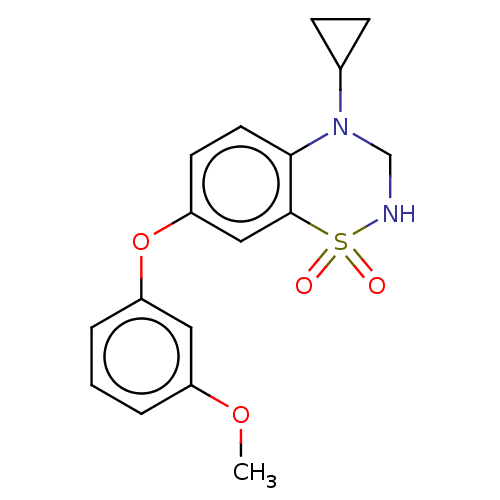
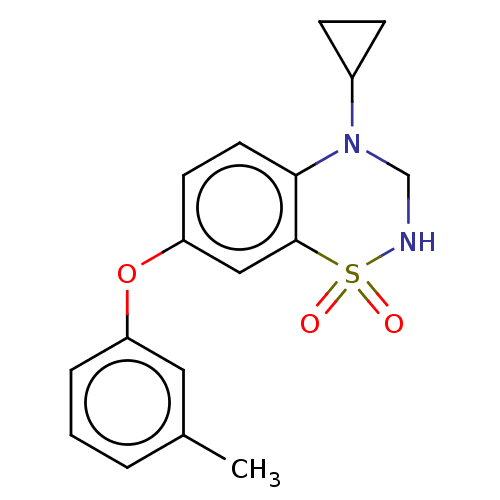
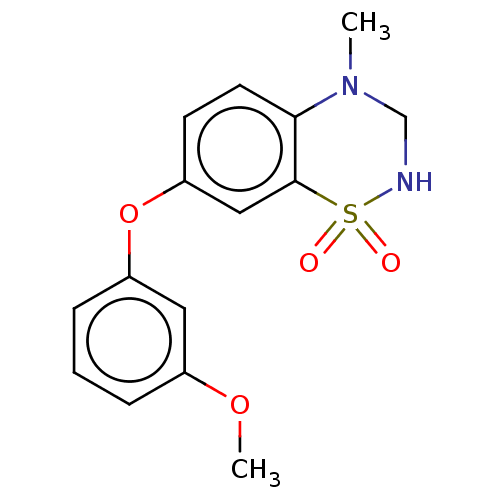
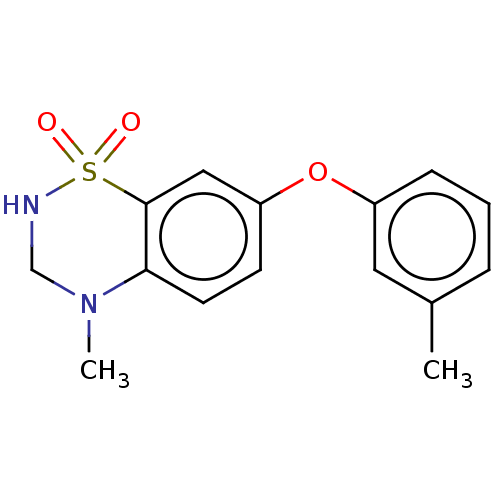
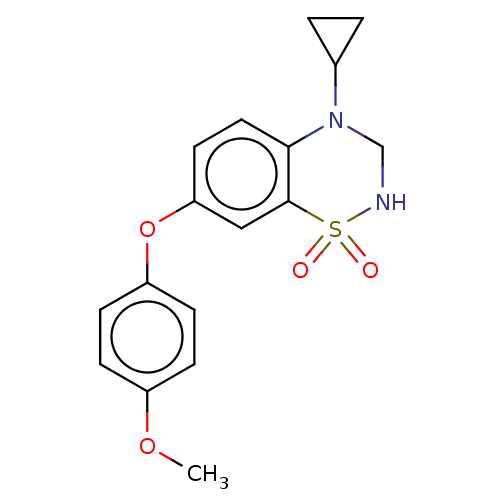
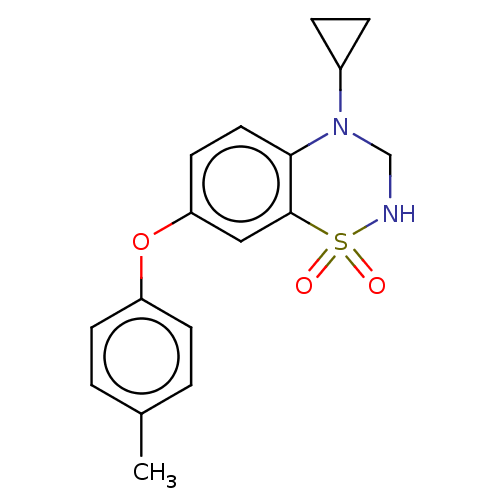
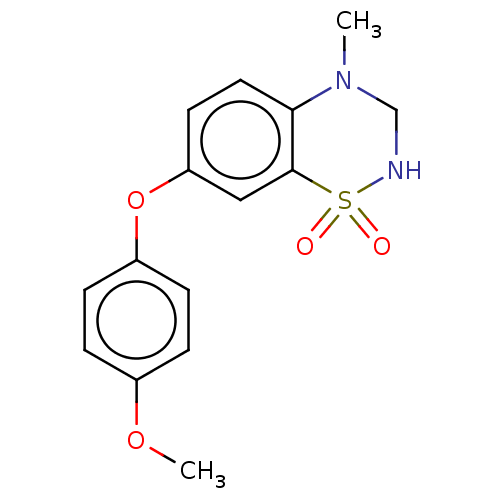
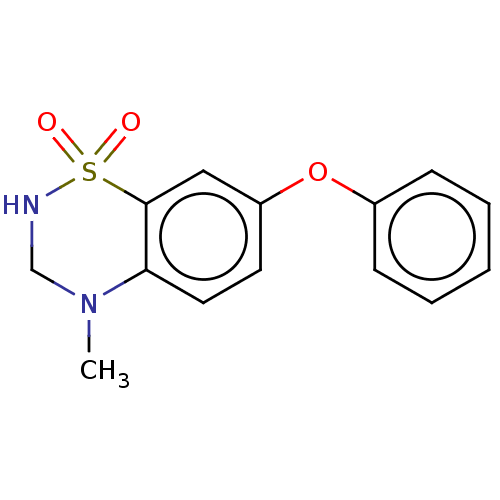
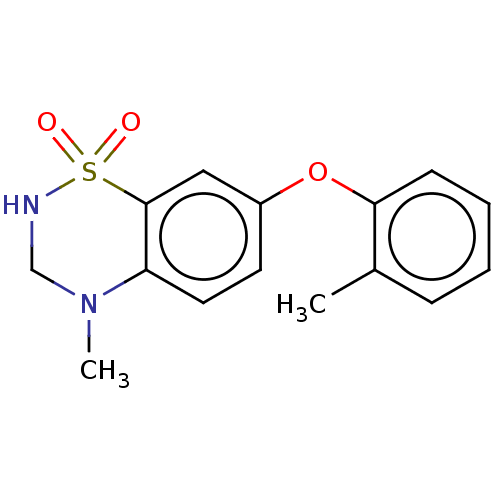
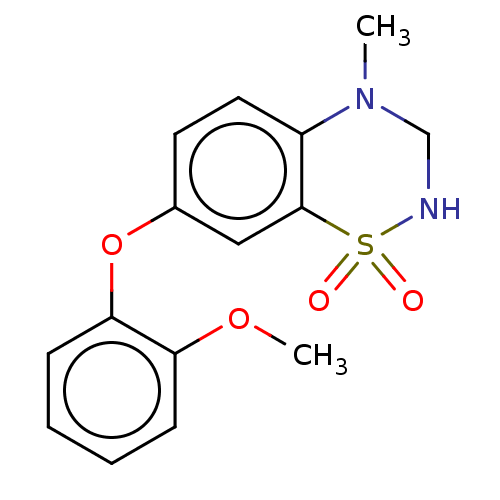
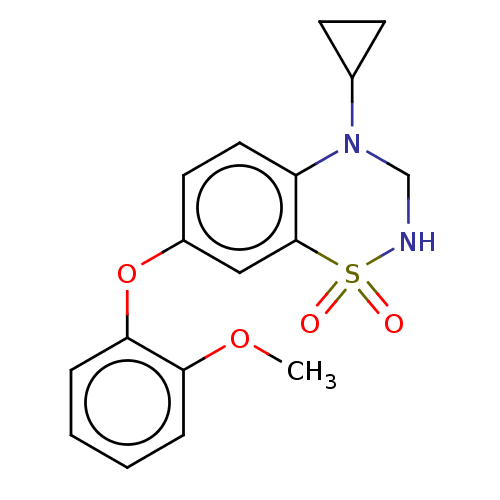
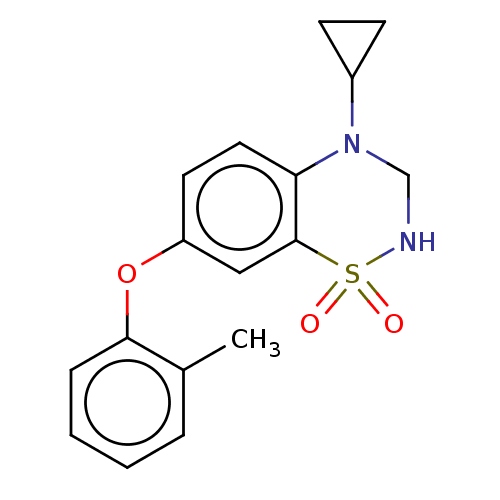
Report error Found 38 Enz. Inhib. hit(s) with all data for entry = 50000481
Affinity DataEC50: 2nMAssay Description:Positive allosteric modulation of GluA2 (Q) flop isoform (unknown origin) expressed in HEK293 cells assessed as potentiation of glutamate-evoked calc...More data for this Ligand-Target Pair
Affinity DataEC50: 2nMAssay Description:Positive allosteric modulation of GluA2 (Q) flop isoform (unknown origin) expressed in HEK293 cells assessed as potentiation of glutamate-evoked calc...More data for this Ligand-Target Pair
Affinity DataEC50: 46nMAssay Description:Positive allosteric modulation of GluA2 (Q) flop isoform (unknown origin) expressed in HEK293 cells assessed as potentiation of glutamate-evoked calc...More data for this Ligand-Target Pair
Affinity DataEC50: 50nMAssay Description:Positive allosteric modulation of GluA2 (Q) flop isoform (unknown origin) expressed in HEK293 cells assessed as potentiation of glutamate-evoked calc...More data for this Ligand-Target Pair
Affinity DataEC50: 370nMAssay Description:Positive allosteric modulation of GluA2 (Q) flop isoform (unknown origin) expressed in HEK293 cells assessed as potentiation of glutamate-evoked calc...More data for this Ligand-Target Pair
Affinity DataEC50: 371nMAssay Description:Positive allosteric modulation of GluA2 (Q) flop isoform (unknown origin) expressed in HEK293 cells assessed as potentiation of glutamate-evoked calc...More data for this Ligand-Target Pair
