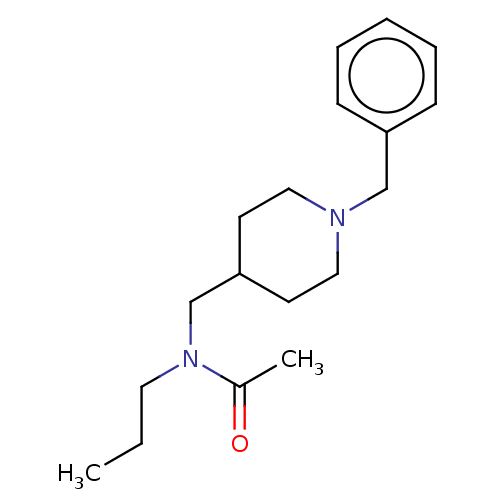
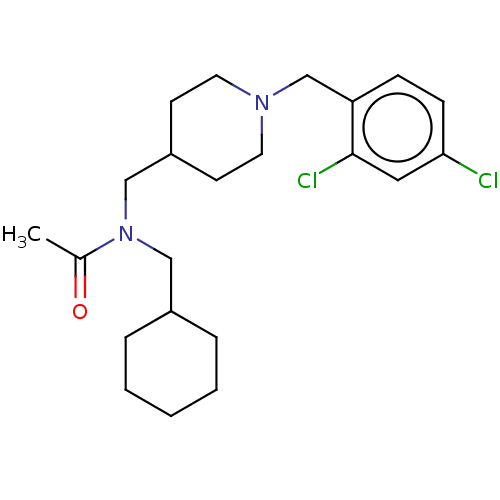
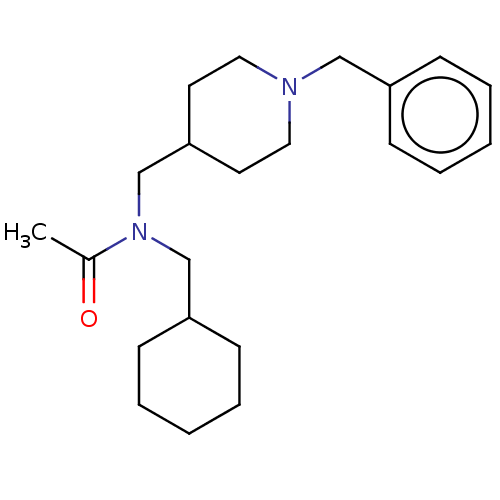
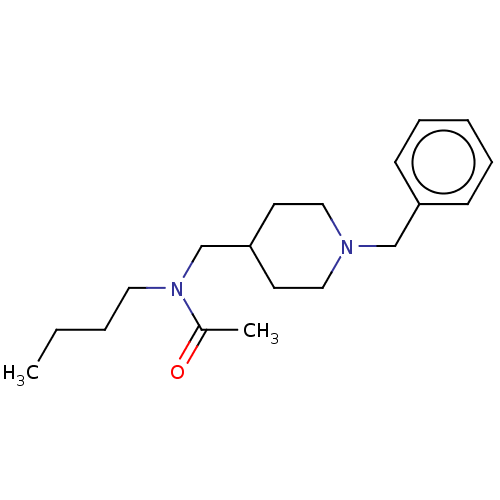
Report error Found 27 Enz. Inhib. hit(s) with all data for entry = 50003100
Affinity DataKi: 17nMAssay Description:Displacement of [3H]-(+)-pentazocine from sigma1 receptor in rat liver membranes after 120 mins by liquid scintillation countingMore data for this Ligand-Target Pair
Affinity DataKi: 20nMAssay Description:Displacement of [3H]-DTG from sigma2 receptor in rat liver membranes after 120 mins by liquid scintillation countingMore data for this Ligand-Target Pair
Affinity DataKi: 32nMAssay Description:Displacement of [3H]-(+)-pentazocine from sigma1 receptor in rat liver membranes after 120 mins by liquid scintillation countingMore data for this Ligand-Target Pair
Affinity DataKi: 34nMAssay Description:Displacement of [3H]-(+)-pentazocine from sigma1 receptor in rat liver membranes after 120 mins by liquid scintillation countingMore data for this Ligand-Target Pair
Affinity DataKi: 37nMAssay Description:Displacement of [3H]-(+)-pentazocine from sigma1 receptor in rat liver membranes after 120 mins by liquid scintillation countingMore data for this Ligand-Target Pair
Affinity DataKi: 52nMAssay Description:Displacement of [3H]-(+)-pentazocine from sigma1 receptor in rat liver membranes after 120 mins by liquid scintillation countingMore data for this Ligand-Target Pair
Affinity DataKi: 59nMAssay Description:Displacement of [3H]-(+)-pentazocine from sigma1 receptor in rat liver membranes after 120 mins by liquid scintillation countingMore data for this Ligand-Target Pair
Affinity DataKi: 67nMAssay Description:Displacement of [3H]-(+)-pentazocine from sigma1 receptor in rat liver membranes after 120 mins by liquid scintillation countingMore data for this Ligand-Target Pair
Affinity DataKi: 76nMAssay Description:Displacement of [3H]-(+)-pentazocine from sigma1 receptor in rat liver membranes after 120 mins by liquid scintillation countingMore data for this Ligand-Target Pair
Affinity DataKi: 77nMAssay Description:Displacement of [3H]-(+)-pentazocine from sigma1 receptor in rat liver membranes after 120 mins by liquid scintillation countingMore data for this Ligand-Target Pair
Affinity DataKi: 95nMAssay Description:Displacement of [3H]-(+)-pentazocine from sigma1 receptor in rat liver membranes after 120 mins by liquid scintillation countingMore data for this Ligand-Target Pair
Affinity DataKi: 97nMAssay Description:Displacement of [3H]-(+)-pentazocine from sigma1 receptor in rat liver membranes after 120 mins by liquid scintillation countingMore data for this Ligand-Target Pair
Affinity DataKi: 143nMAssay Description:Displacement of [3H]-(+)-pentazocine from sigma1 receptor in rat liver membranes after 120 mins by liquid scintillation countingMore data for this Ligand-Target Pair
