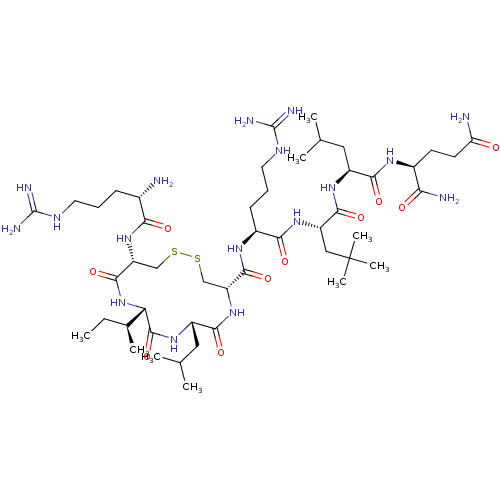
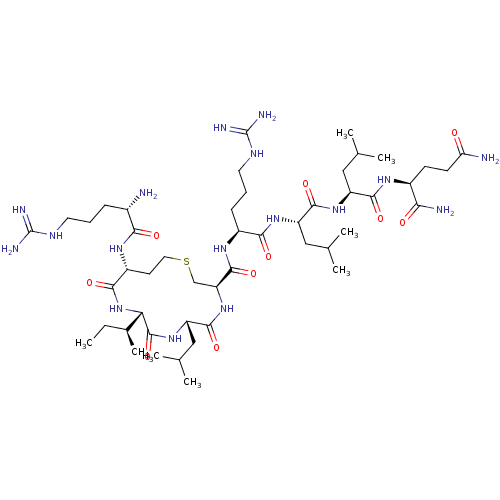
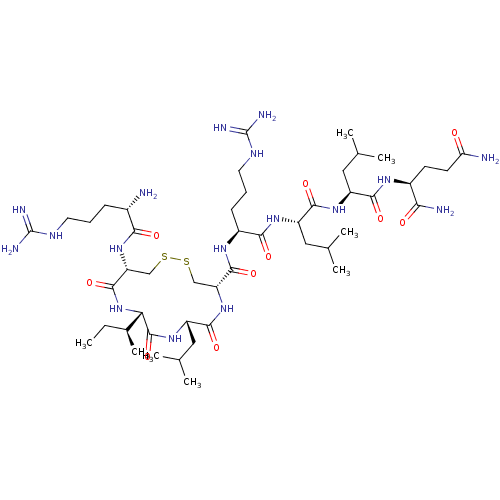
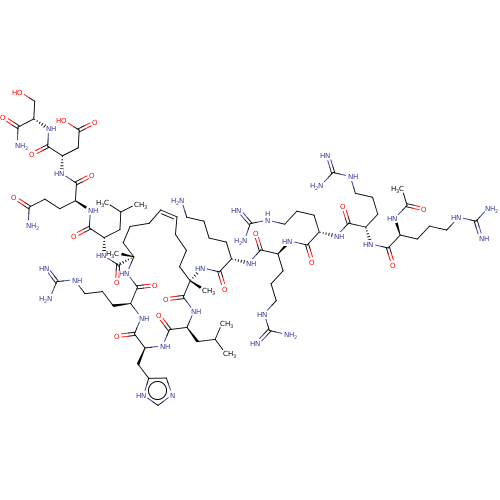
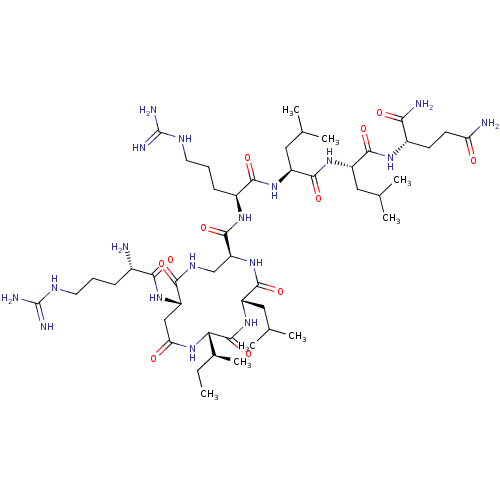
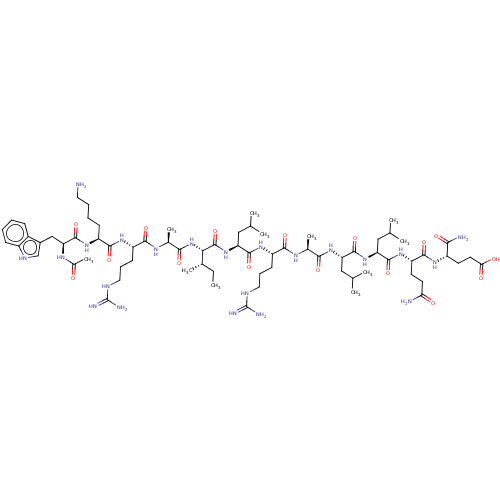
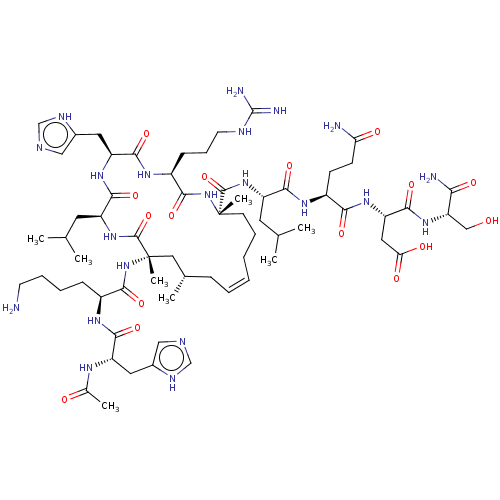
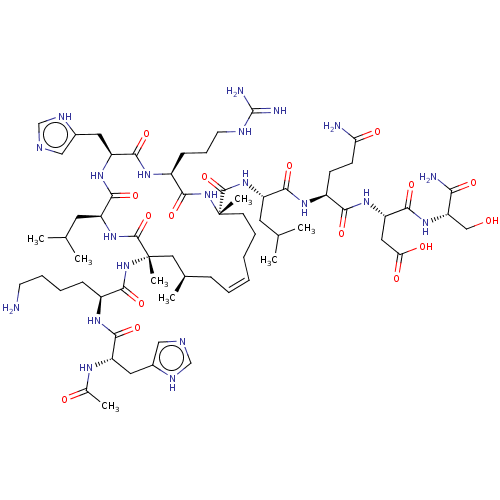
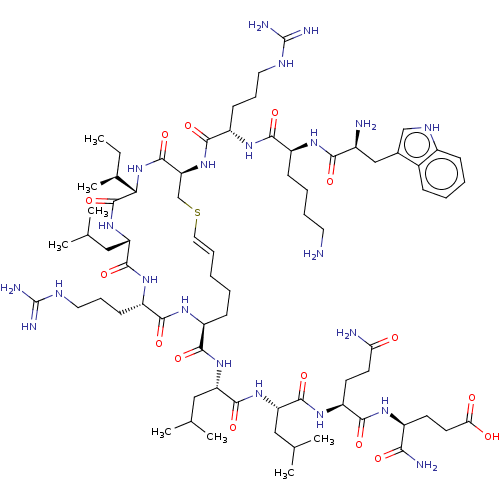
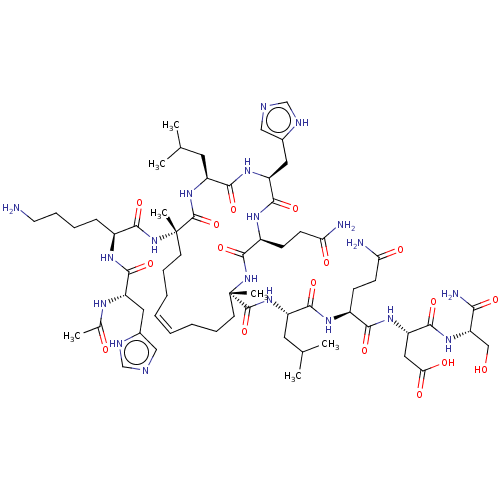
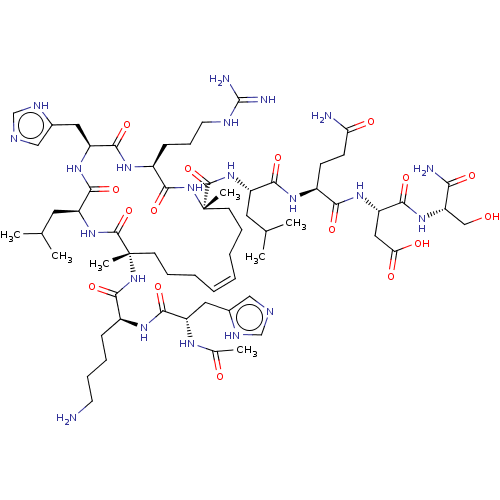
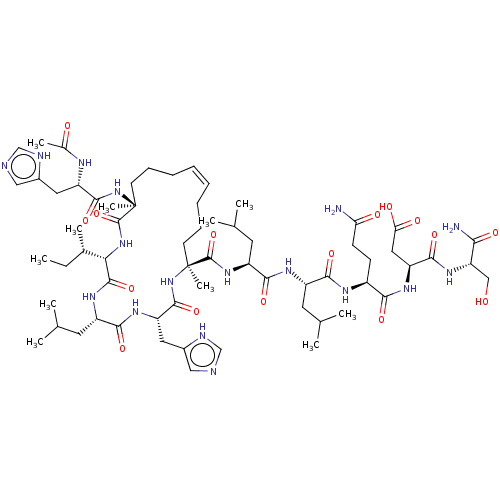
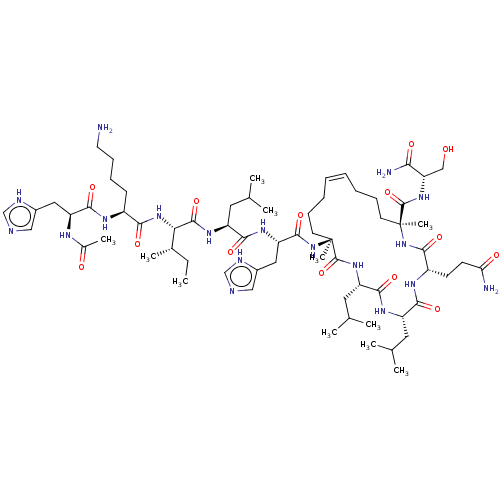
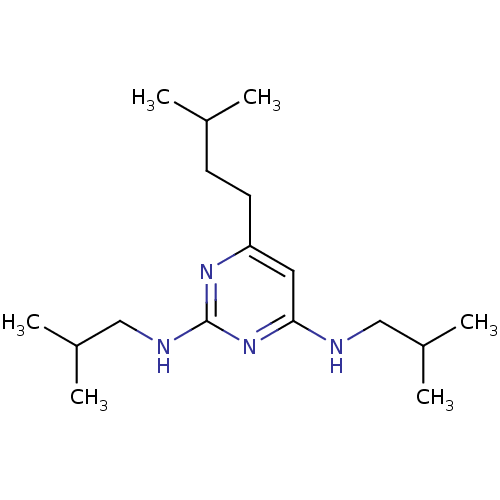
Report error Found 43 Enz. Inhib. hit(s) with all data for entry = 50002786
Affinity DataKi: 0.0700nMAssay Description:Antagonist activity at full length recombinant human ERalpha expressed in baculovirus expression system assessed as inhibition of estradiol-induced E...More data for this Ligand-Target Pair
Affinity DataKi: 1.20nMAssay Description:Antagonist activity at full length recombinant human ERbeta expressed in baculovirus expression system assessed as inhibition of estradiol-induced Eu...More data for this Ligand-Target Pair
Affinity DataKi: 6.90nMAssay Description:Antagonist activity at full length recombinant human ERalpha expressed in baculovirus expression system assessed as inhibition of estradiol-induced E...More data for this Ligand-Target Pair
Affinity DataKi: 11nMAssay Description:Antagonist activity at full length recombinant human ERalpha expressed in baculovirus expression system assessed as inhibition of estradiol-induced E...More data for this Ligand-Target Pair
Affinity DataKd: 19nMAssay Description:Binding affinity to recombinant N-terminal His6-tagged ERalpha LBD (unknown origin) (299 to 554 residues) by surface plasmon resonance methodMore data for this Ligand-Target Pair
Affinity DataKi: 25nMAssay Description:Displacement of FAM-LTERHKILHRLLQEGSPSD from ERbeta (unknown origin) (260 to 502 residues) expressed in Escherichia coli rosetta (DE3) after 1 hr by ...More data for this Ligand-Target Pair
