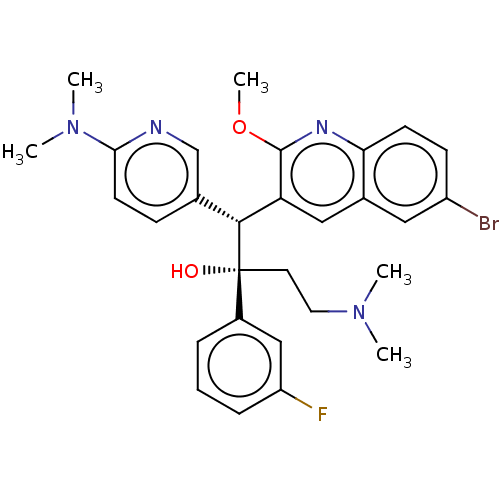
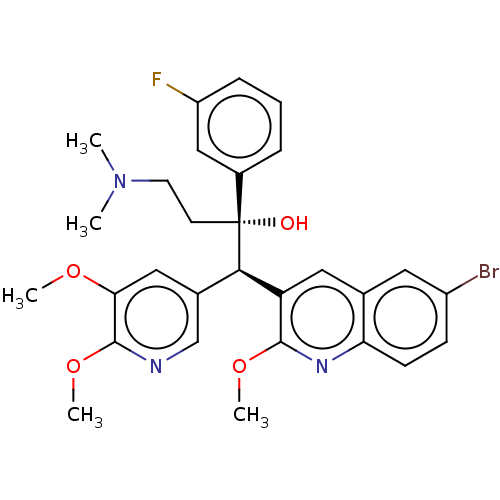
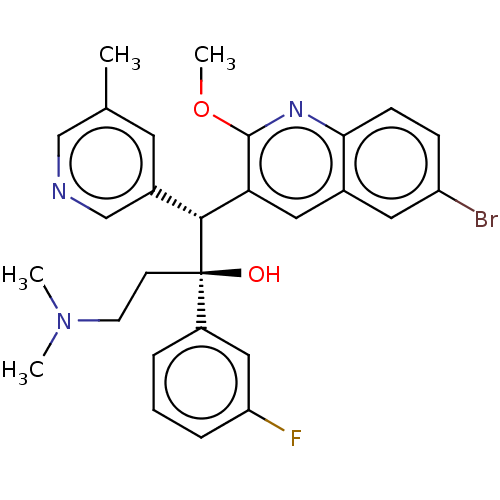
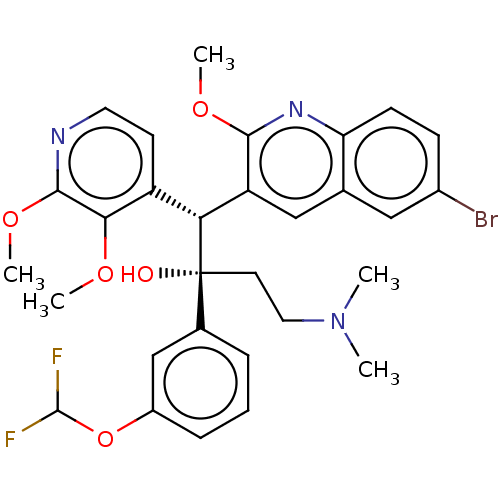
Report error Found 16 Enz. Inhib. hit(s) with all data for entry = 50002014
TargetPotassium voltage-gated channel subfamily H member 2(Human)
University of Auckland
Curated by ChEMBL
University of Auckland
Curated by ChEMBL
Affinity DataIC50: 370nMAssay Description:Inhibition of human ERGMore data for this Ligand-Target Pair
TargetPotassium voltage-gated channel subfamily H member 2(Human)
University of Auckland
Curated by ChEMBL
University of Auckland
Curated by ChEMBL
Affinity DataIC50: 460nMAssay Description:Inhibition of human ERGMore data for this Ligand-Target Pair
TargetPotassium voltage-gated channel subfamily H member 2(Human)
University of Auckland
Curated by ChEMBL
University of Auckland
Curated by ChEMBL
Affinity DataIC50: 530nMAssay Description:Inhibition of human ERGMore data for this Ligand-Target Pair
TargetPotassium voltage-gated channel subfamily H member 2(Human)
University of Auckland
Curated by ChEMBL
University of Auckland
Curated by ChEMBL
Affinity DataIC50: 880nMAssay Description:Inhibition of human ERGMore data for this Ligand-Target Pair
TargetPotassium voltage-gated channel subfamily H member 2(Human)
University of Auckland
Curated by ChEMBL
University of Auckland
Curated by ChEMBL
Affinity DataIC50: 890nMAssay Description:Inhibition of human ERGMore data for this Ligand-Target Pair
TargetPotassium voltage-gated channel subfamily H member 2(Human)
University of Auckland
Curated by ChEMBL
University of Auckland
Curated by ChEMBL
Affinity DataIC50: 1.90E+3nMAssay Description:Inhibition of human ERGMore data for this Ligand-Target Pair
Affinity DataIC50: 4.60E+3nMAssay Description:Inhibition of CYP3A4 (unknown origin) after 20 minsMore data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMAssay Description:Inhibition of CYP3A4 (unknown origin) after 20 minsMore data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMAssay Description:Inhibition of CYP3A4 (unknown origin) after 20 minsMore data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMAssay Description:Inhibition of CYP3A4 (unknown origin) after 20 minsMore data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMAssay Description:Inhibition of CYP3A4 (unknown origin) after 20 minsMore data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMAssay Description:Inhibition of CYP3A4 (unknown origin) after 20 minsMore data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMAssay Description:Inhibition of CYP3A4 (unknown origin) after 20 minsMore data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMAssay Description:Inhibition of CYP3A4 (unknown origin) after 20 minsMore data for this Ligand-Target Pair
Affinity DataIC50: 5.00E+4nMAssay Description:Inhibition of CYP3A4 (unknown origin) after 20 minsMore data for this Ligand-Target Pair
Affinity DataIC50: 5.00E+4nMAssay Description:Inhibition of CYP3A4 (unknown origin) after 20 minsMore data for this Ligand-Target Pair