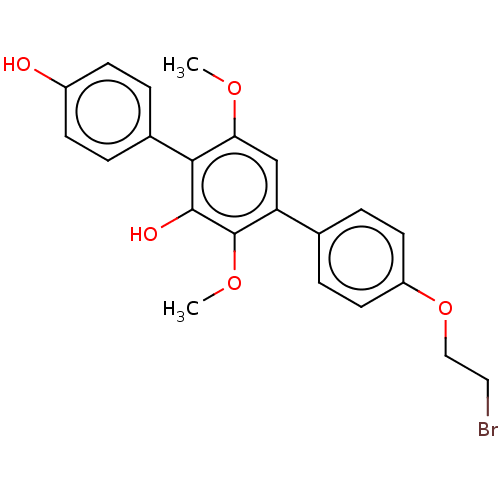
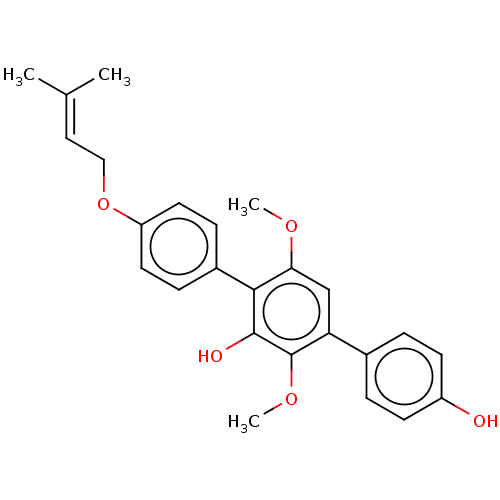
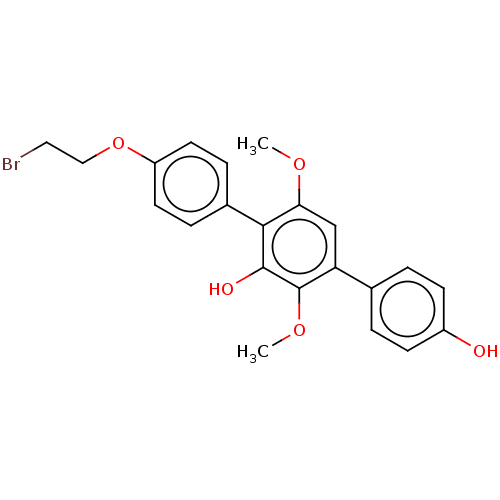
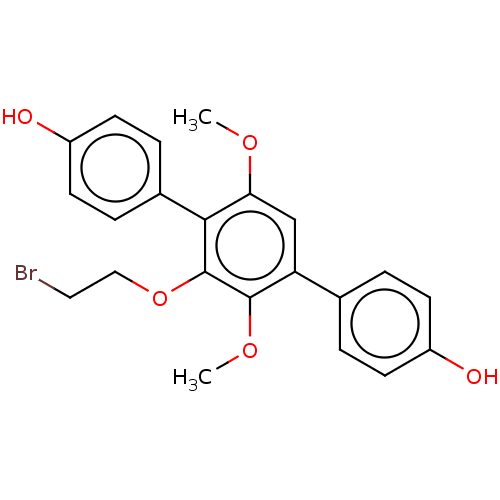
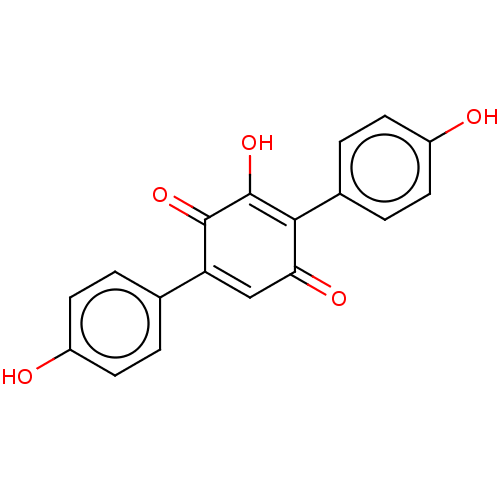
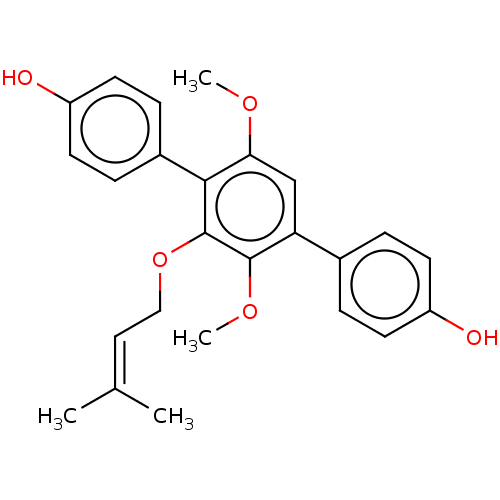
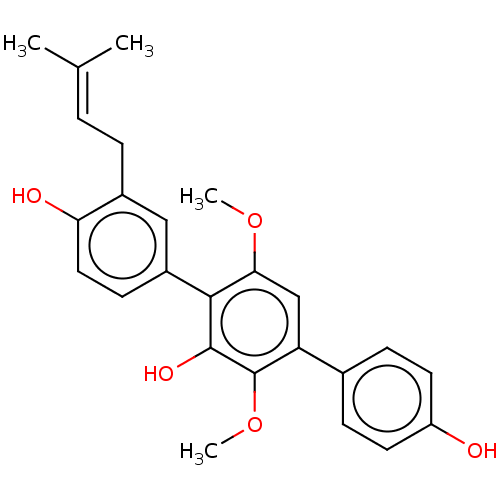
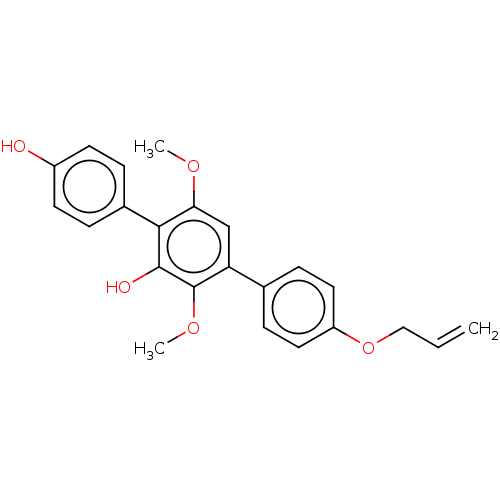
Report error Found 38 Enz. Inhib. hit(s) with all data for entry = 50002377
Affinity DataKi: 1.50E+3nMAssay Description:Non-competitive inhibition of human alpha-glucosidase expressed in Saccharomyces cerevisiae using p-nitrophenyl-alpha-D-glucopyranoside as substrate ...More data for this Ligand-Target Pair
Affinity DataKi: 3.45E+3nMAssay Description:Non-competitive inhibition of human alpha-glucosidase expressed in Saccharomyces cerevisiae using p-nitrophenyl-alpha-D-glucopyranoside as substrate ...More data for this Ligand-Target Pair
Affinity DataIC50: 4.79E+3nMAssay Description:Inhibition of human alpha-glucosidase expressed in Saccharomyces cerevisiae using p-nitrophenyl-alpha-D-glucopyranoside as substrate after 15 minsMore data for this Ligand-Target Pair
Affinity DataIC50: 6.67E+3nMAssay Description:Inhibition of human alpha-glucosidase expressed in Saccharomyces cerevisiae using p-nitrophenyl-alpha-D-glucopyranoside as substrate after 15 minsMore data for this Ligand-Target Pair
Affinity DataIC50: 7.14E+3nMAssay Description:Inhibition of human alpha-glucosidase expressed in Saccharomyces cerevisiae using p-nitrophenyl-alpha-D-glucopyranoside as substrate after 15 minsMore data for this Ligand-Target Pair
Affinity DataIC50: 8.58E+3nMAssay Description:Inhibition of human alpha-glucosidase expressed in Saccharomyces cerevisiae using p-nitrophenyl-alpha-D-glucopyranoside as substrate after 15 minsMore data for this Ligand-Target Pair
Affinity DataIC50: 8.69E+3nMAssay Description:Inhibition of human alpha-glucosidase expressed in Saccharomyces cerevisiae using p-nitrophenyl-alpha-D-glucopyranoside as substrate after 15 minsMore data for this Ligand-Target Pair
Affinity DataIC50: 8.77E+3nMAssay Description:Inhibition of human alpha-glucosidase expressed in Saccharomyces cerevisiae using p-nitrophenyl-alpha-D-glucopyranoside as substrate after 15 minsMore data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMAssay Description:Inhibition of human alpha-glucosidase expressed in Saccharomyces cerevisiae using p-nitrophenyl-alpha-D-glucopyranoside as substrate after 15 minsMore data for this Ligand-Target Pair
