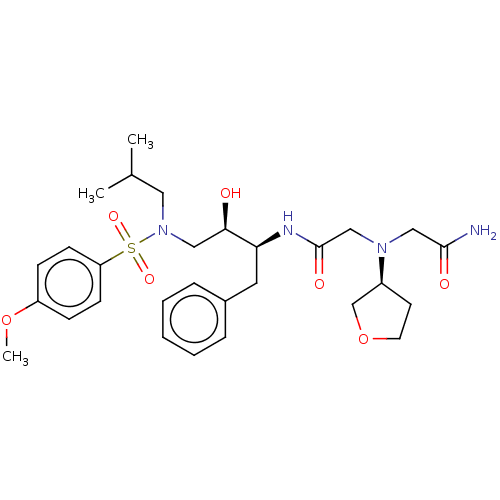
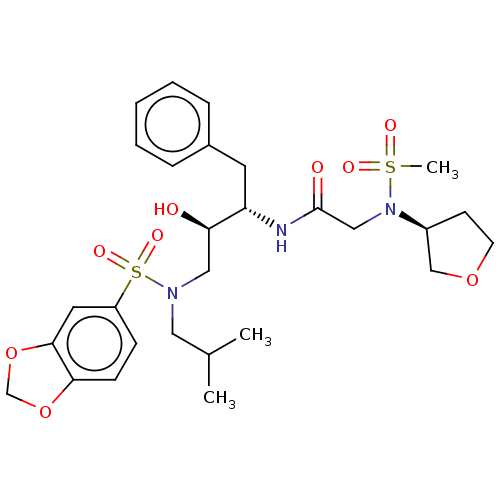
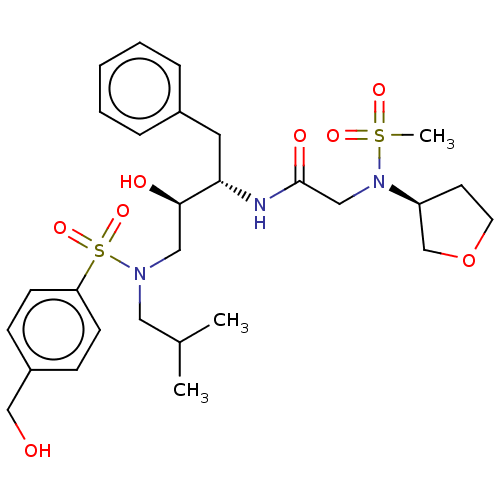
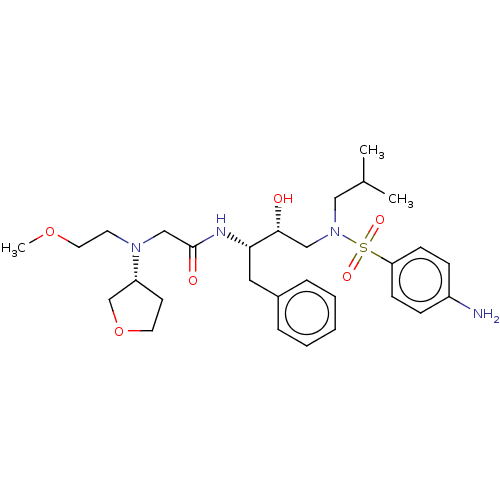
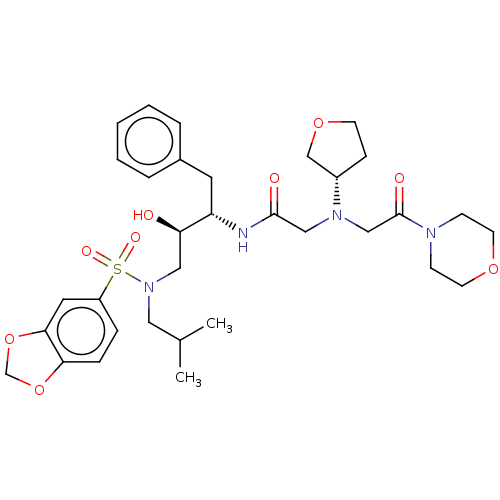
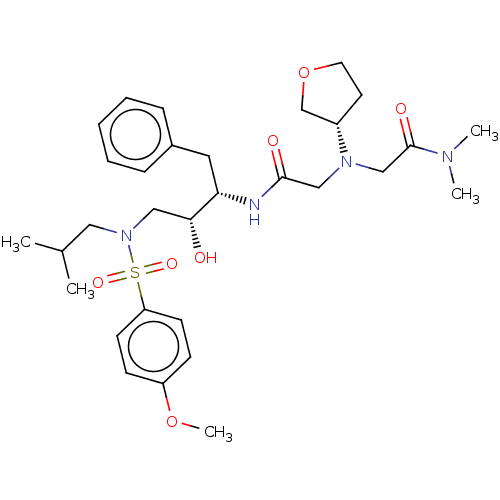
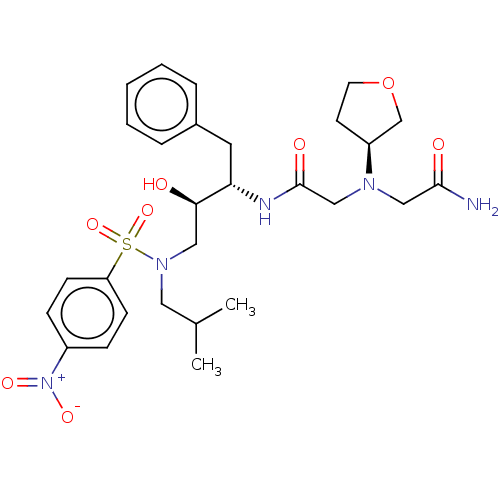
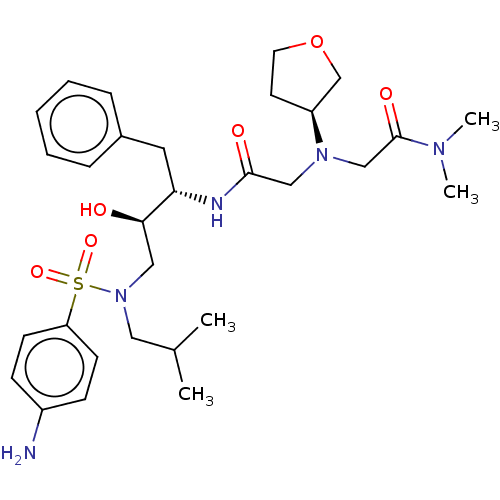
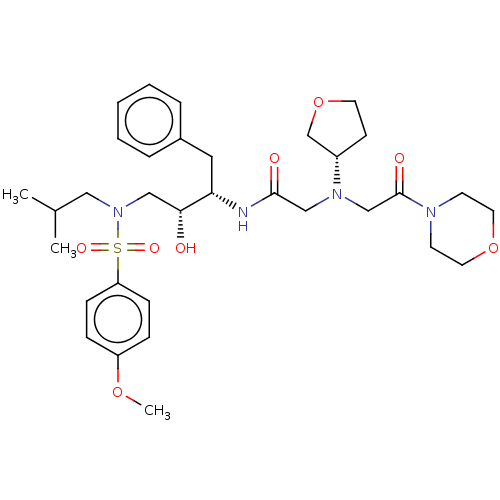
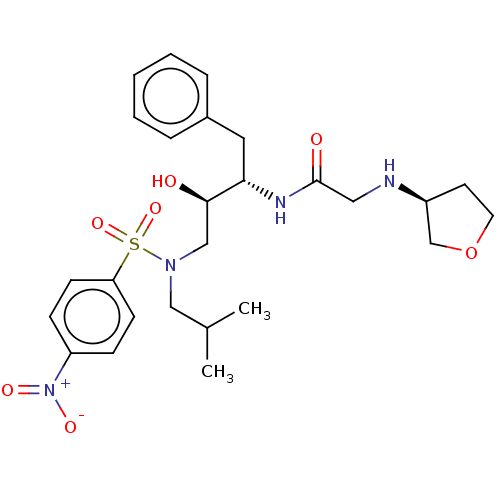
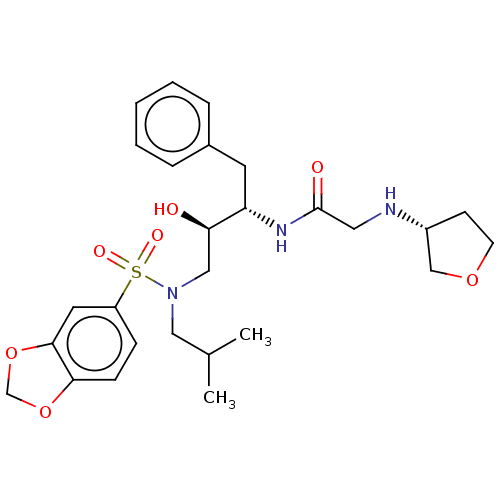
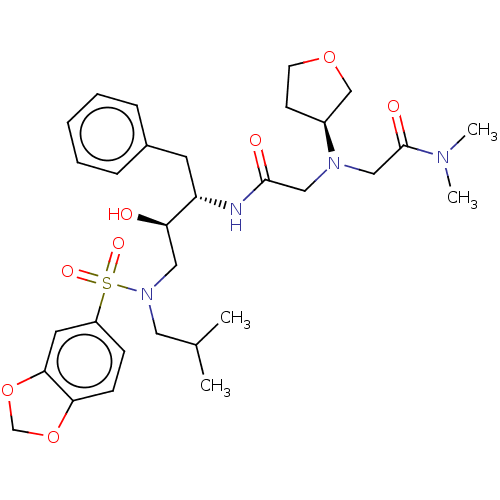
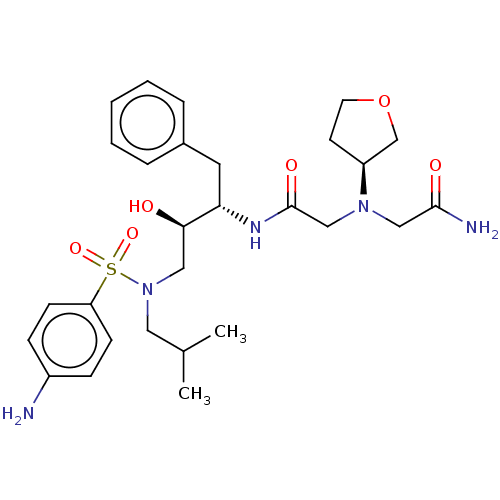
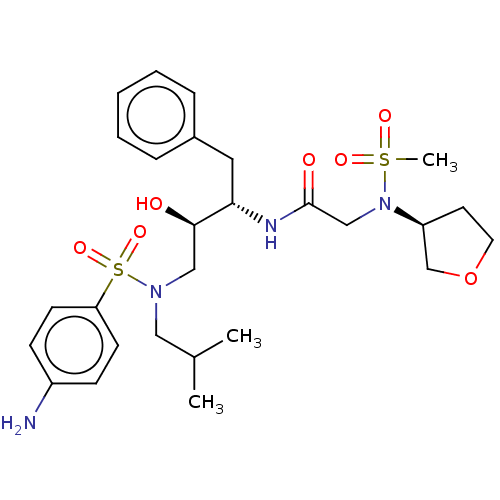
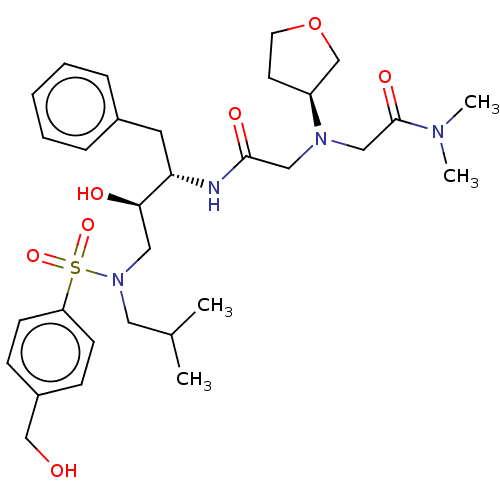
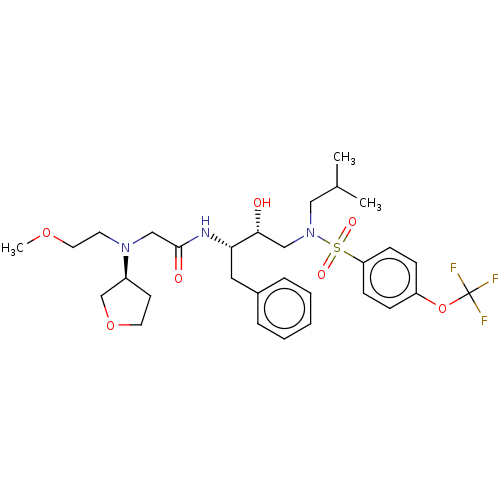
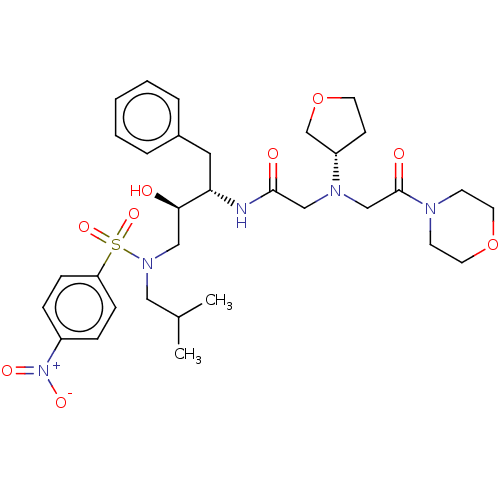
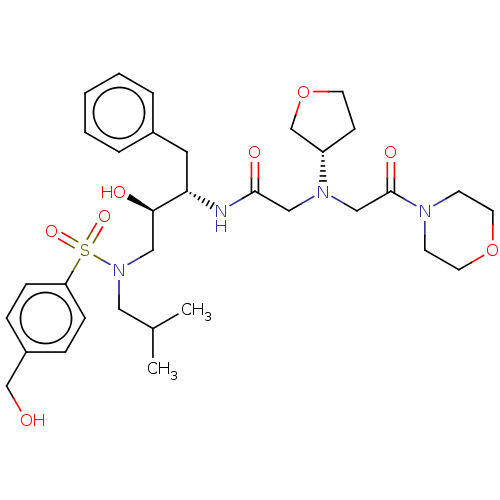
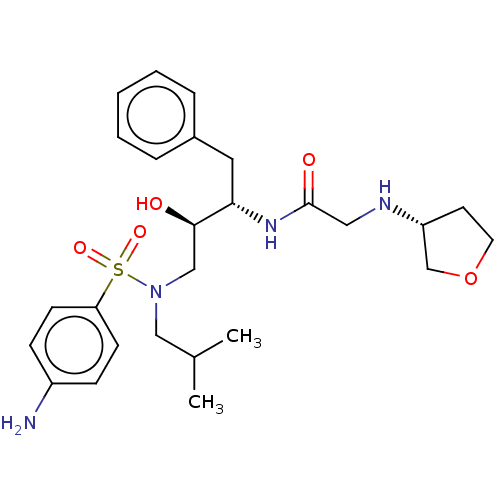
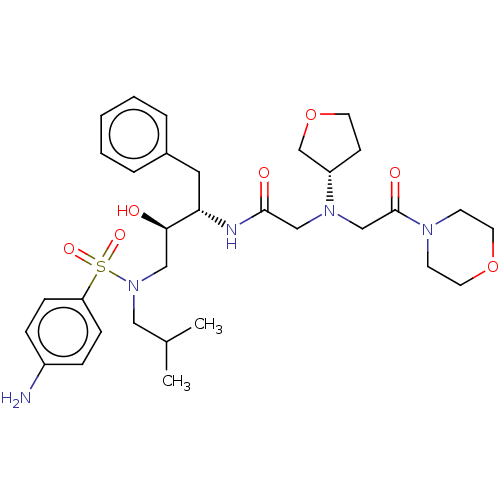
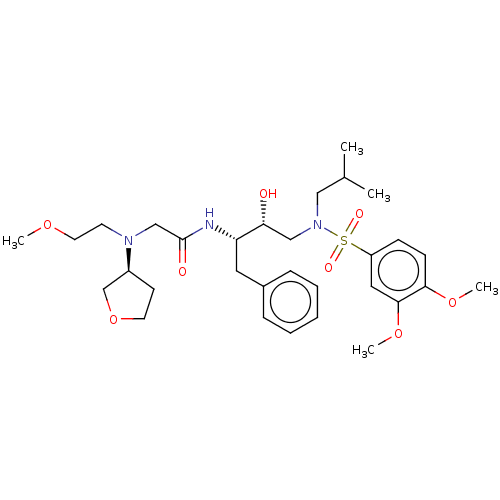
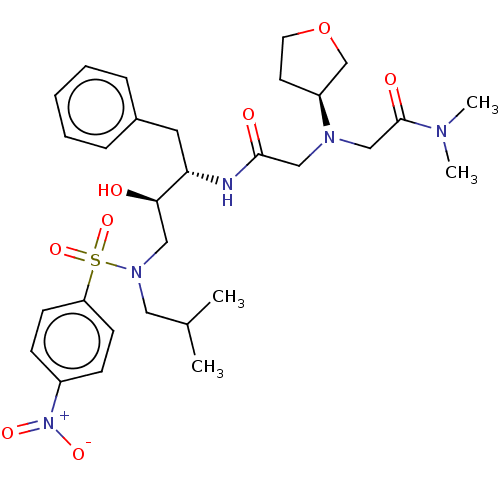
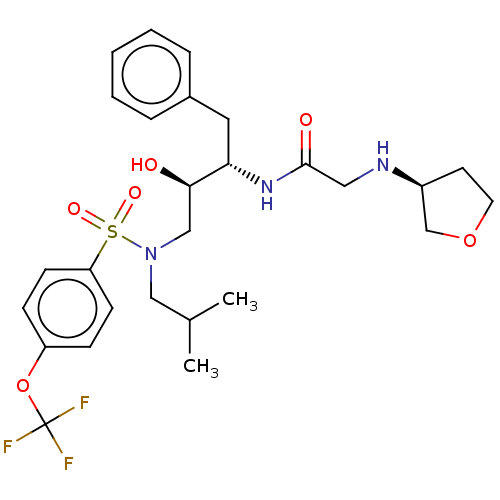
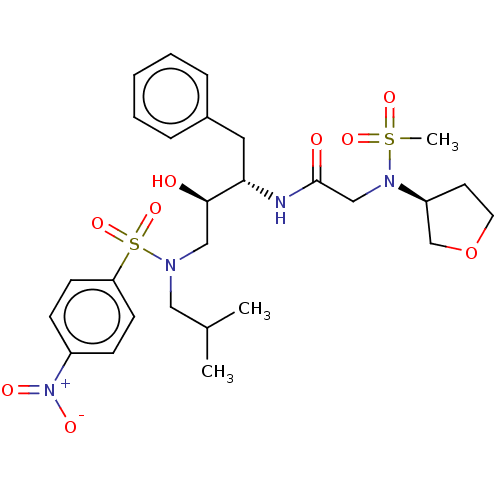
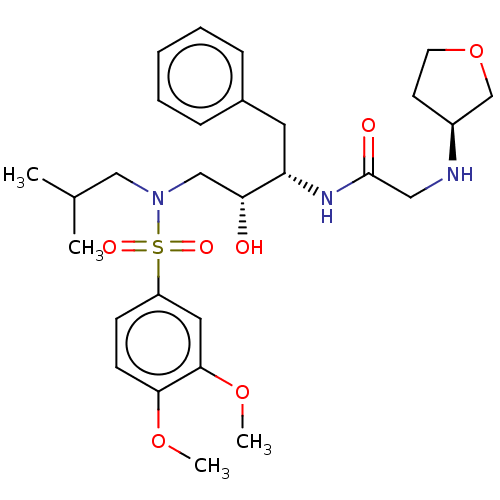
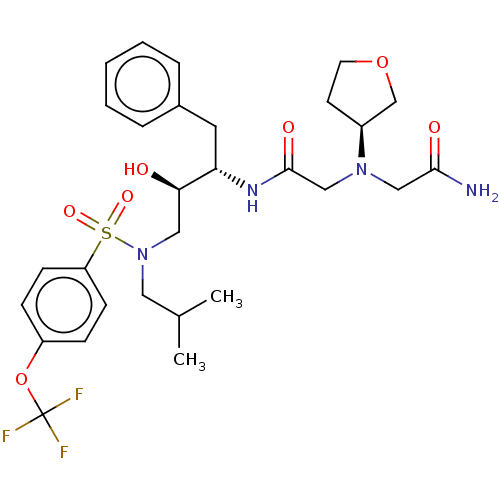
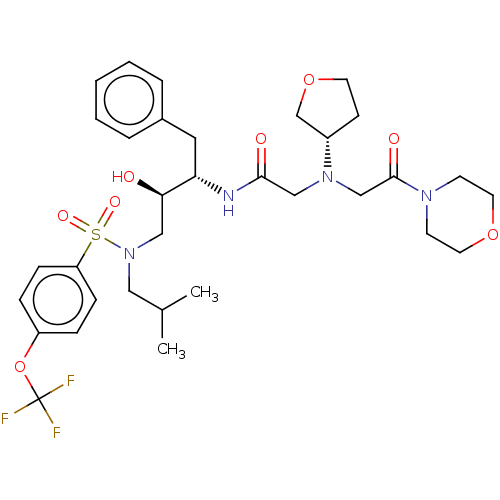
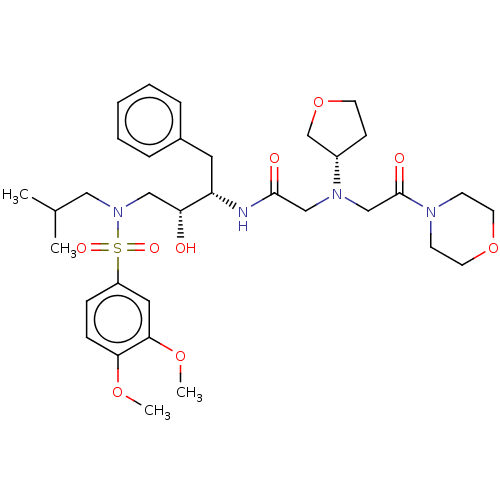
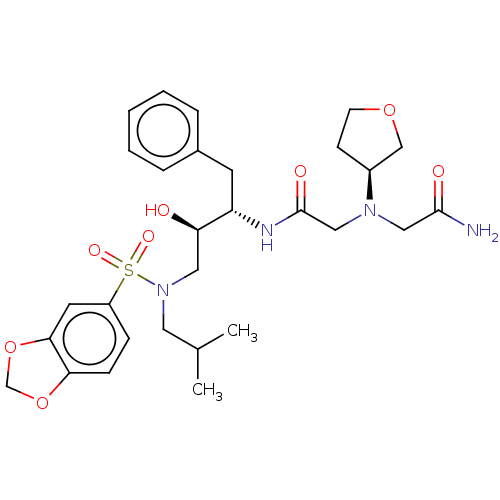
Report error Found 49 Enz. Inhib. hit(s) with all data for entry = 50001154
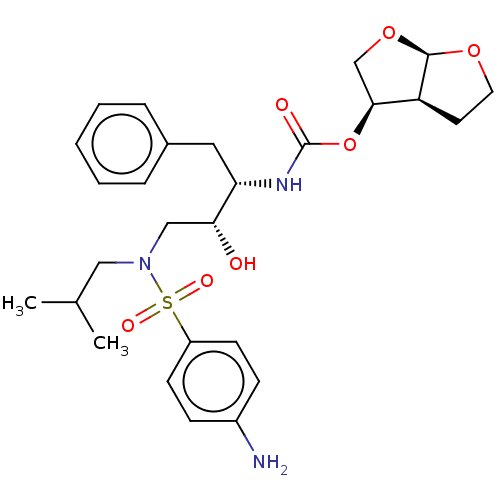
Affinity DataIC50: 0.0600nMAssay Description:Inhibition of HIV1 protease expressed in Escherichia coli using Arg-Glu (EDANS)-Ser-GlnAsn-Tyr-Pro-Ile-Val-Gln-Lys(DABCYL)-Arg as substrate preincuba...More data for this Ligand-Target Pair
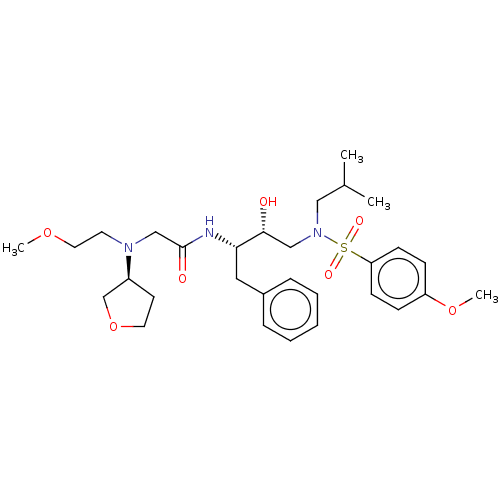