Affinity DataKi: 25nMAssay Description:Antagonist activity at full length recombinant human ERalpha expressed in baculovirus expression system assessed as inhibition of estradiol-induced E...More data for this Ligand-Target Pair
Affinity DataKd: 30nMAssay Description:Binding affinity to recombinant human ERalpha LBD (301 to 553 residues) expressed in Escherichia coli BL21(DE3) using FITC-labelled compound after 1 ...More data for this Ligand-Target Pair
Affinity DataKi: 64nMAssay Description:Antagonist activity at full length recombinant human ERbeta expressed in baculovirus expression system assessed as inhibition of estradiol-induced Eu...More data for this Ligand-Target Pair
Affinity DataKd: 75nMAssay Description:Binding affinity to recombinant human ERalpha LBD (301 to 553 residues) expressed in Escherichia coli BL21(DE3) by surface plasmon resonance methodMore data for this Ligand-Target Pair
Affinity DataKi: 77nMAssay Description:Antagonist activity at full length recombinant human ERbeta expressed in baculovirus expression system assessed as inhibition of estradiol-induced Eu...More data for this Ligand-Target Pair
Affinity DataKd: 85nMAssay Description:Binding affinity to recombinant human ERalpha LBD (301 to 553 residues) expressed in Escherichia coli BL21(DE3) using FITC-labelled compound after 1 ...More data for this Ligand-Target Pair
Affinity DataKd: 89nMAssay Description:Binding affinity to recombinant human ERalpha LBD (301 to 553 residues) expressed in Escherichia coli BL21(DE3) using FAM-labelled compound after 1 h...More data for this Ligand-Target Pair
Affinity DataIC50: 89nMAssay Description:Displacement of 5-iodoacetamidofluorescein-labelled SRC3 from C417-biotin labelled ERalpha (unknown origin) (304 to 554 residues) after 45 mins by TR...More data for this Ligand-Target Pair
Affinity DataIC50: 96nMAssay Description:Displacement of 5-iodoacetamidofluorescein-labelled SRC3 from C417-biotin labelled ERalpha (unknown origin) (304 to 554 residues) after 45 mins by TR...More data for this Ligand-Target Pair
Affinity DataKd: 101nMAssay Description:Binding affinity to recombinant human ERalpha LBD (301 to 553 residues) expressed in Escherichia coli BL21(DE3) using FAM-labelled compound after 1 h...More data for this Ligand-Target Pair