Affinity DataIC50: 1.07E+4nMAssay Description:Inhibition of human alpha-glucosidase expressed in Saccharomyces cerevisiae using p-nitrophenyl-alpha-D-glucopyranoside as substrate after 15 minsMore data for this Ligand-Target Pair
Affinity DataIC50: 1.13E+4nMAssay Description:Inhibition of human alpha-glucosidase expressed in Saccharomyces cerevisiae using p-nitrophenyl-alpha-D-glucopyranoside as substrate after 15 minsMore data for this Ligand-Target Pair
Affinity DataIC50: 1.22E+4nMAssay Description:Inhibition of human alpha-glucosidase expressed in Saccharomyces cerevisiae using p-nitrophenyl-alpha-D-glucopyranoside as substrate after 15 minsMore data for this Ligand-Target Pair
Affinity DataIC50: 1.23E+4nMAssay Description:Inhibition of human alpha-glucosidase expressed in Saccharomyces cerevisiae using p-nitrophenyl-alpha-D-glucopyranoside as substrate after 15 minsMore data for this Ligand-Target Pair
Affinity DataIC50: 1.25E+4nMAssay Description:Inhibition of human alpha-glucosidase expressed in Saccharomyces cerevisiae using p-nitrophenyl-alpha-D-glucopyranoside as substrate after 15 minsMore data for this Ligand-Target Pair
Affinity DataIC50: 1.25E+4nMAssay Description:Inhibition of human alpha-glucosidase expressed in Saccharomyces cerevisiae using p-nitrophenyl-alpha-D-glucopyranoside as substrate after 15 minsMore data for this Ligand-Target Pair
Affinity DataIC50: 1.32E+4nMAssay Description:Inhibition of human alpha-glucosidase expressed in Saccharomyces cerevisiae using p-nitrophenyl-alpha-D-glucopyranoside as substrate after 15 minsMore data for this Ligand-Target Pair
Affinity DataIC50: 1.40E+4nMAssay Description:Inhibition of human alpha-glucosidase expressed in Saccharomyces cerevisiae using p-nitrophenyl-alpha-D-glucopyranoside as substrate after 15 minsMore data for this Ligand-Target Pair
Affinity DataIC50: 1.43E+4nMAssay Description:Inhibition of human alpha-glucosidase expressed in Saccharomyces cerevisiae using p-nitrophenyl-alpha-D-glucopyranoside as substrate after 15 minsMore data for this Ligand-Target Pair