Affinity DataIC50: 0.350nMAssay Description:Inhibition of HIV1 protease expressed in Escherichia coli using Arg-Glu (EDANS)-Ser-GlnAsn-Tyr-Pro-Ile-Val-Gln-Lys(DABCYL)-Arg as substrate preincuba...More data for this Ligand-Target Pair
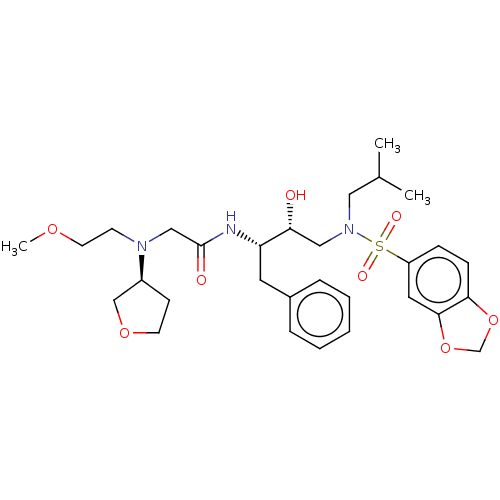
Affinity DataIC50: 0.770nMAssay Description:Inhibition of HIV1 protease expressed in Escherichia coli using Arg-Glu (EDANS)-Ser-GlnAsn-Tyr-Pro-Ile-Val-Gln-Lys(DABCYL)-Arg as substrate preincuba...More data for this Ligand-Target Pair
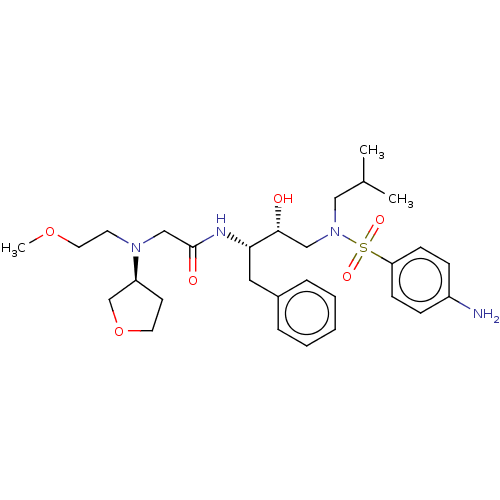
Affinity DataIC50: 0.880nMAssay Description:Inhibition of HIV1 protease expressed in Escherichia coli using Arg-Glu (EDANS)-Ser-GlnAsn-Tyr-Pro-Ile-Val-Gln-Lys(DABCYL)-Arg as substrate preincuba...More data for this Ligand-Target Pair
Affinity DataIC50: 1.70nMAssay Description:Inhibition of HIV1 protease expressed in Escherichia coli using Arg-Glu (EDANS)-Ser-GlnAsn-Tyr-Pro-Ile-Val-Gln-Lys(DABCYL)-Arg as substrate preincuba...More data for this Ligand-Target Pair
Affinity DataIC50: 2.20nMAssay Description:Inhibition of HIV1 protease expressed in Escherichia coli using Arg-Glu (EDANS)-Ser-GlnAsn-Tyr-Pro-Ile-Val-Gln-Lys(DABCYL)-Arg as substrate preincuba...More data for this Ligand-Target Pair
Affinity DataIC50: 4.10nMAssay Description:Inhibition of HIV1 protease expressed in Escherichia coli using Arg-Glu (EDANS)-Ser-GlnAsn-Tyr-Pro-Ile-Val-Gln-Lys(DABCYL)-Arg as substrate preincuba...More data for this Ligand-Target Pair
Affinity DataIC50: 4.30nMAssay Description:Inhibition of HIV1 protease expressed in Escherichia coli using Arg-Glu (EDANS)-Ser-GlnAsn-Tyr-Pro-Ile-Val-Gln-Lys(DABCYL)-Arg as substrate preincuba...More data for this Ligand-Target Pair