Affinity DataEC50: 390nMAssay Description:Positive allosteric modulation of GluA2 (Q) flop isoform (unknown origin) expressed in HEK293 cells assessed as potentiation of glutamate-evoked calc...More data for this Ligand-Target Pair
Affinity DataEC50: 391nMAssay Description:Positive allosteric modulation of GluA2 (Q) flop isoform (unknown origin) expressed in HEK293 cells assessed as potentiation of glutamate-evoked calc...More data for this Ligand-Target Pair
Affinity DataEC50: 450nMAssay Description:Positive allosteric modulation of GluA2 (Q) flop isoform (unknown origin) expressed in HEK293 cells assessed as potentiation of glutamate-evoked calc...More data for this Ligand-Target Pair
Affinity DataEC50: 454nMAssay Description:Positive allosteric modulation of GluA2 (Q) flop isoform (unknown origin) expressed in HEK293 cells assessed as potentiation of glutamate-evoked calc...More data for this Ligand-Target Pair
Affinity DataEC50: 710nMAssay Description:Positive allosteric modulation of GluA2 (Q) flop isoform (unknown origin) expressed in HEK293 cells assessed as potentiation of glutamate-evoked calc...More data for this Ligand-Target Pair
Affinity DataEC50: 713nMAssay Description:Positive allosteric modulation of GluA2 (Q) flop isoform (unknown origin) expressed in HEK293 cells assessed as potentiation of glutamate-evoked calc...More data for this Ligand-Target Pair
Affinity DataEC50: 810nMAssay Description:Positive allosteric modulation of GluA2 (Q) flop isoform (unknown origin) expressed in HEK293 cells assessed as potentiation of glutamate-evoked calc...More data for this Ligand-Target Pair
Affinity DataEC50: 813nMAssay Description:Positive allosteric modulation of GluA2 (Q) flop isoform (unknown origin) expressed in HEK293 cells assessed as potentiation of glutamate-evoked calc...More data for this Ligand-Target Pair