Affinity DataKd: 155nMAssay Description:Binding affinity to recombinant human ERbeta LBD (259 to 498 residues) C334S/C369S/C481S triple mutant expressed in Escherichia coli BL21(DE3) by sur...More data for this Ligand-Target Pair
Affinity DataKd: 178nMAssay Description:Binding affinity to recombinant human ERbeta LBD (259 to 498 residues) C334S/C369S/C481S triple mutant expressed in Escherichia coli BL21(DE3) using ...More data for this Ligand-Target Pair
Affinity DataKd: 179nMAssay Description:Binding affinity to recombinant human ERbeta LBD (259 to 498 residues) C334S/C369S/C481S triple mutant expressed in Escherichia coli BL21(DE3) using ...More data for this Ligand-Target Pair
Affinity DataKi: 185nMAssay Description:Displacement of FAM-LTERHKILHRLLQEGSPSD from ERalpha (unknown origin) (302 to 552 residues) expressed in Escherichia coli rosetta (DE3) after 1 hr by...More data for this Ligand-Target Pair
Affinity DataKd: 352nMAssay Description:Binding affinity to recombinant human ERalpha LBD (301 to 553 residues) expressed in Escherichia coli BL21(DE3) by surface plasmon resonance methodMore data for this Ligand-Target Pair
Affinity DataKi: 390nMAssay Description:Antagonist activity at full length recombinant human ERbeta expressed in baculovirus expression system assessed as inhibition of estradiol-induced Eu...More data for this Ligand-Target Pair
Affinity DataIC50: 390nMAssay Description:Displacement of 5-iodoacetamidofluorescein-labelled SRC3 from C417-biotin labelled ERalpha (unknown origin) (304 to 554 residues) after 45 mins by TR...More data for this Ligand-Target Pair
Affinity DataKd: 632nMAssay Description:Binding affinity to recombinant human ERbeta LBD (259 to 498 residues) C334S/C369S/C481S triple mutant expressed in Escherichia coli BL21(DE3) by sur...More data for this Ligand-Target Pair
Affinity DataKd: 674nMAssay Description:Binding affinity to recombinant human ERalpha LBD (301 to 553 residues) expressed in Escherichia coli BL21(DE3) by surface plasmon resonance methodMore data for this Ligand-Target Pair