Affinity DataIC50: 6nMAssay Description:Inhibition of HIV1 protease expressed in Escherichia coli using Arg-Glu (EDANS)-Ser-GlnAsn-Tyr-Pro-Ile-Val-Gln-Lys(DABCYL)-Arg as substrate preincuba...More data for this Ligand-Target Pair
Affinity DataIC50: 6.90nMAssay Description:Inhibition of HIV1 protease expressed in Escherichia coli using Arg-Glu (EDANS)-Ser-GlnAsn-Tyr-Pro-Ile-Val-Gln-Lys(DABCYL)-Arg as substrate preincuba...More data for this Ligand-Target Pair
Affinity DataIC50: 9.90nMAssay Description:Inhibition of HIV1 protease expressed in Escherichia coli using Arg-Glu (EDANS)-Ser-GlnAsn-Tyr-Pro-Ile-Val-Gln-Lys(DABCYL)-Arg as substrate preincuba...More data for this Ligand-Target Pair
Affinity DataIC50: 10nMAssay Description:Inhibition of HIV1 protease expressed in Escherichia coli using Arg-Glu (EDANS)-Ser-GlnAsn-Tyr-Pro-Ile-Val-Gln-Lys(DABCYL)-Arg as substrate preincuba...More data for this Ligand-Target Pair
Affinity DataIC50: 13nMAssay Description:Inhibition of HIV1 protease expressed in Escherichia coli using Arg-Glu (EDANS)-Ser-GlnAsn-Tyr-Pro-Ile-Val-Gln-Lys(DABCYL)-Arg as substrate preincuba...More data for this Ligand-Target Pair
Affinity DataIC50: 16nMAssay Description:Inhibition of HIV1 protease expressed in Escherichia coli using Arg-Glu (EDANS)-Ser-GlnAsn-Tyr-Pro-Ile-Val-Gln-Lys(DABCYL)-Arg as substrate preincuba...More data for this Ligand-Target Pair