Affinity DataEC50: 950nMAssay Description:Positive allosteric modulation of GluA2 (Q) flop isoform (unknown origin) expressed in HEK293 cells assessed as potentiation of glutamate-evoked calc...More data for this Ligand-Target Pair
Affinity DataEC50: 955nMAssay Description:Positive allosteric modulation of GluA2 (Q) flop isoform (unknown origin) expressed in HEK293 cells assessed as potentiation of glutamate-evoked calc...More data for this Ligand-Target Pair
Affinity DataEC50: 1.82E+3nMAssay Description:Positive allosteric modulation of GluA2 (Q) flop isoform (unknown origin) expressed in HEK293 cells assessed as potentiation of glutamate-evoked calc...More data for this Ligand-Target Pair
Affinity DataEC50: 1.82E+3nMAssay Description:Positive allosteric modulation of GluA2 (Q) flop isoform (unknown origin) expressed in HEK293 cells assessed as potentiation of glutamate-evoked calc...More data for this Ligand-Target Pair
Affinity DataEC50: 3.19E+3nMAssay Description:Positive allosteric modulation of GluA2 (Q) flop isoform (unknown origin) expressed in HEK293 cells assessed as potentiation of glutamate-evoked calc...More data for this Ligand-Target Pair
Affinity DataEC50: 3.19E+3nMAssay Description:Positive allosteric modulation of GluA2 (Q) flop isoform (unknown origin) expressed in HEK293 cells assessed as potentiation of glutamate-evoked calc...More data for this Ligand-Target Pair
Affinity DataEC50: 5.90E+3nMAssay Description:Positive allosteric modulation of GluA2 (Q) flop isoform (unknown origin) expressed in HEK293 cells assessed as potentiation of glutamate-evoked calc...More data for this Ligand-Target Pair
Affinity DataEC50: 5.90E+3nMAssay Description:Positive allosteric modulation of GluA2 (Q) flop isoform (unknown origin) expressed in HEK293 cells assessed as potentiation of glutamate-evoked calc...More data for this Ligand-Target Pair