Affinity DataIC50: 1.61E+4nMAssay Description:Inhibition of human alpha-glucosidase expressed in Saccharomyces cerevisiae using p-nitrophenyl-alpha-D-glucopyranoside as substrate after 15 minsMore data for this Ligand-Target Pair
Affinity DataIC50: 1.75E+4nMAssay Description:Inhibition of human alpha-glucosidase expressed in Saccharomyces cerevisiae using p-nitrophenyl-alpha-D-glucopyranoside as substrate after 15 minsMore data for this Ligand-Target Pair
Affinity DataIC50: 1.80E+4nMAssay Description:Inhibition of human alpha-glucosidase expressed in Saccharomyces cerevisiae using p-nitrophenyl-alpha-D-glucopyranoside as substrate after 15 minsMore data for this Ligand-Target Pair
Affinity DataIC50: 1.88E+4nMAssay Description:Inhibition of human alpha-glucosidase expressed in Saccharomyces cerevisiae using p-nitrophenyl-alpha-D-glucopyranoside as substrate after 15 minsMore data for this Ligand-Target Pair
Affinity DataIC50: 1.91E+4nMAssay Description:Inhibition of human alpha-glucosidase expressed in Saccharomyces cerevisiae using p-nitrophenyl-alpha-D-glucopyranoside as substrate after 15 minsMore data for this Ligand-Target Pair
Affinity DataIC50: 1.93E+4nMAssay Description:Inhibition of human alpha-glucosidase expressed in Saccharomyces cerevisiae using p-nitrophenyl-alpha-D-glucopyranoside as substrate after 15 minsMore data for this Ligand-Target Pair
Affinity DataIC50: 2.01E+4nMAssay Description:Inhibition of human alpha-glucosidase expressed in Saccharomyces cerevisiae using p-nitrophenyl-alpha-D-glucopyranoside as substrate after 15 minsMore data for this Ligand-Target Pair
Affinity DataIC50: 2.01E+4nMAssay Description:Inhibition of human alpha-glucosidase expressed in Saccharomyces cerevisiae using p-nitrophenyl-alpha-D-glucopyranoside as substrate after 15 minsMore data for this Ligand-Target Pair
Affinity DataIC50: 2.27E+4nMAssay Description:Inhibition of human alpha-glucosidase expressed in Saccharomyces cerevisiae using p-nitrophenyl-alpha-D-glucopyranoside as substrate after 15 minsMore data for this Ligand-Target Pair
Affinity DataIC50: 2.34E+4nMAssay Description:Inhibition of human alpha-glucosidase expressed in Saccharomyces cerevisiae using p-nitrophenyl-alpha-D-glucopyranoside as substrate after 15 minsMore data for this Ligand-Target Pair
