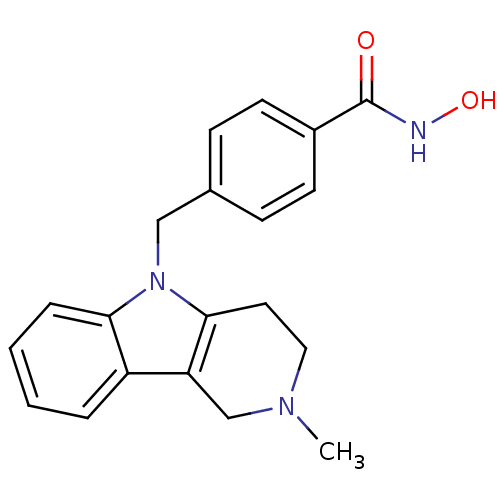
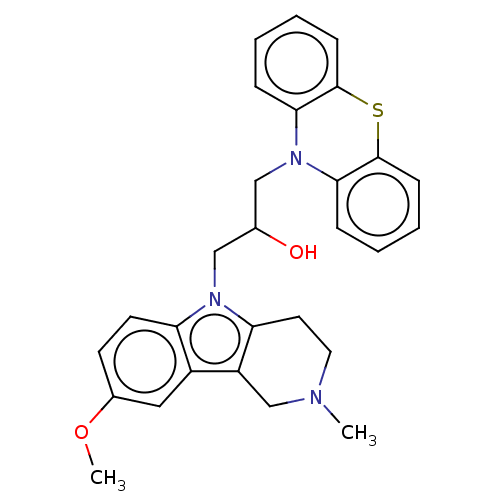
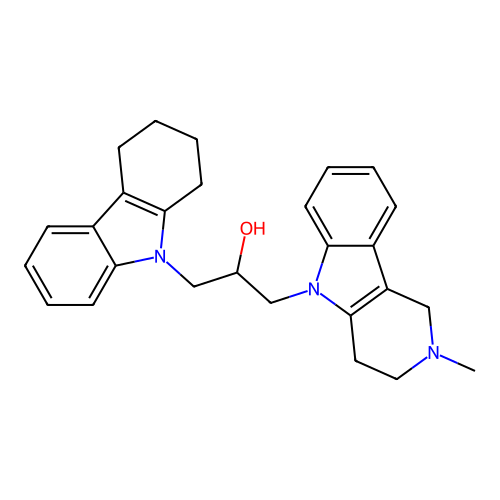
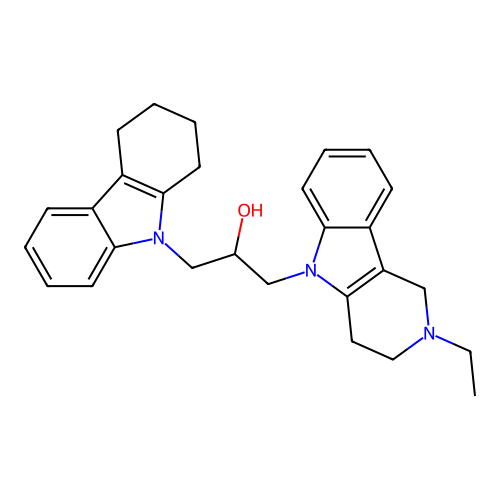
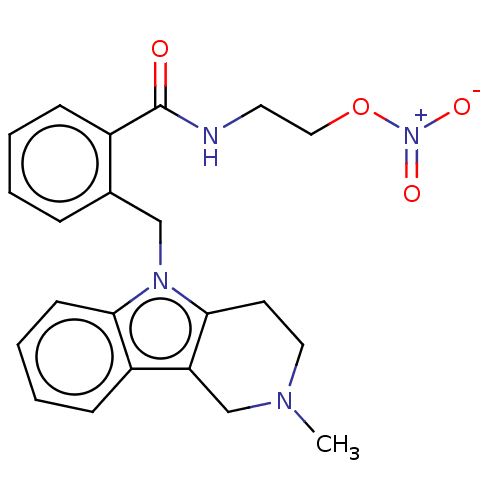
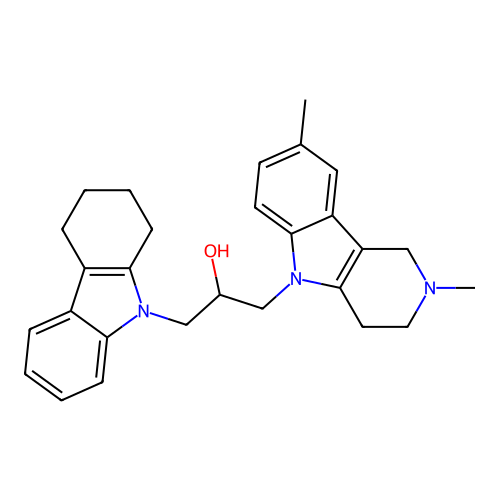
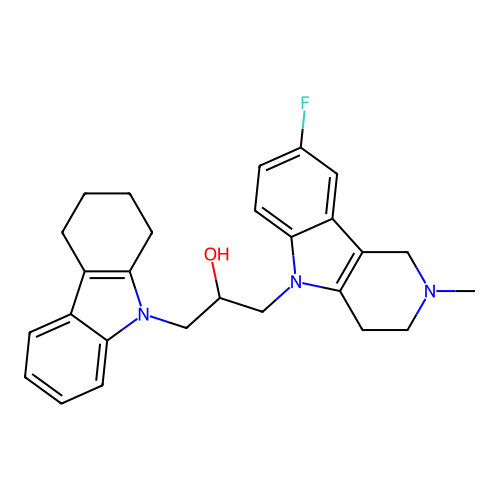
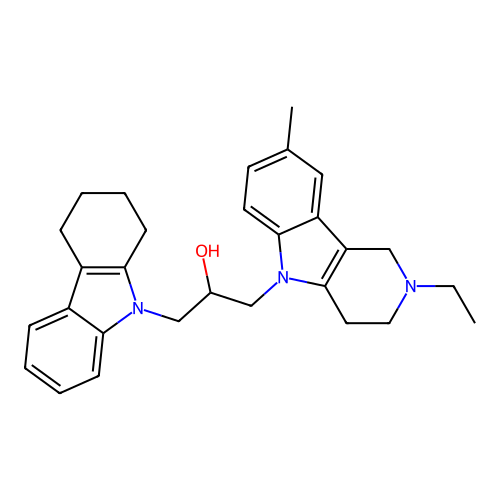
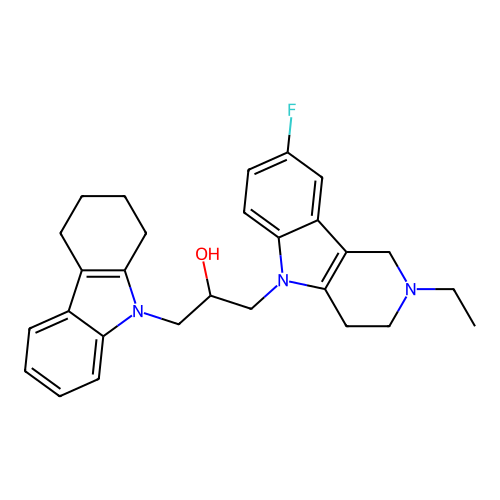
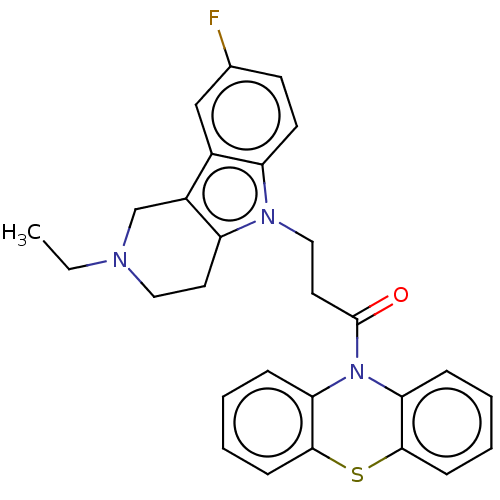
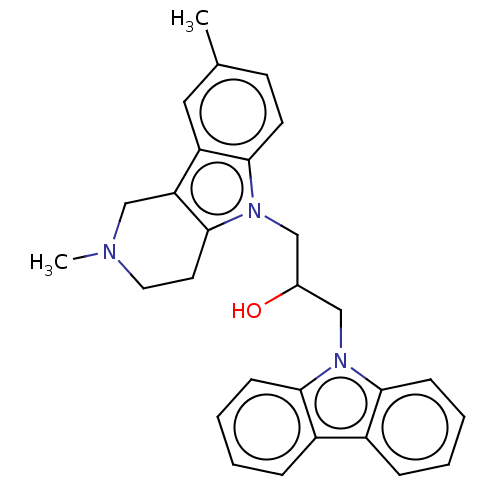
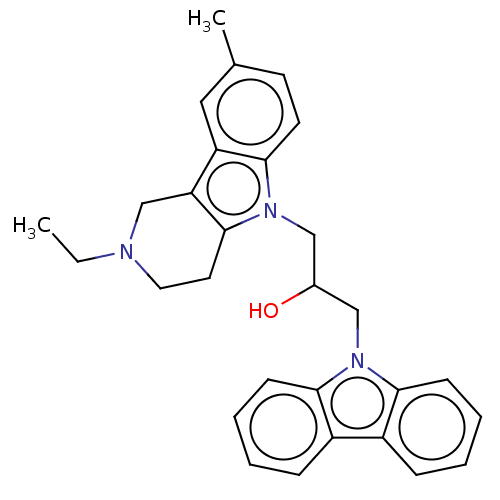
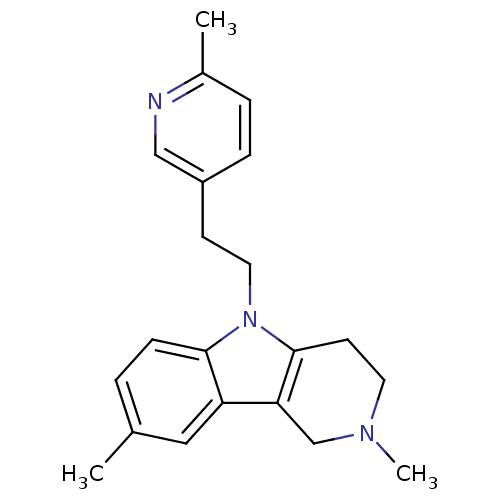
Report error Found 22 Enz. Inhib. hit(s) with all data for entry = 50018910
Affinity DataIC50: 15nMAssay Description:Inhibition of HDAC6 (unknown origin)More data for this Ligand-Target Pair
Affinity DataIC50: 390nMAssay Description:Inhibition of BChE (unknown origin)More data for this Ligand-Target Pair
Affinity DataIC50: 1.06E+3nMAssay Description:Inhibition of BChE (unknown origin)More data for this Ligand-Target Pair
Affinity DataIC50: 2.77E+3nMAssay Description:Inhibition of BChE (unknown origin)More data for this Ligand-Target Pair
Affinity DataIC50: 3.68E+3nMAssay Description:Inhibition of horse serum BChE incubated for 10 mins by Ellman's methodMore data for this Ligand-Target Pair
