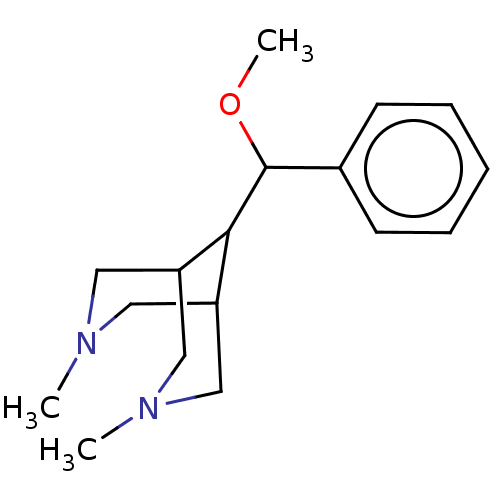
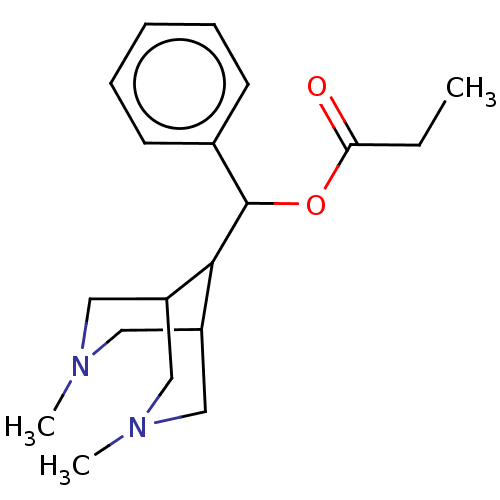
Report error Found 6 Enz. Inhib. hit(s) with all data for entry = 50048707
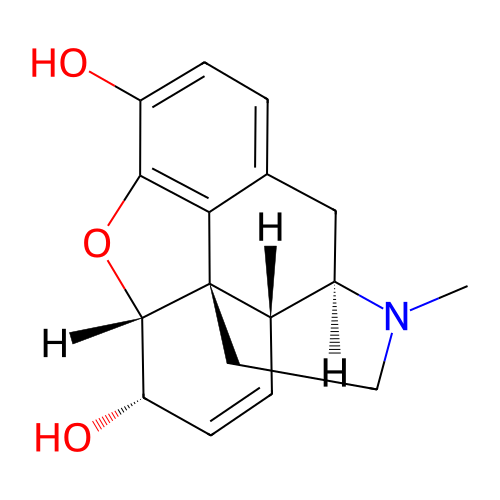
TargetDelta-type opioid receptor/Kappa-type opioid receptor/Mu-type opioid receptor/Sigma non-opioid intracellular receptor 1(Rat)
TBA
Curated by ChEMBL
TBA
Curated by ChEMBL
Affinity DataIC50: 160nMAssay Description:Concentration required to inhibit opiate receptor using radioligand [3H]- EtorphineMore data for this Ligand-Target Pair
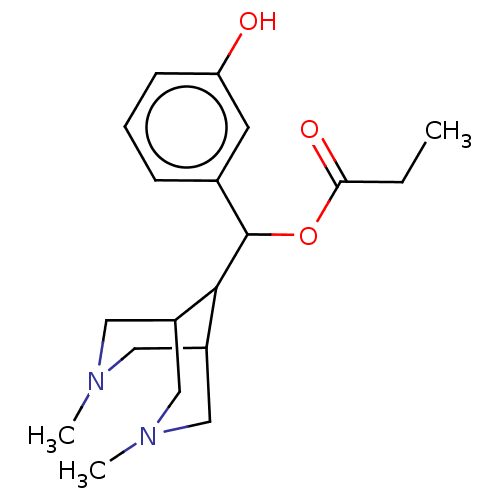
TargetDelta-type opioid receptor/Kappa-type opioid receptor/Mu-type opioid receptor/Sigma non-opioid intracellular receptor 1(Rat)
TBA
Curated by ChEMBL
TBA
Curated by ChEMBL
Affinity DataIC50: 530nMAssay Description:Concentration required to inhibit opiate receptor using radioligand [3H]- EtorphineMore data for this Ligand-Target Pair
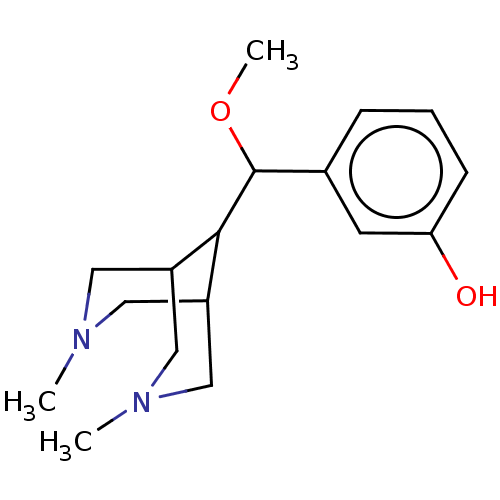
TargetDelta-type opioid receptor/Kappa-type opioid receptor/Mu-type opioid receptor/Sigma non-opioid intracellular receptor 1(Rat)
TBA
Curated by ChEMBL
TBA
Curated by ChEMBL
Affinity DataIC50: 2.30E+3nMAssay Description:Concentration required to inhibit opiate receptor using radioligand [3H]- EtorphineMore data for this Ligand-Target Pair
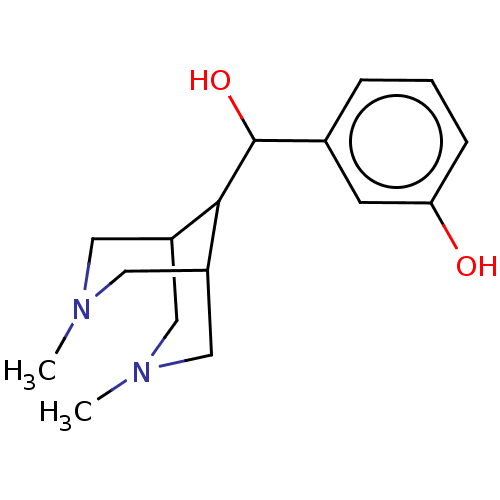
TargetDelta-type opioid receptor/Kappa-type opioid receptor/Mu-type opioid receptor/Sigma non-opioid intracellular receptor 1(Rat)
TBA
Curated by ChEMBL
TBA
Curated by ChEMBL
Affinity DataIC50: 3.00E+3nMAssay Description:Concentration required to inhibit opiate receptor using radioligand [3H]- EtorphineMore data for this Ligand-Target Pair
TargetDelta-type opioid receptor/Kappa-type opioid receptor/Mu-type opioid receptor/Sigma non-opioid intracellular receptor 1(Rat)
TBA
Curated by ChEMBL
TBA
Curated by ChEMBL
Affinity DataIC50: 7.20E+3nMAssay Description:Concentration required to inhibit opiate receptor using radioligand [3H]- EtorphineMore data for this Ligand-Target Pair
TargetDelta-type opioid receptor/Kappa-type opioid receptor/Mu-type opioid receptor/Sigma non-opioid intracellular receptor 1(Rat)
TBA
Curated by ChEMBL
TBA
Curated by ChEMBL
Affinity DataIC50: 7.60E+3nMAssay Description:Concentration required to inhibit opiate receptor using radioligand [3H]- EtorphineMore data for this Ligand-Target Pair