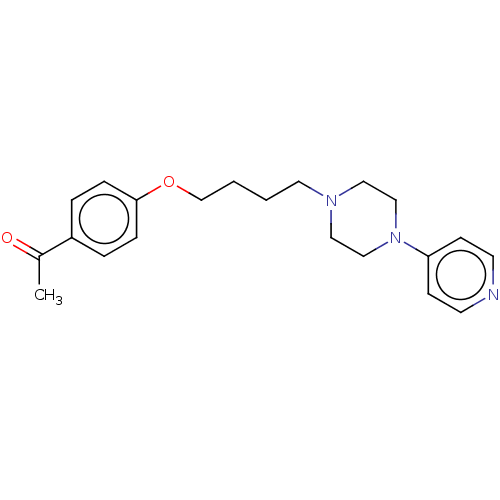
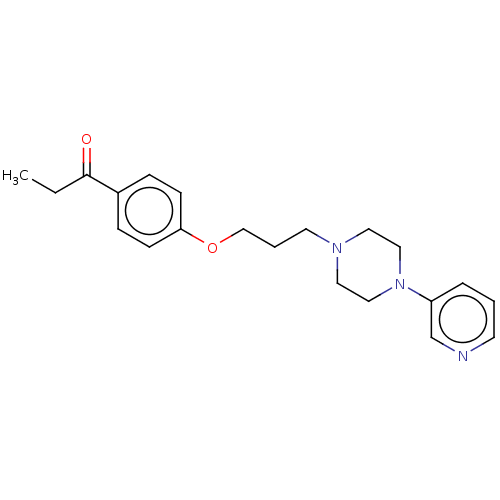
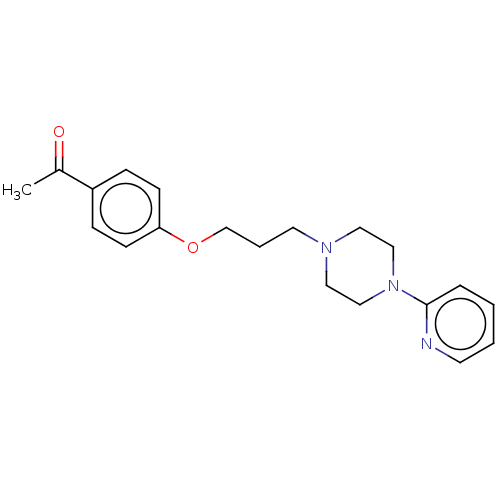
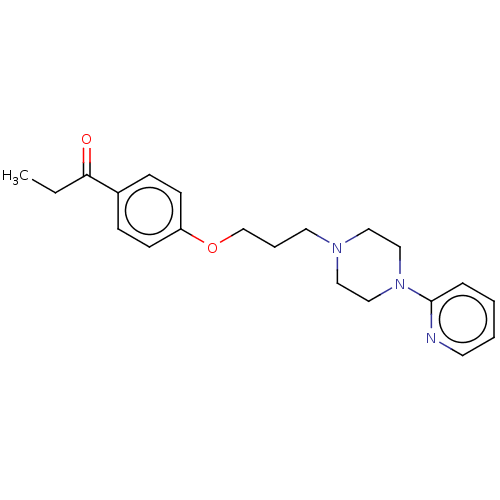
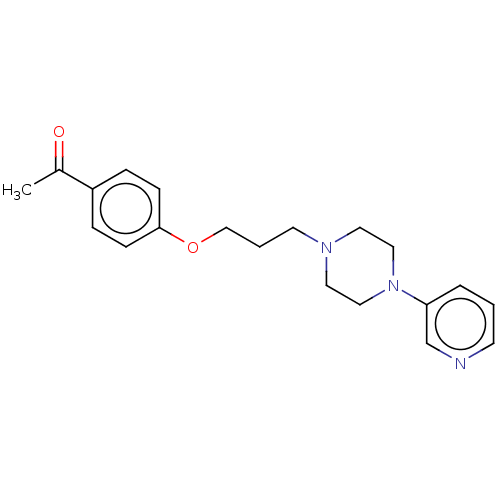
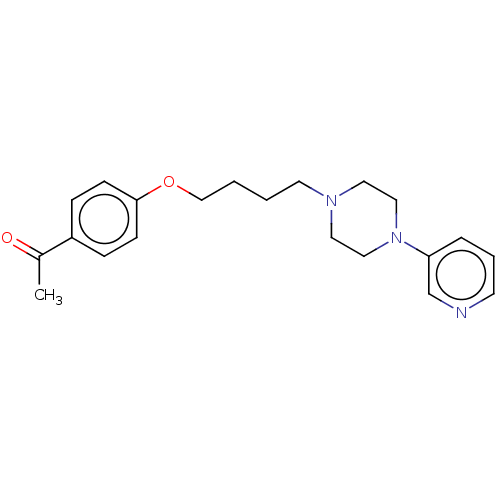
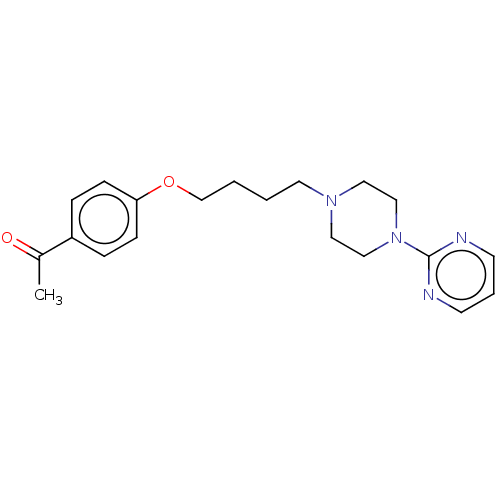
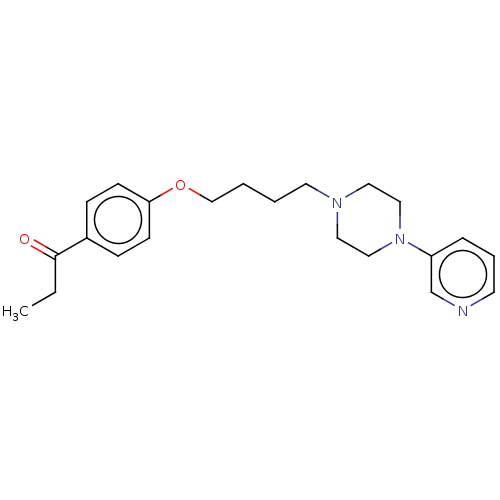
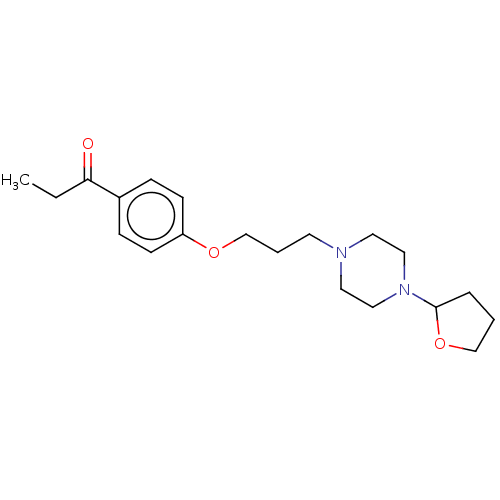
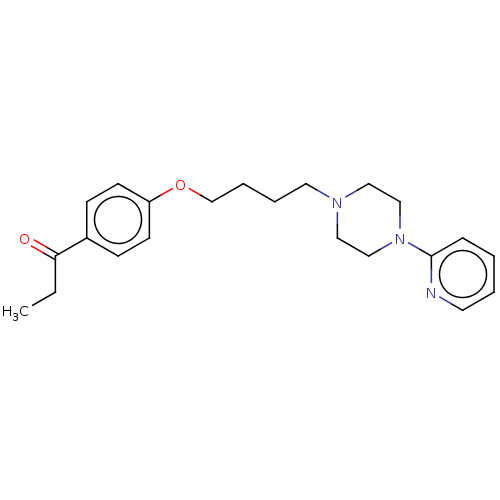
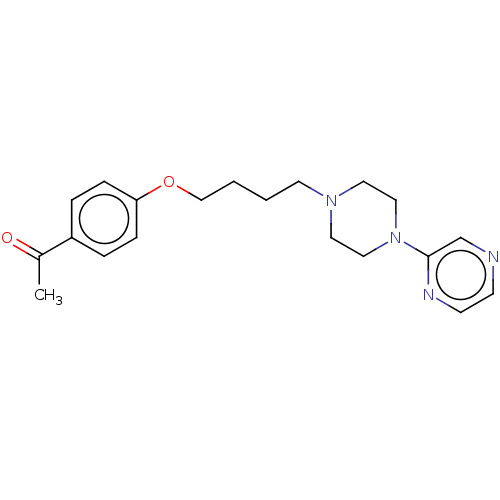
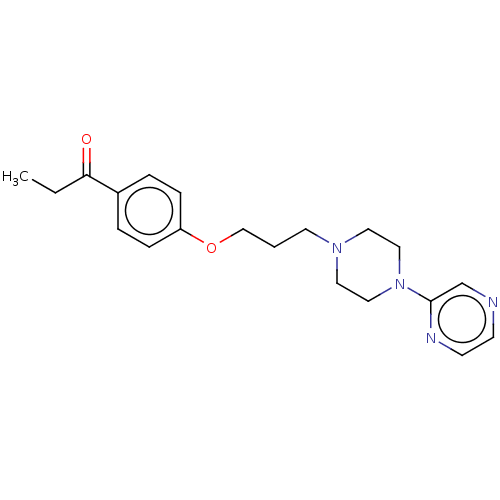
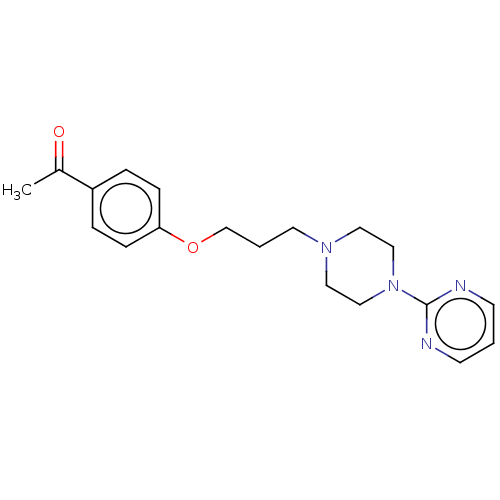
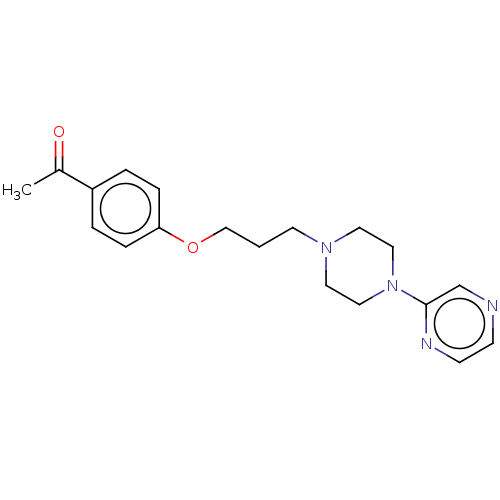
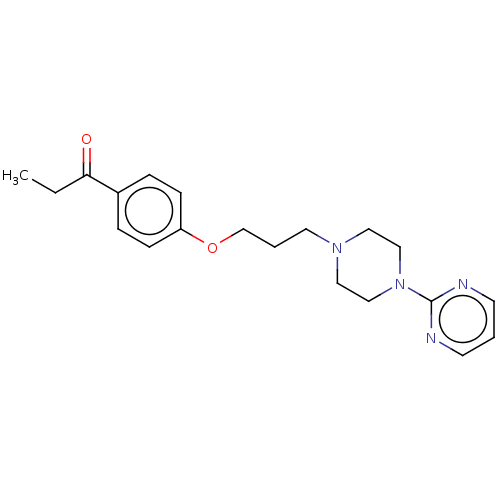
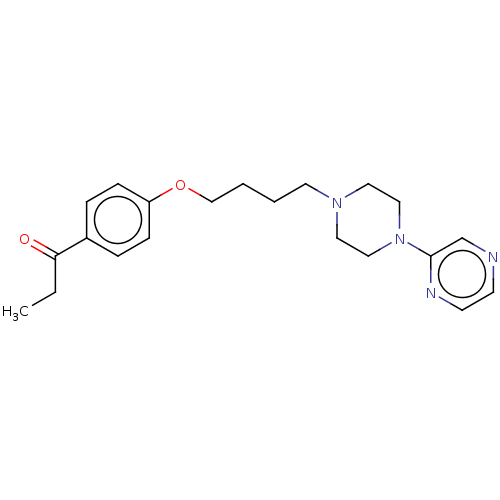
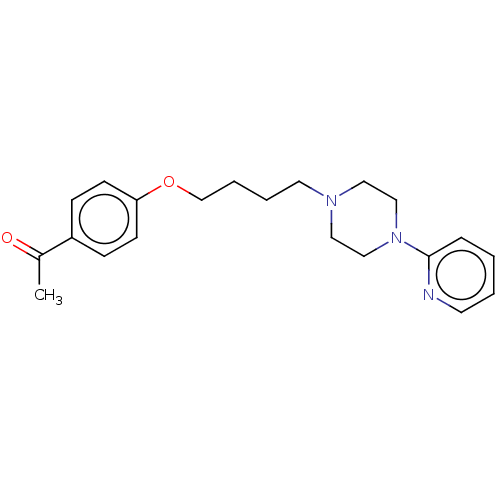
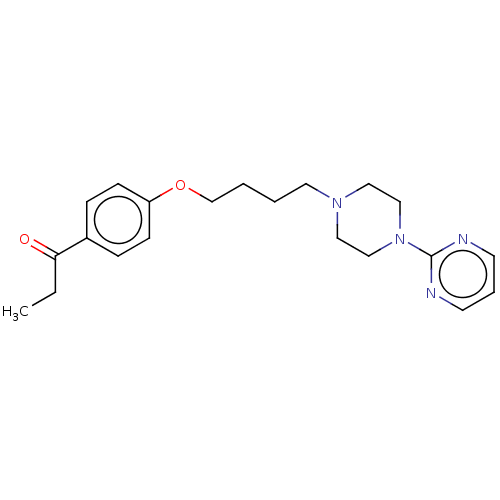
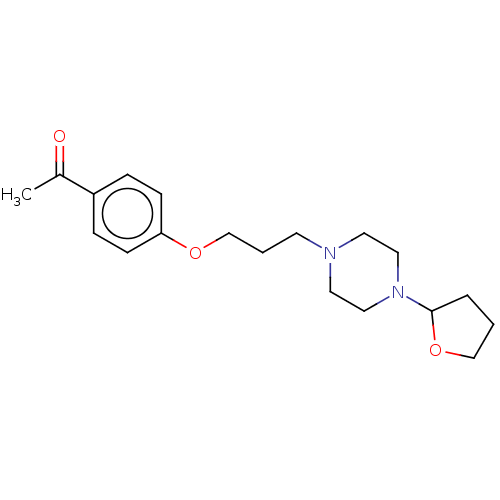
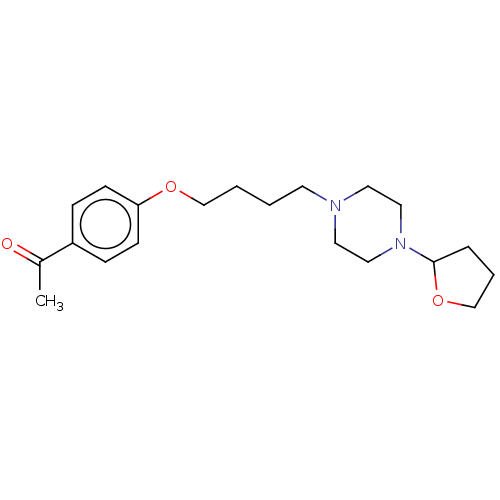
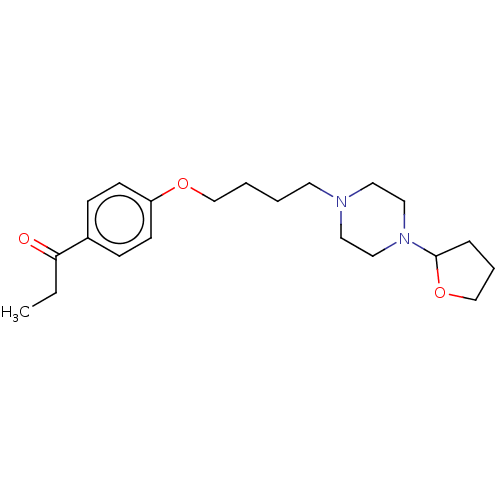
Report error Found 24 Enz. Inhib. hit(s) with all data for entry = 50008624
Affinity DataKi: 5.20nMAssay Description:Displacement of [3H]N-alpha-methylhistamine from full length recombinant human H3R expressed in HEK293 cell membranes after 90 mins by beta scintilla...More data for this Ligand-Target Pair