Affinity DataIC50: 18nMAssay Description:Inhibition of HIV1 protease expressed in Escherichia coli using Arg-Glu (EDANS)-Ser-GlnAsn-Tyr-Pro-Ile-Val-Gln-Lys(DABCYL)-Arg as substrate preincuba...More data for this Ligand-Target Pair
Affinity DataIC50: 19nMAssay Description:Inhibition of HIV1 protease expressed in Escherichia coli using Arg-Glu (EDANS)-Ser-GlnAsn-Tyr-Pro-Ile-Val-Gln-Lys(DABCYL)-Arg as substrate preincuba...More data for this Ligand-Target Pair
Affinity DataIC50: 20nMAssay Description:Inhibition of HIV1 protease expressed in Escherichia coli using Arg-Glu (EDANS)-Ser-GlnAsn-Tyr-Pro-Ile-Val-Gln-Lys(DABCYL)-Arg as substrate preincuba...More data for this Ligand-Target Pair
Affinity DataIC50: 23nMAssay Description:Inhibition of HIV1 protease expressed in Escherichia coli using Arg-Glu (EDANS)-Ser-GlnAsn-Tyr-Pro-Ile-Val-Gln-Lys(DABCYL)-Arg as substrate preincuba...More data for this Ligand-Target Pair
Affinity DataIC50: 23nMAssay Description:Inhibition of HIV1 protease expressed in Escherichia coli using Arg-Glu (EDANS)-Ser-GlnAsn-Tyr-Pro-Ile-Val-Gln-Lys(DABCYL)-Arg as substrate preincuba...More data for this Ligand-Target Pair
Affinity DataIC50: 23nMAssay Description:Inhibition of HIV1 protease expressed in Escherichia coli using Arg-Glu (EDANS)-Ser-GlnAsn-Tyr-Pro-Ile-Val-Gln-Lys(DABCYL)-Arg as substrate preincuba...More data for this Ligand-Target Pair
Affinity DataIC50: 27nMAssay Description:Inhibition of HIV1 protease expressed in Escherichia coli using Arg-Glu (EDANS)-Ser-GlnAsn-Tyr-Pro-Ile-Val-Gln-Lys(DABCYL)-Arg as substrate preincuba...More data for this Ligand-Target Pair