Affinity DataIC50: 760nMAssay Description:Displacement of 5-iodoacetamidofluorescein-labelled SRC3 from C417-biotin labelled ERalpha (unknown origin) (304 to 554 residues) after 45 mins by TR...More data for this Ligand-Target Pair
Affinity DataKd: 900nMAssay Description:Displacement of THC-ketone from human ERalpha LBD (304 to 554 residues) expressed in Escherichia coli BL21(DE3) after 1 hr by fluorescence assayMore data for this Ligand-Target Pair
Affinity DataIC50: 1.10E+3nMAssay Description:Displacement of 5-iodoacetamidofluorescein-labelled SRC3 from C417-biotin labelled ERalpha (unknown origin) (304 to 554 residues) after 45 mins by TR...More data for this Ligand-Target Pair
Affinity DataIC50: 1.20E+3nMAssay Description:Displacement of 5-iodoacetamidofluorescein-labelled SRC3 from C417-biotin labelled ERalpha (unknown origin) (304 to 554 residues) after 45 mins by TR...More data for this Ligand-Target Pair
Affinity DataKi: 1.54E+3nMAssay Description:Displacement of FAM-LTERHKILHRLLQEGSPSD from ERalpha (unknown origin) (302 to 552 residues) expressed in Escherichia coli rosetta (DE3) after 1 hr by...More data for this Ligand-Target Pair
Affinity DataIC50: 1.80E+3nMAssay Description:Displacement of 5-iodoacetamidofluorescein-labelled SRC3 from C417-biotin labelled ERalpha (unknown origin) (304 to 554 residues) after 45 mins by TR...More data for this Ligand-Target Pair
Affinity DataKd: 1.99E+3nMAssay Description:Binding affinity to recombinant human ERbeta LBD (259 to 498 residues) C334S/C369S/C481S triple mutant expressed in Escherichia coli BL21(DE3) by sur...More data for this Ligand-Target Pair
Affinity DataKd: 2.50E+3nMAssay Description:Binding affinity to recombinant human ERalpha LBD (301 to 553 residues) expressed in Escherichia coli BL21(DE3) by surface plasmon resonance methodMore data for this Ligand-Target Pair
Affinity DataKd: 2.60E+3nMAssay Description:Binding affinity to recombinant N-terminal His6-tagged ERalpha LBD (unknown origin) (299 to 554 residues) by surface plasmon resonance methodMore data for this Ligand-Target Pair