Affinity DataIC50: 3.90E+3nMAssay Description:Inhibition of BChE (unknown origin)More data for this Ligand-Target Pair
Affinity DataIC50: 5.03E+3nMAssay Description:Inhibition of BChE (unknown origin)More data for this Ligand-Target Pair
Affinity DataIC50: 5.23E+3nMAssay Description:Inhibition of BChE (unknown origin)More data for this Ligand-Target Pair
Affinity DataIC50: 5.83E+3nMAssay Description:Inhibition of BChE (unknown origin)More data for this Ligand-Target Pair
Affinity DataIC50: 8.80E+3nMAssay Description:Displacement of [3H]ifenprodyl from NMDA receptor NR2B subunit (unknown origin)More data for this Ligand-Target Pair
Affinity DataIC50: 1.17E+4nMAssay Description:Displacement of [3H]ifenprodyl from NMDA receptor (unknown origin)More data for this Ligand-Target Pair
Affinity DataIC50: 1.25E+4nMAssay Description:Displacement of [3H]ifenprodyl from NMDA receptor (unknown origin)More data for this Ligand-Target Pair
Affinity DataIC50: 1.32E+4nMAssay Description:Displacement of [3H]MK801 from NMDA receptor NR2B subunit (unknown origin)More data for this Ligand-Target Pair
Affinity DataIC50: 1.34E+4nMAssay Description:Displacement of [3H]ifenprodyl from NMDA receptor NR2B subunit (unknown origin)More data for this Ligand-Target Pair