Affinity DataIC50: 29nMAssay Description:Inhibition of HIV1 protease expressed in Escherichia coli using Arg-Glu (EDANS)-Ser-GlnAsn-Tyr-Pro-Ile-Val-Gln-Lys(DABCYL)-Arg as substrate preincuba...More data for this Ligand-Target Pair
Affinity DataIC50: 34nMAssay Description:Inhibition of HIV1 protease expressed in Escherichia coli using Arg-Glu (EDANS)-Ser-GlnAsn-Tyr-Pro-Ile-Val-Gln-Lys(DABCYL)-Arg as substrate preincuba...More data for this Ligand-Target Pair
Affinity DataIC50: 38nMAssay Description:Inhibition of HIV1 protease expressed in Escherichia coli using Arg-Glu (EDANS)-Ser-GlnAsn-Tyr-Pro-Ile-Val-Gln-Lys(DABCYL)-Arg as substrate preincuba...More data for this Ligand-Target Pair
Affinity DataIC50: 44nMAssay Description:Inhibition of HIV1 protease expressed in Escherichia coli using Arg-Glu (EDANS)-Ser-GlnAsn-Tyr-Pro-Ile-Val-Gln-Lys(DABCYL)-Arg as substrate preincuba...More data for this Ligand-Target Pair
Affinity DataIC50: 46nMAssay Description:Inhibition of HIV1 protease expressed in Escherichia coli using Arg-Glu (EDANS)-Ser-GlnAsn-Tyr-Pro-Ile-Val-Gln-Lys(DABCYL)-Arg as substrate preincuba...More data for this Ligand-Target Pair
Affinity DataIC50: 53nMAssay Description:Inhibition of HIV1 protease expressed in Escherichia coli using Arg-Glu (EDANS)-Ser-GlnAsn-Tyr-Pro-Ile-Val-Gln-Lys(DABCYL)-Arg as substrate preincuba...More data for this Ligand-Target Pair
Affinity DataIC50: 57nMAssay Description:Inhibition of HIV1 protease expressed in Escherichia coli using Arg-Glu (EDANS)-Ser-GlnAsn-Tyr-Pro-Ile-Val-Gln-Lys(DABCYL)-Arg as substrate preincuba...More data for this Ligand-Target Pair