Affinity DataKi: 187nMAssay Description:Displacement of [3H]-(+)-pentazocine from sigma1 receptor in rat liver membranes after 120 mins by liquid scintillation countingMore data for this Ligand-Target Pair
Affinity DataKi: 228nMAssay Description:Displacement of [3H]-(+)-pentazocine from sigma1 receptor in rat liver membranes after 120 mins by liquid scintillation countingMore data for this Ligand-Target Pair
Affinity DataKi: 285nMAssay Description:Displacement of [3H]-(+)-pentazocine from sigma1 receptor in rat liver membranes after 120 mins by liquid scintillation countingMore data for this Ligand-Target Pair
Affinity DataKi: 422nMAssay Description:Displacement of [3H]-DTG from sigma2 receptor in rat liver membranes after 120 mins by liquid scintillation countingMore data for this Ligand-Target Pair
Affinity DataKi: 825nMAssay Description:Displacement of [3H]-DTG from sigma2 receptor in rat liver membranes after 120 mins by liquid scintillation countingMore data for this Ligand-Target Pair
Affinity DataKi: 987nMAssay Description:Displacement of [3H]-DTG from sigma2 receptor in rat liver membranes after 120 mins by liquid scintillation countingMore data for this Ligand-Target Pair
Affinity DataKi: 1.12E+3nMAssay Description:Displacement of [3H]-DTG from sigma2 receptor in rat liver membranes after 120 mins by liquid scintillation countingMore data for this Ligand-Target Pair
Affinity DataKi: 1.26E+3nMAssay Description:Displacement of [3H]-DTG from sigma2 receptor in rat liver membranes after 120 mins by liquid scintillation countingMore data for this Ligand-Target Pair
Affinity DataKi: 1.38E+3nMAssay Description:Displacement of [3H]-DTG from sigma2 receptor in rat liver membranes after 120 mins by liquid scintillation countingMore data for this Ligand-Target Pair
Affinity DataKi: 1.84E+3nMAssay Description:Displacement of [3H]-DTG from sigma2 receptor in rat liver membranes after 120 mins by liquid scintillation countingMore data for this Ligand-Target Pair
Affinity DataKi: 2.34E+3nMAssay Description:Displacement of [3H]-DTG from sigma2 receptor in rat liver membranes after 120 mins by liquid scintillation countingMore data for this Ligand-Target Pair
Affinity DataKi: 2.43E+3nMAssay Description:Displacement of [3H]-DTG from sigma2 receptor in rat liver membranes after 120 mins by liquid scintillation countingMore data for this Ligand-Target Pair
Affinity DataKi: 2.58E+3nMAssay Description:Displacement of [3H]-DTG from sigma2 receptor in rat liver membranes after 120 mins by liquid scintillation countingMore data for this Ligand-Target Pair
Affinity DataKi: 3.20E+3nMAssay Description:Displacement of [3H]-DTG from sigma2 receptor in rat liver membranes after 120 mins by liquid scintillation countingMore data for this Ligand-Target Pair