Affinity DataIC50: 1.46E+4nMAssay Description:Displacement of [3H]MK801 from NMDA receptor NR2B subunit (unknown origin)More data for this Ligand-Target Pair
Affinity DataIC50: 1.48E+4nMAssay Description:Inhibition of human erythrocyte AChE incubated for 10 mins by Ellman's methodMore data for this Ligand-Target Pair
Affinity DataIC50: 1.54E+4nMAssay Description:Displacement of [3H]ifenprodyl from NMDA receptor NR2B subunit (unknown origin)More data for this Ligand-Target Pair
Affinity DataIC50: 1.58E+4nMAssay Description:Displacement of [3H]MK801 from NMDA receptor NR2B subunit (unknown origin)More data for this Ligand-Target Pair
Affinity DataIC50: 1.64E+4nMAssay Description:Inhibition of HDAC1 (unknown origin)More data for this Ligand-Target Pair
Affinity DataIC50: 3.26E+4nMAssay Description:Displacement of [3H]ifenprodyl from NMDA receptor NR2B subunit (unknown origin)More data for this Ligand-Target Pair
Affinity DataIC50: 3.95E+4nMAssay Description:Displacement of [3H]ifenprodyl from NMDA receptor NR2B subunit (unknown origin)More data for this Ligand-Target Pair
Affinity DataIC50: 8.24E+4nMAssay Description:Displacement of [3H]ifenprodyl from NMDA receptor NR2B subunit (unknown origin)More data for this Ligand-Target Pair