Affinity DataIC50: 2.57E+4nMAssay Description:Inhibition of human alpha-glucosidase expressed in Saccharomyces cerevisiae using p-nitrophenyl-alpha-D-glucopyranoside as substrate after 15 minsMore data for this Ligand-Target Pair
Affinity DataIC50: 2.65E+4nMAssay Description:Inhibition of human alpha-glucosidase expressed in Saccharomyces cerevisiae using p-nitrophenyl-alpha-D-glucopyranoside as substrate after 15 minsMore data for this Ligand-Target Pair
Affinity DataIC50: 3.13E+4nMAssay Description:Inhibition of human alpha-glucosidase expressed in Saccharomyces cerevisiae using p-nitrophenyl-alpha-D-glucopyranoside as substrate after 15 minsMore data for this Ligand-Target Pair
Affinity DataIC50: 3.22E+4nMAssay Description:Inhibition of human alpha-glucosidase expressed in Saccharomyces cerevisiae using p-nitrophenyl-alpha-D-glucopyranoside as substrate after 15 minsMore data for this Ligand-Target Pair
Affinity DataIC50: 3.54E+4nMAssay Description:Inhibition of human alpha-glucosidase expressed in Saccharomyces cerevisiae using p-nitrophenyl-alpha-D-glucopyranoside as substrate after 15 minsMore data for this Ligand-Target Pair
Affinity DataIC50: 5.31E+4nMAssay Description:Inhibition of human alpha-glucosidase expressed in Saccharomyces cerevisiae using p-nitrophenyl-alpha-D-glucopyranoside as substrate after 15 minsMore data for this Ligand-Target Pair
Affinity DataIC50: 8.38E+4nMAssay Description:Inhibition of human alpha-glucosidase expressed in Saccharomyces cerevisiae using p-nitrophenyl-alpha-D-glucopyranoside as substrate after 15 minsMore data for this Ligand-Target Pair
Affinity DataIC50: 1.08E+5nMAssay Description:Inhibition of human alpha-glucosidase expressed in Saccharomyces cerevisiae using p-nitrophenyl-alpha-D-glucopyranoside as substrate after 15 minsMore data for this Ligand-Target Pair
Affinity DataIC50: 1.92E+5nMAssay Description:Inhibition of human alpha-glucosidase expressed in Saccharomyces cerevisiae using p-nitrophenyl-alpha-D-glucopyranoside as substrate after 15 minsMore data for this Ligand-Target Pair
Affinity DataIC50: 7.08E+5nMAssay Description:Inhibition of human alpha-glucosidase expressed in Saccharomyces cerevisiae using p-nitrophenyl-alpha-D-glucopyranoside as substrate after 15 minsMore data for this Ligand-Target Pair