Affinity DataKd: 2.80E+3nMAssay Description:Binding affinity to recombinant human ERbeta LBD (259 to 498 residues) C334S/C369S/C481S triple mutant expressed in Escherichia coli BL21(DE3) by sur...More data for this Ligand-Target Pair
Affinity DataKi: 3.64E+3nMAssay Description:Displacement of FAM-LTERHKILHRLLQEGSPSD from ERbeta (unknown origin) (260 to 502 residues) expressed in Escherichia coli rosetta (DE3) after 1 hr by ...More data for this Ligand-Target Pair
Affinity DataKd: 4.00E+3nMAssay Description:Binding affinity to recombinant human ERalpha LBD (301 to 553 residues) expressed in Escherichia coli BL21(DE3) by surface plasmon resonance methodMore data for this Ligand-Target Pair
Affinity DataKd: 5.00E+3nMAssay Description:Binding affinity to recombinant human ERbeta LBD (259 to 498 residues) C334S/C369S/C481S triple mutant expressed in Escherichia coli BL21(DE3) by sur...More data for this Ligand-Target Pair
Affinity DataKd: 8.00E+3nMAssay Description:Binding affinity to recombinant human ERalpha LBD (301 to 553 residues) expressed in Escherichia coli BL21(DE3) by surface plasmon resonance methodMore data for this Ligand-Target Pair
Affinity DataKd: 8.30E+3nMAssay Description:Binding affinity to recombinant human ERbeta LBD (259 to 498 residues) C334S/C369S/C481S triple mutant expressed in Escherichia coli BL21(DE3) by sur...More data for this Ligand-Target Pair
Affinity DataKd: >1.50E+4nMAssay Description:Binding affinity to recombinant human ERalpha LBD (301 to 553 residues) expressed in Escherichia coli BL21(DE3) by surface plasmon resonance methodMore data for this Ligand-Target Pair
Affinity DataKd: >1.50E+4nMAssay Description:Binding affinity to recombinant human ERbeta LBD (259 to 498 residues) C334S/C369S/C481S triple mutant expressed in Escherichia coli BL21(DE3) by sur...More data for this Ligand-Target Pair
Affinity DataKi: 3.00E+4nMAssay Description:Displacement of TMR-labelled ILRKLLQE peptide from His6-tagged ERalpha LBD (unknown origin) (304 to 554 residues) expressed in Escherichia coli BL21(...More data for this Ligand-Target Pair