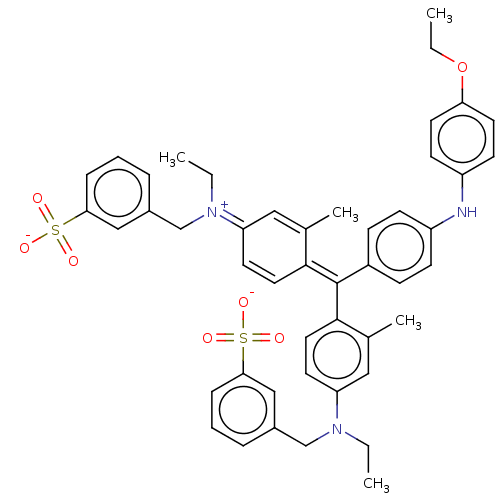
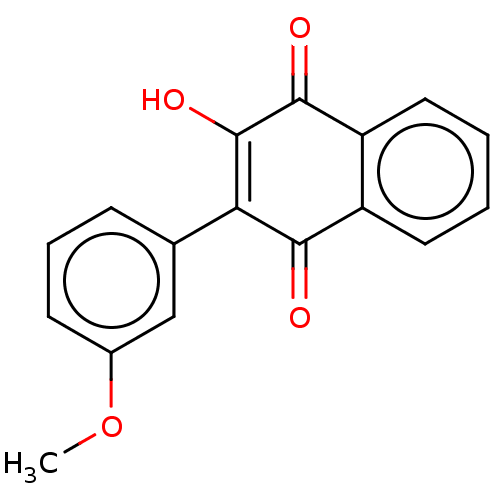
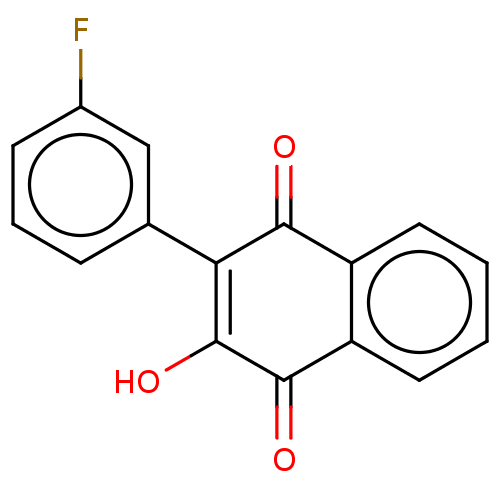
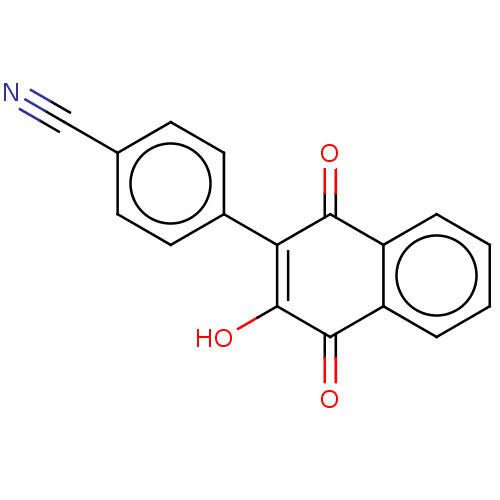
Report error Found 16 Enz. Inhib. hit(s) with all data for entry = 50002892
Affinity DataIC50: 22nMAssay Description:Antagonist activity at human P2X7R expressed in HEK293 cells assessed as inhibition of ATP-induced ethidium iodide uptake preincubated for 10 mins fo...More data for this Ligand-Target Pair
Affinity DataIC50: 23nMAssay Description:Antagonist activity at P2X7R in Swiss Webster mouse peritoneal macrophages assessed as inhibition of ATP-induced propidium iodide uptake preincubated...More data for this Ligand-Target Pair
Affinity DataIC50: 48nMAssay Description:Antagonist activity at P2X7R in Swiss Webster mouse peritoneal macrophages assessed as inhibition of ATP-induced propidium iodide uptake preincubated...More data for this Ligand-Target Pair
Affinity DataIC50: 71nMAssay Description:Antagonist activity at human P2X7R expressed in HEK293 cells assessed as inhibition of ATP-induced ethidium iodide uptake preincubated for 10 mins fo...More data for this Ligand-Target Pair
Affinity DataIC50: 109nMAssay Description:Antagonist activity at human P2X7R expressed in HEK293 cells assessed as inhibition of ATP-induced ethidium iodide uptake preincubated for 10 mins fo...More data for this Ligand-Target Pair
Affinity DataIC50: 414nMAssay Description:Antagonist activity at human P2X7R expressed in HEK293 cells assessed as inhibition of ATP-induced ethidium iodide uptake preincubated for 10 mins fo...More data for this Ligand-Target Pair
Affinity DataIC50: 536nMAssay Description:Antagonist activity at human P2X7R expressed in HEK293 cells assessed as inhibition of ATP-induced ethidium iodide uptake preincubated for 10 mins fo...More data for this Ligand-Target Pair
Affinity DataIC50: 712nMAssay Description:Antagonist activity at P2X7R in Swiss Webster mouse peritoneal macrophages assessed as inhibition of ATP-induced propidium iodide uptake preincubated...More data for this Ligand-Target Pair
Affinity DataIC50: 897nMAssay Description:Antagonist activity at P2X7R in Swiss Webster mouse peritoneal macrophages assessed as inhibition of ATP-induced propidium iodide uptake preincubated...More data for this Ligand-Target Pair
Affinity DataIC50: 3.17E+3nMAssay Description:Antagonist activity at P2X7R in Swiss Webster mouse peritoneal macrophages assessed as inhibition of ATP-induced propidium iodide uptake preincubated...More data for this Ligand-Target Pair
Affinity DataIC50: 1.52E+4nMAssay Description:Antagonist activity at P2X7R in Swiss Webster mouse peritoneal macrophages assessed as inhibition of ATP-induced propidium iodide uptake preincubated...More data for this Ligand-Target Pair
Affinity DataIC50: 3.35E+4nMAssay Description:Antagonist activity at P2X7R in Swiss Webster mouse peritoneal macrophages assessed as inhibition of ATP-induced propidium iodide uptake preincubated...More data for this Ligand-Target Pair
Affinity DataIC50: 4.58E+4nMAssay Description:Antagonist activity at P2X7R in Swiss Webster mouse peritoneal macrophages assessed as inhibition of ATP-induced propidium iodide uptake preincubated...More data for this Ligand-Target Pair
Affinity DataIC50: 7.55E+4nMAssay Description:Antagonist activity at P2X7R in Swiss Webster mouse peritoneal macrophages assessed as inhibition of ATP-induced propidium iodide uptake preincubated...More data for this Ligand-Target Pair
Affinity DataIC50: 8.01E+4nMAssay Description:Antagonist activity at P2X7R in Swiss Webster mouse peritoneal macrophages assessed as inhibition of ATP-induced propidium iodide uptake preincubated...More data for this Ligand-Target Pair
Affinity DataIC50: 8.13E+4nMAssay Description:Antagonist activity at P2X7R in Swiss Webster mouse peritoneal macrophages assessed as inhibition of ATP-induced propidium iodide uptake preincubated...More data for this Ligand-Target Pair