Affinity DataIC50: 57nMAssay Description:Inhibition of HIV1 protease expressed in Escherichia coli using Arg-Glu (EDANS)-Ser-GlnAsn-Tyr-Pro-Ile-Val-Gln-Lys(DABCYL)-Arg as substrate preincuba...More data for this Ligand-Target Pair
Affinity DataIC50: 60nMAssay Description:Inhibition of HIV1 protease expressed in Escherichia coli using Arg-Glu (EDANS)-Ser-GlnAsn-Tyr-Pro-Ile-Val-Gln-Lys(DABCYL)-Arg as substrate preincuba...More data for this Ligand-Target Pair
Affinity DataIC50: 65nMAssay Description:Inhibition of HIV1 protease expressed in Escherichia coli using Arg-Glu (EDANS)-Ser-GlnAsn-Tyr-Pro-Ile-Val-Gln-Lys(DABCYL)-Arg as substrate preincuba...More data for this Ligand-Target Pair
Affinity DataIC50: 68nMAssay Description:Inhibition of HIV1 protease expressed in Escherichia coli using Arg-Glu (EDANS)-Ser-GlnAsn-Tyr-Pro-Ile-Val-Gln-Lys(DABCYL)-Arg as substrate preincuba...More data for this Ligand-Target Pair
Affinity DataIC50: 98nMAssay Description:Inhibition of HIV1 protease expressed in Escherichia coli using Arg-Glu (EDANS)-Ser-GlnAsn-Tyr-Pro-Ile-Val-Gln-Lys(DABCYL)-Arg as substrate preincuba...More data for this Ligand-Target Pair
Affinity DataIC50: 110nMAssay Description:Inhibition of HIV1 protease expressed in Escherichia coli using Arg-Glu (EDANS)-Ser-GlnAsn-Tyr-Pro-Ile-Val-Gln-Lys(DABCYL)-Arg as substrate preincuba...More data for this Ligand-Target Pair
Affinity DataIC50: 141nMAssay Description:Inhibition of HIV1 protease expressed in Escherichia coli using Arg-Glu (EDANS)-Ser-GlnAsn-Tyr-Pro-Ile-Val-Gln-Lys(DABCYL)-Arg as substrate preincuba...More data for this Ligand-Target Pair