Affinity DataKi: 10nMAssay Description:Displacement of [3H]N-alpha-methylhistamine from full length recombinant human H3R expressed in HEK293 cell membranes after 90 mins by beta scintilla...More data for this Ligand-Target Pair
Affinity DataKi: 15nMAssay Description:Displacement of [3H]N-alpha-methylhistamine from full length recombinant human H3R expressed in HEK293 cell membranes after 90 mins by beta scintilla...More data for this Ligand-Target Pair
Affinity DataKi: 115nMAssay Description:Displacement of [3H]N-alpha-methylhistamine from full length recombinant human H3R expressed in HEK293 cell membranes after 90 mins by beta scintilla...More data for this Ligand-Target Pair
Affinity DataKi: 192nMAssay Description:Displacement of [3H]N-alpha-methylhistamine from full length recombinant human H3R expressed in HEK293 cell membranes after 90 mins by beta scintilla...More data for this Ligand-Target Pair
Affinity DataKi: 315nMAssay Description:Displacement of [3H]N-alpha-methylhistamine from full length recombinant human H3R expressed in HEK293 cell membranes after 90 mins by beta scintilla...More data for this Ligand-Target Pair
Affinity DataKi: 377nMAssay Description:Displacement of [3H]N-alpha-methylhistamine from full length recombinant human H3R expressed in HEK293 cell membranes after 90 mins by beta scintilla...More data for this Ligand-Target Pair
Affinity DataKi: 499nMAssay Description:Displacement of [3H]N-alpha-methylhistamine from full length recombinant human H3R expressed in HEK293 cell membranes after 90 mins by beta scintilla...More data for this Ligand-Target Pair
Affinity DataKi: 908nMAssay Description:Displacement of [3H]N-alpha-methylhistamine from full length recombinant human H3R expressed in HEK293 cell membranes after 90 mins by beta scintilla...More data for this Ligand-Target Pair
Affinity DataKi: 1.23E+3nMAssay Description:Displacement of [3H]N-alpha-methylhistamine from full length recombinant human H3R expressed in HEK293 cell membranes after 90 mins by beta scintilla...More data for this Ligand-Target Pair