Affinity DataEC50: 7.62E+3nMAssay Description:Positive allosteric modulation of GluA2 (Q) flop isoform (unknown origin) expressed in HEK293 cells assessed as potentiation of glutamate-evoked calc...More data for this Ligand-Target Pair
Affinity DataEC50: 7.62E+3nMAssay Description:Positive allosteric modulation of GluA2 (Q) flop isoform (unknown origin) expressed in HEK293 cells assessed as potentiation of glutamate-evoked calc...More data for this Ligand-Target Pair
Affinity DataEC50: <1.00E+4nMAssay Description:Positive allosteric modulation of GluA2 (Q) flop isoform (unknown origin) expressed in HEK293 cells assessed as potentiation of glutamate-evoked calc...More data for this Ligand-Target Pair
Affinity DataEC50: <1.00E+4nMAssay Description:Positive allosteric modulation of GluA2 (Q) flop isoform (unknown origin) expressed in HEK293 cells assessed as potentiation of glutamate-evoked calc...More data for this Ligand-Target Pair
Affinity DataEC50: 1.00E+4nMAssay Description:Positive allosteric modulation of GluA2 (Q) flop isoform (unknown origin) expressed in HEK293 cells assessed as potentiation of glutamate-evoked calc...More data for this Ligand-Target Pair
Affinity DataEC50: >1.00E+4nMAssay Description:Positive allosteric modulation of GluA2 (Q) flop isoform (unknown origin) expressed in HEK293 cells assessed as potentiation of glutamate-evoked calc...More data for this Ligand-Target Pair
Affinity DataEC50: 1.56E+4nMAssay Description:Positive allosteric modulation of GluA2 (Q) flop isoform (unknown origin) expressed in HEK293 cells assessed as potentiation of glutamate-evoked calc...More data for this Ligand-Target Pair
Affinity DataEC50: 1.56E+4nMAssay Description:Positive allosteric modulation of GluA2 (Q) flop isoform (unknown origin) expressed in HEK293 cells assessed as potentiation of glutamate-evoked calc...More data for this Ligand-Target Pair