Affinity DataIC50: 160nMAssay Description:Inhibition of HIV1 protease expressed in Escherichia coli using Arg-Glu (EDANS)-Ser-GlnAsn-Tyr-Pro-Ile-Val-Gln-Lys(DABCYL)-Arg as substrate preincuba...More data for this Ligand-Target Pair
Affinity DataIC50: 190nMAssay Description:Inhibition of HIV1 protease expressed in Escherichia coli using Arg-Glu (EDANS)-Ser-GlnAsn-Tyr-Pro-Ile-Val-Gln-Lys(DABCYL)-Arg as substrate preincuba...More data for this Ligand-Target Pair
Affinity DataIC50: 270nMAssay Description:Inhibition of HIV1 protease expressed in Escherichia coli using Arg-Glu (EDANS)-Ser-GlnAsn-Tyr-Pro-Ile-Val-Gln-Lys(DABCYL)-Arg as substrate preincuba...More data for this Ligand-Target Pair
Affinity DataIC50: 290nMAssay Description:Inhibition of HIV1 protease expressed in Escherichia coli using Arg-Glu (EDANS)-Ser-GlnAsn-Tyr-Pro-Ile-Val-Gln-Lys(DABCYL)-Arg as substrate preincuba...More data for this Ligand-Target Pair
Affinity DataIC50: 291nMAssay Description:Inhibition of HIV1 protease expressed in Escherichia coli using Arg-Glu (EDANS)-Ser-GlnAsn-Tyr-Pro-Ile-Val-Gln-Lys(DABCYL)-Arg as substrate preincuba...More data for this Ligand-Target Pair
Affinity DataIC50: 310nMAssay Description:Inhibition of HIV1 protease expressed in Escherichia coli using Arg-Glu (EDANS)-Ser-GlnAsn-Tyr-Pro-Ile-Val-Gln-Lys(DABCYL)-Arg as substrate preincuba...More data for this Ligand-Target Pair
Affinity DataIC50: 530nMAssay Description:Inhibition of HIV1 protease expressed in Escherichia coli using Arg-Glu (EDANS)-Ser-GlnAsn-Tyr-Pro-Ile-Val-Gln-Lys(DABCYL)-Arg as substrate preincuba...More data for this Ligand-Target Pair