Affinity DataKi: 1.39E+3nMAssay Description:Displacement of [3H]N-alpha-methylhistamine from full length recombinant human H3R expressed in HEK293 cell membranes after 90 mins by beta scintilla...More data for this Ligand-Target Pair
Affinity DataKi: 1.39E+3nMAssay Description:Displacement of [3H]N-alpha-methylhistamine from full length recombinant human H3R expressed in HEK293 cell membranes after 90 mins by beta scintilla...More data for this Ligand-Target Pair
Affinity DataKi: 1.44E+3nMAssay Description:Displacement of [3H]N-alpha-methylhistamine from full length recombinant human H3R expressed in HEK293 cell membranes after 90 mins by beta scintilla...More data for this Ligand-Target Pair
Affinity DataKi: 1.63E+3nMAssay Description:Displacement of [3H]N-alpha-methylhistamine from full length recombinant human H3R expressed in HEK293 cell membranes after 90 mins by beta scintilla...More data for this Ligand-Target Pair
Affinity DataKi: 1.72E+3nMAssay Description:Displacement of [3H]N-alpha-methylhistamine from full length recombinant human H3R expressed in HEK293 cell membranes after 90 mins by beta scintilla...More data for this Ligand-Target Pair
Affinity DataKi: 2.32E+3nMAssay Description:Displacement of [3H]N-alpha-methylhistamine from full length recombinant human H3R expressed in HEK293 cell membranes after 90 mins by beta scintilla...More data for this Ligand-Target Pair
Affinity DataKi: 2.45E+3nMAssay Description:Displacement of [3H]N-alpha-methylhistamine from full length recombinant human H3R expressed in HEK293 cell membranes after 90 mins by beta scintilla...More data for this Ligand-Target Pair
Affinity DataKi: 3.39E+3nMAssay Description:Displacement of [3H]N-alpha-methylhistamine from full length recombinant human H3R expressed in HEK293 cell membranes after 90 mins by beta scintilla...More data for this Ligand-Target Pair
Affinity DataKi: 3.63E+3nMAssay Description:Displacement of [3H]N-alpha-methylhistamine from full length recombinant human H3R expressed in HEK293 cell membranes after 90 mins by beta scintilla...More data for this Ligand-Target Pair
Affinity DataKi: 3.70E+3nMAssay Description:Displacement of [3H]N-alpha-methylhistamine from full length recombinant human H3R expressed in HEK293 cell membranes after 90 mins by beta scintilla...More data for this Ligand-Target Pair
Affinity DataKi: 4.04E+3nMAssay Description:Displacement of [3H]N-alpha-methylhistamine from full length recombinant human H3R expressed in HEK293 cell membranes after 90 mins by beta scintilla...More data for this Ligand-Target Pair
Affinity DataKi: 4.89E+3nMAssay Description:Displacement of [3H]N-alpha-methylhistamine from full length recombinant human H3R expressed in HEK293 cell membranes after 90 mins by beta scintilla...More data for this Ligand-Target Pair
Affinity DataKi: >1.00E+4nMAssay Description:Displacement of [3H]N-alpha-methylhistamine from full length recombinant human H3R expressed in HEK293 cell membranes after 90 mins by beta scintilla...More data for this Ligand-Target Pair
Affinity DataKi: >1.00E+4nMAssay Description:Displacement of [3H]N-alpha-methylhistamine from full length recombinant human H3R expressed in HEK293 cell membranes after 90 mins by beta scintilla...More data for this Ligand-Target Pair