Affinity DataEC50: 1.56E+4nMAssay Description:Positive allosteric modulation of GluA2 (Q) flop isoform (unknown origin) expressed in HEK293 cells assessed as potentiation of glutamate-evoked calc...More data for this Ligand-Target Pair
Affinity DataEC50: 1.56E+4nMAssay Description:Positive allosteric modulation of GluA2 (Q) flop isoform (unknown origin) expressed in HEK293 cells assessed as potentiation of glutamate-evoked calc...More data for this Ligand-Target Pair
Affinity DataEC50: 1.87E+4nMAssay Description:Positive allosteric modulation of GluA2 (Q) flop isoform (unknown origin) expressed in HEK293 cells assessed as potentiation of glutamate-evoked calc...More data for this Ligand-Target Pair
Affinity DataEC50: 1.87E+4nMAssay Description:Positive allosteric modulation of GluA2 (Q) flop isoform (unknown origin) expressed in HEK293 cells assessed as potentiation of glutamate-evoked calc...More data for this Ligand-Target Pair
Affinity DataEC50: 2.04E+4nMAssay Description:Positive allosteric modulation of GluA2 (Q) flop isoform (unknown origin) expressed in HEK293 cells assessed as potentiation of glutamate-evoked calc...More data for this Ligand-Target Pair
Affinity DataEC50: 2.04E+4nMAssay Description:Positive allosteric modulation of GluA2 (Q) flop isoform (unknown origin) expressed in HEK293 cells assessed as potentiation of glutamate-evoked calc...More data for this Ligand-Target Pair
Affinity DataEC50: 5.06E+4nMAssay Description:Positive allosteric modulation of GluA2 (Q) flop isoform (unknown origin) expressed in HEK293 cells assessed as potentiation of glutamate-evoked calc...More data for this Ligand-Target Pair
Affinity DataEC50: 5.06E+4nMAssay Description:Positive allosteric modulation of GluA2 (Q) flop isoform (unknown origin) expressed in HEK293 cells assessed as potentiation of glutamate-evoked calc...More data for this Ligand-Target Pair