Affinity DataIC50: 910nMAssay Description:Inhibition of HIV1 protease expressed in Escherichia coli using Arg-Glu (EDANS)-Ser-GlnAsn-Tyr-Pro-Ile-Val-Gln-Lys(DABCYL)-Arg as substrate preincuba...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+3nMAssay Description:Inhibition of HIV1 protease expressed in Escherichia coli using Arg-Glu (EDANS)-Ser-GlnAsn-Tyr-Pro-Ile-Val-Gln-Lys(DABCYL)-Arg as substrate preincuba...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+3nMAssay Description:Inhibition of HIV1 protease expressed in Escherichia coli using Arg-Glu (EDANS)-Ser-GlnAsn-Tyr-Pro-Ile-Val-Gln-Lys(DABCYL)-Arg as substrate preincuba...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+3nMAssay Description:Inhibition of HIV1 protease expressed in Escherichia coli using Arg-Glu (EDANS)-Ser-GlnAsn-Tyr-Pro-Ile-Val-Gln-Lys(DABCYL)-Arg as substrate preincuba...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+3nMAssay Description:Inhibition of HIV1 protease expressed in Escherichia coli using Arg-Glu (EDANS)-Ser-GlnAsn-Tyr-Pro-Ile-Val-Gln-Lys(DABCYL)-Arg as substrate preincuba...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+3nMAssay Description:Inhibition of HIV1 protease expressed in Escherichia coli using Arg-Glu (EDANS)-Ser-GlnAsn-Tyr-Pro-Ile-Val-Gln-Lys(DABCYL)-Arg as substrate preincuba...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+3nMAssay Description:Inhibition of HIV1 protease expressed in Escherichia coli using Arg-Glu (EDANS)-Ser-GlnAsn-Tyr-Pro-Ile-Val-Gln-Lys(DABCYL)-Arg as substrate preincuba...More data for this Ligand-Target Pair